Abstract
Selective targeting of aneuploid cells is an attractive strategy for cancer treatment1. However, it is unclear whether aneuploidy generates any clinically relevant vulnerabilities in cancer cells. Here we mapped the aneuploidy landscapes of about 1,000 human cancer cell lines, and analysed genetic and chemical perturbation screens2,3,4,5,6,7,8,9 to identify cellular vulnerabilities associated with aneuploidy. We found that aneuploid cancer cells show increased sensitivity to genetic perturbation of core components of the spindle assembly checkpoint (SAC), which ensures the proper segregation of chromosomes during mitosis10. Unexpectedly, we also found that aneuploid cancer cells were less sensitive than diploid cells to short-term exposure to multiple SAC inhibitors. Indeed, aneuploid cancer cells became increasingly sensitive to inhibition of SAC over time. Aneuploid cells exhibited aberrant spindle geometry and dynamics, and kept dividing when the SAC was inhibited, resulting in the accumulation of mitotic defects, and in unstable and less-fit karyotypes. Therefore, although aneuploid cancer cells could overcome inhibition of SAC more readily than diploid cells, their long-term proliferation was jeopardized. We identified a specific mitotic kinesin, KIF18A, whose activity was perturbed in aneuploid cancer cells. Aneuploid cancer cells were particularly vulnerable to depletion of KIF18A, and KIF18A overexpression restored their response to SAC inhibition. Our results identify a therapeutically relevant, synthetic lethal interaction between aneuploidy and the SAC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
The code used to generate and/or analyse the data are publicly available, or available upon request.
Data availability
All datasets are available within the article and its Supplementary Information, or from the Corresponding Author upon request. Cell line aneuploidy profiles and scores are available at the DepMap portal (https://depmap.org/portal/). The analysed CCLE genomic data are available at https://doi.org/10.6084/m9.figshare.11384241.v2. LP-WGS data have been deposited to SRA with BioProject accession number PRJNA672256.
References
Ben-David, U. & Amon, A. Context is everything: aneuploidy in cancer. Nat. Rev. Genet. 21, 44–62 (2020).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e16 (2017).
McDonald, E. R. III et al. Project DRIVE: a compendium of cancer dependencies and synthetic lethal relationships uncovered by large-scale, deep RNAi screening. Cell 170, 577–592.e10 (2017).
Aguirre, A. J. et al. Genomic copy number dictates a gene-independent cell response to CRISPR/Cas9 targeting. Cancer Discov. 6, 914–929 (2016).
Yang, W. et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 41, D955–D961 (2013).
Basu, A. et al. An interactive resource to identify cancer genetic and lineage dependencies targeted by small molecules. Cell 154, 1151–1161 (2013).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Corsello, S. M. et al. Discovering the anticancer potential of non-oncology drugs by systematic viability profiling. Nat. Cancer 1, 235–248 (2020).
Musacchio, A. & Salmon, E. D. The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8, 379–393 (2007).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33, 676–689.e3 (2018).
Knouse, K. A., Wu, J., Whittaker, C. A. & Amon, A. Single cell sequencing reveals low levels of aneuploidy across mammalian tissues. Proc. Natl Acad. Sci. USA 111, 13409–13414 (2014).
Oromendia, A. B., Dodgson, S. E. & Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 26, 2696–2708 (2012).
Torres, E. M. et al. Effects of aneuploidy on cellular physiology and cell division in haploid yeast. Science 317, 916–924 (2007).
Hwang, S. et al. Serine-dependent sphingolipid synthesis is a metabolic liability of aneuploid cells. Cell Rep. 21, 3807–3818 (2017).
Storchová, Z. et al. Genome-wide genetic analysis of polyploidy in yeast. Nature 443, 541–547 (2006).
Ben-David, U. et al. Patient-derived xenografts undergo mouse-specific tumor evolution. Nat. Genet. 49, 1567–1575 (2017).
Kops, G. J. P. L., Foltz, D. R. & Cleveland, D. W. Lethality to human cancer cells through massive chromosome loss by inhibition of the mitotic checkpoint. Proc. Natl Acad. Sci. USA 101, 8699–8704 (2004).
Wild, T. et al. The spindle assembly checkpoint is not essential for viability of human cells with genetically lowered APC/C activity. Cell Rep. 14, 1829–1840 (2016).
Chunduri, N. K. & Storchová, Z. The diverse consequences of aneuploidy. Nat. Cell Biol. 21, 54–62 (2019).
Baker, D. J., Jin, F., Jeganathan, K. B. & van Deursen, J. M. Whole chromosome instability caused by Bub1 insufficiency drives tumorigenesis through tumor suppressor gene loss of heterozygosity. Cancer Cell 16, 475–486 (2009).
