Abstract
A non-enveloped virus requires a membrane lesion to deliver its genome into a target cell1. For rotaviruses, membrane perforation is a principal function of the viral outer-layer protein, VP42,3. Here we describe the use of electron cryomicroscopy to determine how VP4 performs this function and show that when activated by cleavage to VP8* and VP5*, VP4 can rearrange on the virion surface from an ‘upright’ to a ‘reversed’ conformation. The reversed structure projects a previously buried ‘foot’ domain outwards into the membrane of the host cell to which the virion has attached. Electron cryotomograms of virus particles entering cells are consistent with this picture. Using a disulfide mutant of VP4, we have also stabilized a probable intermediate in the transition between the two conformations. Our results define molecular mechanisms for the first steps of the penetration of rotaviruses into the membranes of target cells and suggest similarities with mechanisms postulated for other viruses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The maps of the cryo-EM reconstructions have been deposited in the Electron Microscopy Data Bank (EMDB) (accession numbers: EMD-21955, upright conformation; EMD-21956, intermediate conformation; EMD-21957, reversed conformation), and the refined coordinates in the Protein Data Bank (PDB) (PDB IDs: 6WXE, upright conformation; 6WXF, intermediate conformation; 6WXG, reversed conformation). We obtained previously published rotavirus structures from the PDB (PDB IDs: 4V7Q and 1SLQ). The rotavirus protein sequences used to prepare the sequence alignments (shown in Supplementary Data 1–3) were retrieved from GenBank46. The accession numbers for VP4 are: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. The accession numbers for VP7 are: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. The accession numbers for VP6 are: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. All data are available from the corresponding author upon reasonable request.
Code availability
The software programs used to generate and analyse the data of this study are publicly available. Custom-written C shell and Python scripts used to run the programs are available from the corresponding author on reasonable request.
References
Harrison, S. C. in Fields Virology 6th edn (eds Knipe, D. M. & Howley, P. M.) 52–86 (Lippincott Williams and Wilkins, 2013).
Estes, M. K. & Greenberg, H. in Fields Virology 6th edn (eds Knipe, D. M. & Howley, P. M.) 1347–1401 (Lippincott Williams and Wilkins, 2013).
Trask, S. D., Ogden, K. M. & Patton, J. T. Interactions among capsid proteins orchestrate rotavirus particle functions. Curr. Opin. Virol. 2, 373–379 (2012).
Tihova, M., Dryden, K. A., Bellamy, A. R., Greenberg, H. B. & Yeager, M. Localization of membrane permeabilization and receptor binding sites on the VP4 hemagglutinin of rotavirus: implications for cell entry. J. Mol. Biol. 314, 985–992 (2001).
Kim, I. S., Trask, S. D., Babyonyshev, M., Dormitzer, P. R. & Harrison, S. C. Effect of mutations in VP5 hydrophobic loops on rotavirus cell entry. J. Virol. 84, 6200–6207 (2010).
Settembre, E. C., Chen, J. Z., Dormitzer, P. R., Grigorieff, N. & Harrison, S. C. Atomic model of an infectious rotavirus particle. EMBO J. 30, 408–416 (2011).
Aoki, S. T. et al. Structure of rotavirus outer-layer protein VP7 bound with a neutralizing Fab. Science 324, 1444–1447 (2009).
Abdelhakim, A. H. et al. Structural correlates of rotavirus cell entry. PLoS Pathog. 10, e1004355 (2014).
Trask, S. D., Kim, I. S., Harrison, S. C. & Dormitzer, P. R. A rotavirus spike protein conformational intermediate binds lipid bilayers. J. Virol. 84, 1764–1770 (2010).
Dormitzer, P. R. et al. Specificity and affinity of sialic acid binding by the rhesus rotavirus VP8* core. J. Virol. 76, 10512–10517 (2002).
Delorme, C. et al. Glycosphingolipid binding specificities of rotavirus: identification of a sialic acid-binding epitope. J. Virol. 75, 2276–2287 (2001).
Martínez, M. A., López, S., Arias, C. F. & Isa, P. Gangliosides have a functional role during rotavirus cell entry. J. Virol. 87, 1115–1122 (2013).
Ramani, S. et al. The VP8* domain of neonatal rotavirus strain G10P[11] binds to type II precursor glycans. J. Virol. 87, 7255–7264 (2013).
Hu, L. et al. Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen. Nature 485, 256–259 (2012).
Dormitzer, P. R., Greenberg, H. B. & Harrison, S. C. Purified recombinant rotavirus VP7 forms soluble, calcium-dependent trimers. Virology 277, 420–428 (2000).
Salgado, E. N., Garcia Rodriguez, B., Narayanaswamy, N., Krishnan, Y. & Harrison, S. C. Visualization of calcium ion loss from rotavirus during cell entry. J. Virol. 92, e01327-18 (2018).
Rodríguez, J. M. et al. New insights into rotavirus entry machinery: stabilization of rotavirus spike conformation is independent of trypsin cleavage. PLoS Pathog. 10, e1004157 (2014).
Dormitzer, P. R., Nason, E. B., Prasad, B. V. & Harrison, S. C. Structural rearrangements in the membrane penetration protein of a non-enveloped virus. Nature 430, 1053–1058 (2004).
Pesavento, J. B., Crawford, S. E., Roberts, E., Estes, M. K. & Prasad, B. V. pH-induced conformational change of the rotavirus VP4 spike: implications for cell entry and antibody neutralization. J. Virol. 79, 8572–8580 (2005).
Salgado, E. N., Upadhyayula, S. & Harrison, S. C. Single-particle detection of transcription following rotavirus entry. J. Virol. 91, e00651-17 (2017).
Smith, R. E., Zweerink, H. J. & Joklik, W. K. Polypeptide components of virions, top component and cores of reovirus type 3. Virology 39, 791–810 (1969).
Street, J. E., Croxson, M. C., Chadderton, W. F. & Bellamy, A. R. Sequence diversity of human rotavirus strains investigated by northern blot hybridization analysis. J. Virol. 43, 369–378 (1982).
Fiore, L. et al. Antigenicity, immunogenicity and passive protection induced by immunization of mice with baculovirus-expressed VP7 protein from rhesus rotavirus. J. Gen. Virol. 76, 1981–1988 (1995).
Mackow, E. R., Barnett, J. W., Chan, H. & Greenberg, H. B. The rhesus rotavirus outer capsid protein VP4 functions as a hemagglutinin and is antigenically conserved when expressed by a baculovirus recombinant. J. Virol. 63, 1661–1668 (1989).
Trask, S. D. & Dormitzer, P. R. Assembly of highly infectious rotavirus particles recoated with recombinant outer capsid proteins. J. Virol. 80, 11293–11304 (2006).
Greenberg, H. B. et al. Production and preliminary characterization of monoclonal antibodies directed at two surface proteins of rhesus rotavirus. J. Virol. 47, 267–275 (1983).
Padilla-Noriega, L. et al. Serologic analysis of human rotavirus serotypes P1A and P2 by using monoclonal antibodies. J. Clin. Microbiol. 31, 622–628 (1993).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Bell, J. M., Chen, M., Baldwin, P. R. & Ludtke, S. J. High resolution single particle refinement in EMAN2.1. Methods 100, 25–34 (2016).
Jenni, S. et al. In situ structure of rotavirus VP1 RNA-dependent RNA polymerase. J. Mol. Biol. 431, 3124–3138 (2019).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014).
Ding, K. et al. In situ structures of rotavirus polymerase in action and mechanism of mRNA transcription and release. Nat. Commun. 10, 2216 (2019).
Mastronarde, D. N. & Held, S. R. Automated tilt series alignment and tomographic reconstruction in IMOD. J. Struct. Biol. 197, 102–113 (2017).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Iancu, C. V. et al. Electron cryotomography sample preparation using the Vitrobot. Nat. Protoc. 1, 2813–2819 (2006).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Galaz-Montoya, J. G., Flanagan, J., Schmid, M. F. & Ludtke, S. J. Single particle tomography in EMAN2. J. Struct. Biol. 190, 279–290 (2015).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 46, D41–D47 (2018).
Cock, P. J. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Gouet, P., Courcelle, E., Stuart, D. I. & Métoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
Gorziglia, M., Larralde, G., Kapikian, A. Z. & Chanock, R. M. Antigenic relationships among human rotaviruses as determined by outer capsid protein VP4. Proc. Natl Acad. Sci. USA 87, 7155–7159 (1990).
Martella, V. et al. Molecular analysis of the VP7, VP4, VP6, NSP4, and NSP5/6 genes of a buffalo rotavirus strain: identification of the rare P[3] rhesus rotavirus-like VP4 gene allele. J. Clin. Microbiol. 41, 5665–5675 (2003).
Matthijnssens, J. et al. Full genome-based classification of rotaviruses reveals a common origin between human Wa-like and porcine rotavirus strains and human DS-1-like and bovine rotavirus strains. J. Virol. 82, 3204–3219 (2008).
Patton, J. T. Rotavirus diversity and evolution in the post-vaccine world. Discov. Med. 13, 85–97 (2012).