Ricke, R. M., Jeganathan, K. B. & van Deursen, J. M. Bub1 overexpression induces aneuploidy and tumor formation through Aurora B kinase hyperactivation. J. Cell Biol. 193, 1049–1064 (2011).
Foijer, F. et al. Chromosome instability induced by Mps1 and p53 mutation generates aggressive lymphomas exhibiting aneuploidy-induced stress. Proc. Natl Acad. Sci. USA 111, 13427–13432 (2014).
Foijer, F. et al. Deletion of the MAD2L1 spindle assembly checkpoint gene is tolerated in mouse models of acute T-cell lymphoma and hepatocellular carcinoma. eLife 6, e20873 (2017).
Sotillo, R. et al. Mad2 overexpression promotes aneuploidy and tumorigenesis in mice. Cancer Cell 11, 9–23 (2007).
Sotillo, R., Schvartzman, J. M., Socci, N. D. & Benezra, R. Mad2-induced chromosome instability leads to lung tumour relapse after oncogene withdrawal. Nature 464, 436–440 (2010).
Mason, J. M. et al. Functional characterization of CFI-402257, a potent and selective Mps1/TTK kinase inhibitor, for the treatment of cancer. Proc. Natl Acad. Sci. USA 114, 3127–3132 (2017).
Pauer, L. R. et al. Phase I study of oral CI-994 in combination with carboplatin and paclitaxel in the treatment of patients with advanced solid tumors. Cancer Invest. 22, 886–896 (2004).
Bielski, C. M. et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat. Genet. 50, 1189–1195 (2018).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Hieronymus, H. et al. Tumor copy number alteration burden is a pan-cancer prognostic factor associated with recurrence and death. eLife 7, e37294 (2018).
Smith, J. C. & Sheltzer, J. M. Systematic identification of mutations and copy number alterations associated with cancer patient prognosis. eLife 7, e39217 (2018).
Soto, M., García-Santisteban, I., Krenning, L., Medema, R. H. & Raaijmakers, J. A. Chromosomes trapped in micronuclei are liable to segregation errors. J. Cell Sci. 131, jcs214742 (2018).
Zhang, C. Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
Burgess, A., Rasouli, M. & Rogers, S. Stressing mitosis to death. Front. Oncol. 4, 140 (2014).
Dominguez-Brauer, C. et al. Targeting mitosis in cancer: emerging strategies. Mol. Cell 60, 524–536 (2015).
Santaguida, S., Tighe, A., D’Alise, A. M., Taylor, S. S. & Musacchio, A. Dissecting the role of MPS1 in chromosome biorientation and the spindle checkpoint through the small molecule inhibitor reversine. J. Cell Biol. 190, 73–87 (2010).
Kuznetsova, A. Y. et al. Chromosomal instability, tolerance of mitotic errors and multidrug resistance are promoted by tetraploidization in human cells. Cell Cycle 14, 2810–2820 (2015).
Santaguida, S. et al. Chromosome mis-segregation generates cell-cycle-arrested cells with complex karyotypes that are eliminated by the immune system. Dev. Cell 41, 638–651.e5 (2017).
Bakker, B. et al. Single-cell sequencing reveals karyotype heterogeneity in murine and human malignancies. Genome Biol. 17, 115 (2016).
Weaver, L. N. et al. Kif18A uses a microtubule binding site in the tail for plus-end localization and spindle length regulation. Curr. Biol. 21, 1500–1506 (2011).
Czechanski, A. et al. Kif18a is specifically required for mitotic progression during germ line development. Dev. Biol. 402, 253–262 (2015).
Wordeman, L., Decarreau, J., Vicente, J. J. & Wagenbach, M. Divergent microtubule assembly rates after short- versus long-term loss of end-modulating kinesins. Mol. Biol. Cell 27, 1300–1309 (2016).
Marquis, C. et al. Chromosomally unstable tumor cells specifically require KIF18A for proliferation. Nat. Comm. (in the press).
Quinton, R. J. et al. Whole-genome doubling confers unique genetic vulnerabilities on tumour cells. Nature https://doi.org/10.1038/s41586-020-03133-3 (2021).
Braun, J. et al. Synthesis and biological evaluation of optimized inhibitors of the mitotic kinesin Kif18A. ACS Chem. Biol. 10, 554–560 (2015).
McFarland, J. M. et al. Improved estimation of cancer dependencies from large-scale RNAi screens using model-based normalization and data integration. Nat. Commun. 9, 4610 (2018).
Seashore-Ludlow, B. et al. Harnessing connectivity in a large-scale small-molecule sensitivity dataset. Cancer Discov. 5, 1210–1223 (2015).
Rees, M. G. et al. Correlating chemical sensitivity and basal gene expression reveals mechanism of action. Nat. Chem. Biol. 12, 109–116 (2016).
Iorio, F. et al. A landscape of pharmacogenomic interactions in cancer. Cell 166, 740–754 (2016).
Nusinow, D. P. et al. Quantitative proteomics of the cancer cell line encyclopedia. Cell 180, 387–402.e16 (2020).
Chan, E. M. et al. WRN helicase is a synthetic lethal target in microsatellite unstable cancers. Nature 568, 551–556 (2019).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Sheltzer, J. M. A transcriptional and metabolic signature of primary aneuploidy is present in chromosomally unstable cancer cells and informs clinical prognosis. Cancer Res. 73, 6401–6412 (2013).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
R core team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2018).
Huang, D. W. et al. DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35, W169–W175 (2007).
Dempster, J. M. et al. Extracting biological insights from the Project Achilles Genome-Scale CRISPR Screens in cancer cell lines. Preprint at https://doi.org/10.1101/720243 (2019).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Mayr, M. I. et al. The human kinesin Kif18A is a motile microtubule depolymerase essential for chromosome congression. Curr. Biol. 17, 488–498 (2007).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452.e17 (2017).
Ben-David, U. et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560, 325–330 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at https://doi.org/10.1101/060012 (2016).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Dürrbaum, M. et al. Unique features of the transcriptional response to model aneuploidy in human cells. BMC Genomics 15, 139 (2014).
Lee, K. & Rhee, K. PLK1 phosphorylation of pericentrin initiates centrosome maturation at the onset of mitosis. J. Cell Biol. 195, 1093–1101 (2011).
Lai, D. & Shah, S. HMMcopy: copy number prediction with correction for GC and mappability bias for HTS data. https://rdrr.io/bioc/HMMcopy/ (2012).
Knouse, K. A., Wu, J. & Amon, A. Assessment of megabase-scale somatic copy number variation using single-cell sequencing. Genome Res. 26, 376–384 (2016).
van den Bos, H. et al. in Cellular Senescence. Methods in Molecular Biology (ed. Demaria, M.) (Humana, 2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Jun, G., Wing, M. K., Abecasis, G. R. & Kang, H. M. An efficient and scalable analysis framework for variant extraction and refinement from population-scale DNA sequence data. Genome Res. 25, 918–925 (2015).
van den Bos, H. et al. Single-cell whole genome sequencing reveals no evidence for common aneuploidy in normal and Alzheimer’s disease neurons. Genome Biol. 17, 116 (2016).
Acknowledgements
We acknowledge J. Bryan, J. Roth and S. Bender for assistance with PRISM; A. Cherniack and M. Kocak for helpful discussions; D. Lam and O. Enache for assistance with the L1000 assay; A. Y. Kuznetsova for the initial characterization of the HPT and RPT cell lines; H. van den Bos for assistance with cell sorting; and S. Tsach for assistance with figure preparation. This work was supported by the Azrieli Foundation (U.B.-D.), the Richard Eimert Research Fund on Solid Tumors (U.B.-D.), the Tel-Aviv University Cancer Biology Research Center (U.B.-D.), the Israel Cancer Association (U.B.-D.), and the DoD CDMRP career development award (CA191148 to U.B.-D.). Work in the Santaguida laboratory is supported by grants from the Italian Association for Cancer Research (MFAG 2018, ID. 21665), Ricerca Finalizzata (GR-2018-12367077), Fondazione Cariplo, the Rita-Levi Montalcini program from MIUR, and the Italian Ministry of Health. Work in the Stumpff laboratory is supported by a Susan G. Komen grant (CCR16377648 to J.S.), an NIH grant (GM121491 to J.S.), and a DoD PRCRP Horizon Award (W81XWH-17-1-0371 to H.L.H.M.). We thank the late A. Amon for groundbreaking research on aneuploidy, and for support and inspiration.