Afonine, P. V. phenix.mtriage: a tool for analysis and validation of cryo-EM 3D reconstructions. Comput. Crystallogr. Newsl. 8, 25 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Acknowledgements
We thank Z. Li, S. Sterling, R. Walsh and S. Rawson for assistance and guidance at the Harvard Medical School Cryo-EM Center for Structural Biology and the Harvard Medical School Molecular Electron Microscopy Suite; C. Xu for assistance at the Brandeis University cryo-EM facility; and H. B. Greenberg for the gift of the HS1 and HS2 antibodies. The work was supported by National Institutes of Health grant CA-13202 (to S.C.H.). S.C.H. is an Investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.H., E.N.S., D.N., S.J. and S.C.H. designed the experiments; T.H., R.T., E.N.S. and C.B. conducted the experiments and recorded data; T.H., R.T., D.S. and S.J. analysed the data; T.H. and S.J. determined structures and built models; T.H., S.J. and S.C.H. wrote the paper; and T.H., C.B., D.S., D.N., S.J. and S.C.H. revised and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks David Bhella, John Patton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sample preparation and cryo-EM data collection.
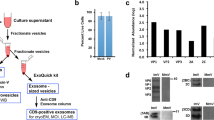
a, Schematic protocol for recoating of double-layer particles (DLPs) with recombinant VP4 and VP7. b, Time course of the digestion of wild-type recoated triple-layer particles (wt rcTLPs) with 5 μg ml−1 trypsin at 37 °C. Samples were analysed after the shown incubation times by SDS–PAGE. The experiment was repeated independently twice with similar results. For gel source data, see Supplementary Fig. 1. c, Representative micrograph (aligned and summed movie frames) of wt rcTLPs recorded with a Polara F30 electron microscope equipped with a K2 summit detector (magnification, 40,650). Scale bar, 100 nm. d, Power spectrum of the micrograph shown in c.
Extended Data Fig. 2 Data processing for local reconstructions.
Full workflow for local reconstructions of rotavirus spike proteins.
Extended Data Fig. 3 Resolution analysis of the local cryo-EM reconstructions.
Left, Fourier shell correlation (FSC) curves for the reconstructions and refined models calculated with phenix.mtriage54. Correlations for the two half maps are shown as solid blue lines after applying a mask encompassing the models. Correlations between the refined model and final map are shown as red solid lines. Nominal resolution estimates at conventional FSC values are indicated by arrows. Right, local resolution of the reconstructions calculated with ResMap55. a, Upright conformation. Dashed lines (left) are the FSC analysis for the reconstruction of the distal VP5*/VP8* dimeric density that was obtained through alignment by classification (see Methods). The images on the right show the local resolution of this reconstruction. b, Intermediate conformation. c, Reversed conformation. For source data of the FSC plots, see Supplementary Data 5.
Extended Data Fig. 4 Cryo-EM density and structure comparison.
a, Magnified views of representative regions of the cryo-EM density maps obtained by local reconstruction. Density is shown as grey mesh; polypeptide-chain backbone as ribbon; side-chain atoms as sticks (carbon, main color; nitrogen, blue; oxygen, red; sulfur, orange). b, Per-residue Cα distances after subunit-wise superposition of VP4, VP7 and VP6 subunits from the upright and reversed conformation structures.
Extended Data Fig. 5 Comparison of penetration protein conformations on rcTLPs and native TLPs.
Relative subparticle amounts, and corresponding cryo-EM reconstructions obtained from rcTLPs (top row) and native TLPs (bottom row). Local spike reconstructions of rcTLPs were obtained from two cryo-EM samples prepared from two independent recoating reactions. Local spike reconstructions of native TLPs were obtained from one cryo-EM sample.
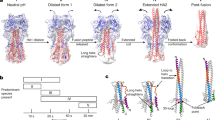
Extended Data Fig. 6 Rearrangements of the VP8* and VP5* penetration proteins during transition from upright to reversed conformation on the virion surface.
a, Distinct domains of the VP8* and VP5* spike proteins are coloured separately to illustrate their conformational change during transition from upright (top row) to reversed (bottom row) conformation. Domains that were not observed in our cryo-EM maps because of flexible attachment are drawn schematically. VP8*, magenta; VP5*, red, orange and salmon. b, Formation of the trimeric coiled coil and extrusion of the foot domains. Top row, magnified views of the VP5* foot domain exit sites as observed in the intermediate conformation structure. A partially cut surface representation is shown. VP5*, red, orange and salmon; VP7, yellow. The last modelled residues of the VP5* β-barrel domains—482 (chain 1), 480 (chain 2) and 481 (chain 3)—are located on the outside. The connections to the first modelled VP5* foot domain residue—494 (chain 1), 494 (chain 2) and 498 (chain 3)—are indicated by dashed lines (fuzzy density in the cryo-EM map). Arrows indicate a suggested mechanism for foot-domain reversal, involving zipping-up of the trimeric coiled coil and unfolding and extrusion of foot-domain residues. Bottom row, proposed transition from the intermediate structure (left) to the reversed structure by zipping up of the coiled coil and unfolding and extrusion of the foot domains.