Author information
Authors and Affiliations
Contributions
U.B.-D. conceived the project. Y.C.-S., M.A., H.T. and U.B.-D. performed the cell culture experiments. K.L and J.Z. assisted with cell culture experiments. C.M., H.L.H.M. and J.S. performed microscopy experiments. Z.S. provided the HCT116/HPT and RPE1/RPT cell lines, and together with S.V.B. and L.-M.S. characterized them and performed microscopy experiments. S.S. provided aneuploid RPE1 cell lines, and together with M.R.I. characterized them and examined their sensitivity to SAC inhibition. I.B., R.W., D.C.J.S. and F.F. generated, processed and assisted in the analysis of the single-cell DNA sequencing data. J.M.M., M.K. and U.B.-D. performed the computational analyses. N.L. assisted with the generation of the gene expression data, and A.J. assisted with their analysis. A.N. and A.J.B. shared data. F.F., R.B., S.S. T.R.G., J.S., Z.S. and U.B.-D. supervised the experiments and analyses that were conducted in their respective laboratories. U.B.-D. directed the project and wrote the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
T.R.G. is a consultant to GlaxoSmithKline and is a founder of Sherlock Biosciences. R.B. owns shares in Ampressa and receives grant funding from Novartis. A.J.B. receives funding from Merck, Bayer and Novartis, and is an advisor to Earli and Helix Nano and a co-founder of Signet Therapeutics. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks Ryoma Ohi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Increased sensitivity of aneuploid cancer cells to genetic inhibition of the spindle assembly checkpoint.
a, Copy number profiles of 5 representative breast cancer cell lines from the highly-aneuploid cell line group (top quartile of AS) and 5 representative breast cancer cell lines from the near-euploid cell line group (bottom quartile of AS). b, A volcano plot showing the differential genetic dependencies between the near-euploid and highly-aneuploid cancer cell lines (top vs. bottom quartiles), based on the genome-wide DRIVE RNAi screen4. BUB1B and MAD2, core members of the SAC, are highlighted in red. c, A Venn diagram showing the overlap of the differentially-dependent genes (q<0.25) between the Achilles and DRIVE RNAi screens. ****P = 1e-16, two-tailed Fisher’s exact test. d, The pathways enriched in the list of genes that are more essential in near-euploid than in highly-aneuploid cancer cell lines (effect size <-0.1, q < 0.1) in the DRIVE RNAi screen, based on DAVID functional annotation enrichment analysis59. The full list is available in Supplementary Table 3. *, Benjamini-corrected p-value <0.1; one-tailed Fisher’s Exact Test. e, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (left) and MAD2 (right) in the DRIVE RNAi screen. The more negative a value, the more essential the gene is in that cell line. ****P = 2e-06 and P = 1e-04 for BUB1B and MAD2, respectively; two-tailed t-test. f, A volcano plot showing the differential genetic dependencies between the near-euploid and highly-aneuploid cancer cell lines (top vs. bottom 10% of cell lines), based on the genome-wide DRIVE RNAi screen4. BUB1B, MAD2 and KIF18A are highlighted in red. g, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (left) and MAD2 (right) in the DRIVE RNAi screen (top vs. bottom 10% of cell lines). The more negative a value, the more essential the gene is in that cell line. *P = 0.037; ***P = 5e-04; two-tailed t-test. h, Comparison of protein expression levels of BUB1B (left) and MAD2 (right) between near-euploid and highly-aneuploid cancer cell lines. n.s., P > 0.05; **P = 0.001; for BUB1B and MAD2, respectively; two-tailed t-test. i, The correlations between the mRNA expression levels of BUB1B (top) and MAD2 (bottom) and the genetic dependency on these genes in the Achilles (left) and DRIVE (right) RNAi screens. Spearman’s ρ = 0.36 (P = 3e-08), 0.31 (P = 2e-06), 0.26 (P = 4e-04) and 0.40 (P = 2e-08), respectively. j, The correlations between the protein expression levels of BUB1B (top) and MAD2 (bottom) and the genetic dependency on these genes in the Achilles (left) and DRIVE (right) RNAi screens. Spearman’s ρ = 0.11 (P = 0.09), 0.26 (P = 4e-05), 0.14 (P = 0.016) and 0.24 (P = 5e-05), respectively. k, The mRNA expression levels of BUB1B (left) and MAD2 (right) in near-euploid and highly-aneuploid cancer cell lines across multiple cell lineages. *P < 0.05; two-tailed t-test.
Extended Data Fig. 2 Genomic and phenotypic features associated with the degree of aneuploidy in human cancer cell lines.
a, The AS distribution across 23 cancer types. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR, individual cell lines. b, Comparison of AS between cancer cell lines with distinct TP53 mutation status (based on CCLE annotations)2. **** P = 6e-15 and P = 1e-22 for the comparisons between TP53-WT and ‘damaging’ and TP53-WT and ‘hotspot’ mutations, respectively; two-tailed t-test. c, Comparison of AS between cancer cell lines with distinct genome doubling (WGD) status. ****P = 1e-192, P = 2e-96 and P = 6e-13 for the comparisons between WGD = 0 and WGD = 1, WGD = 0 and WGD = 2, and WGD = 1 and WGD = 2, respectively; two-tailed t-test. d, Comparison of the HET70 score, a measure of karyotypic instability56, between the near-diploid and highly-aneuploid cell line groups. ****P = 2e-08; two-tailed t-test. e, Comparison of doubling time between the near-diploid and highly-aneuploid cell line groups. **P = 0.005; two-tailed t-test.
Extended Data Fig. 3 Increased sensitivity of aneuploid cancer cells to SACi remains significant when associated genomic and phenotypic features are controlled for.
a, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens across multiple cell lineages. *P < 0.05; **P < 0.01; two-tailed t-test. b, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens, after accounting for lineage-specific differences in gene dependency scores using linear regression. ***P = 2e-04; *P = 0.013; for RNAi-Achilles BUB1B and MAD2 dependencies, respectively; ***P = 5e-04; *P = 0.044; RNAi-DRIVE BUB1B and MAD2 dependencies, respectively; one-tailed t-test. c, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens, across TP53 mutation classes. *P < 0.05; **P < 0.01, ***P < 0.001; two-tailed t-test. d, The correlations between AS and the dependency on BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens, for cell lines that have not undergone whole-genome duplication (that is, cell lines with basal ploidy of n = 2). Spearman’s ρ = -0.32 (P = 1e-05), -0.36 (P = 7e-07), -0.30 (P = 1e-04) and -0.28 (P = 4e-04), respectively. e, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens, after removing the effect of doubling time on gene dependency scores using linear regression. ****P = 1e-05 and P = 9e-07, for RNAi-Achilles BUB1B and MAD2 dependencies, respectively; ****P = 1e-07; ** P = 0.002; for RNAi-DRIVE BUB1B and MAD2 dependencies, respectively; two-tailed t-test. f, The sensitivity of near-euploid and highly-aneuploid cancer cell lines without microsatellite instability (MSS lines only) to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens. ****P = 7e-07 and P = 2e-07, for RNAi-Achilles BUB1B and MAD2 dependencies, respectively; ****P = 6e-07, for RNAi-DRIVE BUB1B dependency; ***, P = 1e-04; two-tailed t-test. g, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens, in cell lines that are WT for the 4 genes most selectively mutated in aneuploid human tumours (after TP53)12. **P < 0.01, ***P < 0.001; ****P < 1e-04; two-tailed t-test. h, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top) and MAD2 (bottom) in the Achilles (left) and DRIVE (right) RNAi screens, after removing the effect of lineage subtype on gene dependency scores using linear regression. ***P = 4e-04; *P = 0.015, for RNAi-Achilles BUB1B and MAD2 dependencies, respectively; **P = 0.002; *P = 0.045, for RNAi-DRIVE BUB1B and MAD2 dependencies, respectively; one-tailed t-test. i, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of BUB1B (top two plots) and MAD2 (bottom two plots) in the Achilles (top) and DRIVE (bottom) RNAi screens, after removing the effect of HET70 scores on gene dependency scores using linear regression. ****P = 9e-07, P = 8e-06 and P = 5e-07 for RNAi-Achilles BUB1B, RNAi-Achilles MAD2 and RNAi-DRIVE BUB1B dependencies, respectively; **P = 0.001; two-tailed t-test.
Extended Data Fig. 4 Reduced sensitivity of aneuploid cancer cells to chemical inhibition of the spindle assembly checkpoint.
a, Volcano plots showing the differential drug sensitivities between the near-euploid and highly-aneuploid cancer cell lines, based on the large-scale GDSC6 and PRISM screens9. MPS1-IN-1 and MPI-0479605, the only SAC inhibitors included in each screen, respectively, are highlighted in red. b, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the SAC inhibitors MPS1-IN-1 and MPI-0479605 in the GDSC (left) and PRISM (right) screens. ****P = 1e-0.5; n.s., P = 0.23; two-tailed t-test. c, Experimental validation of the response of 5 near-euploid (CAL51, EN, MHHNB11, SW48 and VMCUB1) and 5 highly-aneuploid (MDAMB468, NCIH1693, PANC0813, SH10TC and A101D) cell lines to 72h exposure to the SAC inhibitor reversine. *P = 0.016, two-tailed Wilcoxon rank-sum test; n = 5 cell lines in each group. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR. d, Comparison of the sensitivity to reversine between near-euploid and highly-aneuploid cancer cell lines subjected to the PRISM cell viability assay, confirming the reduced sensitivity of highly-aneuploid cells to a 120h exposure to SAC inhibitors. n.s., P > 0.05; *P < 0.05; **P < 0.01; two-tailed t-test. e, An association analysis failed to identify a genomic biomarker of reversine sensitivity. Shown are the top 1000 genomic features identified by our model (see Methods). No feature stands out in terms of importance and/or correlation, and the overall predictive value is poor.