Extended Data Fig. 7 Molecular details of the VP5*–VP7 interfaces for upright and reversed conformations.
a, Upright conformation; b, reversed conformation. In each part of the figure, the left-hand image is a view from outside the TLP, with circles in black and blue corresponding to coloured outlines of the detailed, side-view panels on the right. VP5*, salmon; VP7, yellow. The following VP5* interface residues are conserved (Supplementary Data 1): N268, N376, R467, S469. The VP5* β-barrel N terminus (residues 248–250) is not strictly conserved, but the interaction is based on main-chain hydrogen bonds and can probably be maintained for different side chains as well. The following VP7 interface residues are conserved (Supplementary Data 2): S201, T210, L172–Y175.
Extended Data Fig. 8 Inducing penetration protein reversal at alkaline pH and VP8* association with TLPs.
a, Analysis of rotavirus particles without (lanes 1 and 2) and with (lanes 3 and 4) high pH-induced conformational change of VP8* or VP5* and without (lanes 1 and 3) and with (lanes 2 and 4) EDTA-induced uncoating. Pelleted fractions were analysed by SDS–PAGE and silver staining (left) and by western blotting with the VP8*-specific antibody HS127 (right) (see Methods). The experiment was repeated independently three times with similar results. For gel and western blot source data, see Supplementary Fig. 1. b, Relative VP4 subparticle amounts, and corresponding cryo-EM reconstructions obtained from wild-type rcTLPs. c, Relative VP4 subparticle amounts, and corresponding cryo-EM reconstructions obtained from rcTLPs containing VP4(S567C/A590C). Recoating reactions for all samples were carried out at the same time and with the same VP7 and DLP stock solutions. All cryo-EM samples were prepared in the same blotting session.
Extended Data Fig. 9 Cryo-ET analysis of RRVs entering BSC-1 cells.
a, Sections of the tomogram from which the reconstructions (icosahedral average of single virion subtomograms) in Fig. 4a were obtained. Left, virus with loose membrane contact indicated by an arrow. Right, virus with tight (close) membrane contact indicated by an arrow. Images were low-pass filtered and contrast enhanced for display. Scale bar, 100 nm. b, Tomographic slices of manually selected viruses (not including particle in a) from several tomograms with tight membrane contacts (yellow arrowheads). Images were low-pass filtered and contrast-enhanced for display. Those particles were selected to detect VP4 reversed conformations in additional viruses (other than particle in Fig. 4a). c, Sub-subtomogram classification of VP4 positions extracted from the RRV particles shown in b yielded a class with the upright conformation (left) and a class with the reversed conformation (right). VP4 positions were extracted from 26 selected particles chosen to have an easily identified region of close membrane contact. The particle shown in Fig. 4a was excluded from this selection. Note that the reconstructions shown in Fig. 4a are from single particles with imposed icosahedral symmetry, whereas we show here classified averaged individual volumes extracted at VP4 positions.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion.
Supplementary Figure
Supplementary Figure 1: This file shows the Western blot and gel source data for Fig. 3d, Extended Data Fig. 1b, and Extended Data Fig. 8a.
Supplementary Data
Supplementary Data 1: Rotavirus VP4 multiple sequence alignment from GenBank accession identifiers: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. Modeled residues are shown as solid bars.
Supplementary Data
Supplementary Data 2: Rotavirus VP7 multiple sequence alignment from GenBank accession identifiers: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. Modeled residues are shown as solid bars.
Supplementary Data
Supplementary Data 3: Rotavirus VP6 multiple sequence alignment from GenBank accession identifiers: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. Modeled residues are shown as solid bars.
Supplementary Data
Supplementary Data 4: This file contains the source data for the bar plot shown in Fig. 3a.
Supplementary Data
Supplementary Data 5: This file contains the source data for the Fourier shell correlation (FSC) plots shown in Extended Data Fig. 3.
Rights and permissions
About this article
Cite this article
Herrmann, T., Torres, R., Salgado, E.N. et al. Functional refolding of the penetration protein on a non-enveloped virus. Nature 590, 666–670 (2021). https://doi.org/10.1038/s41586-020-03124-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-03124-4
This article is cited by
-
Cryo-EM structures of Banna virus in multiple states reveal stepwise detachment of viral spikes
Nature Communications (2024)
-
mRNA-based VP8* nanoparticle vaccines against rotavirus are highly immunogenic in rodents
npj Vaccines (2023)
-
Visualizing molecular interactions that determine assembly of a bullet-shaped vesicular stomatitis virus particle
Nature Communications (2022)
-
Novel fold of rotavirus glycan-binding domain predicted by AlphaFold2 and determined by X-ray crystallography
Communications Biology (2022)
-
Multiple conformations of trimeric spikes visualized on a non-enveloped virus
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.