Extended Data Fig. 5 Isogenic model systems of near-diploid and aneuploid cell lines.
a, scDNaseq-based copy number profiling of the HPT1 and HPT2 aneuploid cell lines. b, Karyotyping-based chromosome count of the near-diploid HCT116 cells and their highly-aneuploid HPT derivatives. Each dot represents a metaphase spread. Average chromosome number: n = 45, n = 75 and n = 78, for HCT116-GFP, HPT1 and HPT2, respectively. c, Karyotyping-based chromosome count of the near-diploid RPE1 cells and their highly-aneuploid RPT derivatives. Each dot represents a metaphase spread. Average chromosome number: n = 46, n = 80 and n = 76.5, for RPE1-GFP, RPT1 and RPT2, respectively. d, Low-pass whole-genome sequencing-based karyotyping of near-diploid and aneuploid RPE1 clones. No karyotypic changes have been observed between passage 0 (p0) and passage 10 (p10) of each clone. Red, large (>5Mb) gains (log2CN >0.3); blue, large (>5Mb) losses (log2CN <-0.3).
Extended Data Fig. 6 The effect of aneuploidy on cellular sensitivity to SACi in isogenic human cell lines.
a, Left: dose response curves of the response of near-diploid HCT116 cells and their highly-aneuploid derivatives HPT cells, to the SAC inhibitor reversine following 120h of drug exposure. EC50 = 0.11μM, 0.11μM, 2.32μM and 1.06μM, for HCT116-WT, HCT116-GFP, HPT1 and HPT2, respectively. Right: dose response curves of the response of near-diploid RPE1 cells and their highly-aneuploid derivatives RPT cells, to the SAC inhibitor reversine following 120h of drug exposure. EC50 = 0.13μM, 1.82μM, 0.57μM and 2.07μM, for RPE1-GFP, RPT1, RPT3 and RPT4, respectively. *P < 0.05; **P < 0.01; ***P < 0.001; two-tailed t-test. Data represent the mean ± s.d.; n = 3 biological replicates. b, Time-lapse imaging-based proliferation curves of HCT116 and HPT cells under standard culture conditions. Data represent the mean ± s.d.; n = 3 biological replicates. c, Dose response curves of the response of HCT116 and HPT cells to three drugs with unrelated mechanisms of action. Doxorubicin EC50 = 0.61μM, 0.32μM, 1.2μM and 0.89μM; Nutlin-3 EC50 = 11.88μM, 19.28μM, 15.26μM and 65.11μM; Imatinib EC50 = 17.94μM, 19.08μM, 18.77μM and 23.31μM; for HCT116-WT, HCT116-GFP, HPT1 and HPT2, respectively. d, Relative mRNA expression levels of BUB1B, MAD2 and TTK, confirming successful siRNA-mediated knockdown of each gene in all cell lines. *P = 0.011, P = 0.012 for HCT116-GFP and HPT1, respectively; **P = 0.0019, P = 0.0015, P = 0.0039 for BUB1B in HCT116-WT, HPT1 and HPT2, respectively; P = 0.0021, P = 0.0013 for MAD2 in HCT116-WT and HPT2, respectively; P = 0.0011, P = 0.0012 for TTK in HCT116-WT and HPT2, respectively; ***P = 0004, P = 0.0005 for MAD2 and TTK in HCT116-GFP, respectively; ****P = 9e-05; one-tailed t-test; n = 3 biological replicates. Data represent the mean ± s.e.m. e, The relative viability of 4 near-diploid (CAL51, EN, MHHNB11, VMCUB1) and 3 highly-aneuploid (MDAMB468, PANC0813, SH0TA) cancer cell lines following 72h of siRNA-mediated knockdown of 3 SAC components: BUB1B, MAD2 and TTK. Results are normalized to a non-targeting siRNA control. *P = 0.010, P = 0.016, and P = 0.015, for BUB1B, MAD2 and TTK, respectively; two-tailed t-test. Error bars, s.d. f, Dose response curves of the response of the near-diploid RPE1 clone SS48 and its isogenic aneuploid clones SS51 (+Ts7, +Ts22) and SS111 (+Ts8, +Ts9, +Ts18), to the SAC inhibitor reversine following 120h of drug exposure. EC50 = 0.66μM, 1.03μM and 1.03μM, for SS48, SS51 and SS111, respectively *P < 0.05; **P < 0.01; ***P < 0.001; two-tailed t-test. Data represent the mean ± s.d.; n = 3 biological replicates.
Extended Data Fig. 7 Time-dependent increased sensitivity of aneuploid cancer cells to genetic and chemical SACi.
a, Comparison of the doubling times of HCT116 and HPT cells exposed to siRNAs against BUB1B, MAD2 or TTK. The drug effect of SACi is stronger in the near-diploid HCT116 cells at d5, but is stronger in the highly-aneuploid HPT cells at d21. b, Comparison of the doubling times of HCT116 and HPT cells exposed to the SAC inhibitors MPI-0479605 or reversine. The drug effect of SACi is stronger in the near-diploid HCT116 cells at d5, but at d21 it becomes stronger in the highly-aneuploid HPT cells. *P = 0.034, P = 0.046 and P = 0.049 for MPI-0479605 and reversine at d5 and d14, respectively; **P = 0.0015 ; one-tailed t-test; n = 2 independent cell lines. c, Representative images of cells from the drug experiment (same images as in Fig. 2g), with cell masking performed using the image analysis software ilastik61. Scale bar, 100μm. d, The relative viability of the aneuploid RPE1 clones, SS111 and SS51, following reversine exposure. The viability effect was normalized to the effect of the drug in the near-diploid RPE1 clone, SS48. The drug effect of SACi is comparable during the first week of drug exposure, but the highly-aneuploid cells become significantly more sensitive with time. *P = 0.045, **P = 0.002, P = 0.001 and P = 0.005 for the comparisons between d3 and d14, d5 and d14 and d7 and d14, respectively; two-tailed t-test. e, The relative viability of 5 near-euploid (CAL51, EN, MHHNB11, SW48 and VMCUB1) and 5 highly-aneuploid (MDAMB468, NCIH1693, PANC0813, SH10TC and A101D) cell lines to 72h and 14 days exposure to the SAC inhibitor reversine. *P = 0.012 and P = 0.037, for 3d and 14d time points, respectively; two-tailed Wilcoxon rank-sum test.
Extended Data Fig. 8 Transcriptional, cellular and karyotypic characterization of SACi in aneuploid cells.
a, The top 10 results of a Connectivity Map (CMap) query63 of the transcriptional response of HCT116 and HPT cells to the SAC inhibitors, reversine (250nM and 500nM) and MPI-0479605 (250nM). The top connection is “Cell cycle inhibition”, correctly identifying the expected mechanism of action of these compounds. GOF, gain of function; OE, overexpression; KD, knockdown. b, Functional enrichment of gene sets related to cell cycle regulation. Shown are the gene sets that were significantly more affected by SACi in the highly-aneuploid HPT1 and HPT2 cells than in the nearly-diploid HCT116-WT and HCT116-GFP cells. *P < 0.05, one-tailed Fisher’s exact test. c, Functional enrichment of gene sets related to cell death. Shown are the gene sets that were significantly more affected by SACi in the highly-aneuploid HPT1 and HPT2 cells than in the nearly-diploid HCT116-WT and HCT116-GFP cells. *P < 0.05, one-tailed Fisher’s exact test. d, The mitotic index of HCT116 and HPT cells cultured under standard conditions or exposed to the SAC inhibitor reversine (500nM) for 24h. *P = 0.035; n.s., P = 0.17; two-tailed t-test; Error bars, s.d.; n = 3 biological replicates. e, Imaging-based quantification of the prevalence of cell divisions with multipolar spindles in HCT116 and HPT cell lines cultured under standard conditions or treated with reversine (500nM) for 24hr; n = 3 biological replicates. Error bars, s.d. f, The prevalence of premature mitotic exit (cytokinesis failure) in HCT116 and HPT cells exposed to the SAC inhibitor reversine (500nM) for 24h. *P = 0.047; two-tailed Fisher’s exact test. g, Representative images of premature mitotic exit in HPT2 cells exposed to reversine (500nM). T = 0 defines nuclear envelope breakdown (NEB). Scale bar, 10μm. h, The prevalence of micronuclei formation in RPE1 and RPT cells cultured under standard conditions or exposed to the SAC inhibitor reversine (500nM) for 24h. n.s., P > 0.05; *P = 0.013 and P = 0.015 for the differences between the treated and untreated RPT1 and RPT3 cells, respectively; **P = 0.004; ***P < 0.0002; two-tailed t-test. i, The prevalence of cell divisions with multipolar spindles in RPE1 and RPT cells cultured under standard conditions or exposed to the SAC inhibitor reversine (500nM) for 24h. n.s., P > 0.05; *P = 0.028; two-tailed t-test. Error bars, s.d. j, The prevalence of premature mitotic exit (cytokinesis failure) in RPE1 and RPT cells exposed to the SAC inhibitor reversine (500nM) for 24h. *P = 0.044 and P = 0.019 for the comparisons between RPE1 and RPT1 or RPT3, respectively; two-tailed t-test. k, Chromosomal copy number states of HCT116 and HPT cells at each of the 3 time points that were sequenced by scDNaseq. Differences between the pre-treated (d0) and post-treated (d14+3) populations are highlighted. l, Chromosomal heterogeneity scores of the HCT116 and HPT cells at each of the 3 time points. Highly-heterogeneous chromosomes in the post-treated populations (d14+3) are highlighted; n = 23 chromosomes. m, Comparison of the chromosomal heterogeneity scores between the near-diploid HCT116 cells and the highly-aneuploid HPT cells. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR; circles, individual chromosomes. ****P = 2e-09; two-tailed t-test.
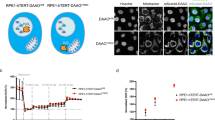
Extended Data Fig. 9 Increased sensitivity of aneuploid cancer cells to perturbation of the mitotic kinesin KIF18A.
a, Left: western blot of KIF18A protein expression levels in HCT116 and HPT cell lines. Right: Quantification of KIF18A expression levels (normalized to GAPDH). **P = 0.002; two-tailed t-test; n = 5 biological replicates. b, Left: Imaging kinetochore-bound KIF18A protein levels in HCT116-GFP, HPT1 and HPT2 cells, Scale bars, 10μm. Right: Immunofluorescence-based quantification of KIF18A protein levels. **P < 0.01, two-tailed t-test. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR. c, Schematics of the definitions of spindle length, width and angle. d, Left: Imaging-based quantification of microtubule polymerization rate in HCT116 and HPT cells cultured under standard conditions. Right: Imaging-based quantification of microtubule regrowth following complete depolymerization in HCT116 and HPT cells. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR; circles, individual cell lines. *P < 0.05; **P < 0.01; ****P < 1e-4; two-tailed t-test. e, Imaging-based quantification of EB1α-tubulin co-localization in HCT116 and HPT cells cultured under standard conditions. **P < 0.01. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR. f, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockout of KIF18A in the CRISPR-Achilles data set. The more negative a value, the more essential the gene is in that cell line. *P = 0.034; two-tailed t-test. g, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen (top vs. bottom 10% of cell lines). The more negative a value, the more essential the gene is in that cell line. ****P = 3e-06; two-tailed t-test. h, Comparison of the mRNA expression levels of KIF18A between near-euploid and highly-aneuploid cancer cell lines. n.s., P > 0.05; two-tailed t-test. i, Comparison of the protein expression levels of KIF18A between near-euploid and highly-aneuploid cancer cell lines. n.s., P > 0.05; two-tailed t-test. j, The correlation between KIF18A mRNA expression and the genetic dependency on this gene in the Achilles-RNAi screen. Spearman’s ρ = 0.17 (P = 0.004). k, The correlation between KIF18A protein expression and the genetic dependency on this gene in the Achilles-RNAi screen. Spearman’s ρ = 0.25 (P = 0.009). l, Relative mRNA expression levels of KIF18A, confirming successful siRNA-mediated KD in all cell lines 72h post-transfection. **P = 0.006, P = 0.003 and P = 0.002 for HCT116-GFP, HPT1 and HPT2, respectively; ***P = 0.0007; one-tailed t-test. m, Proliferation curves of HCT116 and HPT1 cells cultured in the presence of a KIF18A-targeting siRNA, or a non-targeting control siRNA. n, Comparison of the doubling times of HCT116 and HPT cells following siRNA-mediated KIF18A knockdown. **P = 0.001; two-tailed t-test. o, Time-lapse imaging-based quantification of the time from nuclear envelope breakdown (NEBD) to anaphase onset in HCT116 and HPT cell lines exposed to non-targeting or KIF18A-targeting siRNAs for 72h. n.s, P > 0.05; ** P = 0.003; two-tailed t-test. Bar, median; box, 25th and 75th percentile; whiskers, 1.5 X IQR; circles, individual cell lines. p, The prevalence of micronuclei formation in HCT116 and HPT cells exposed to non-targeting or KIF18A-targeting siRNAs for 72h. n.s., P > 0.05; ***P < 0.001; two-tailed Fisher’s exact test. q, Relative protein expression levels of KIF18A, confirming successful KIF18A overexpression in the highly-aneuploid HPT1 and HPT2 cell lines 48h post-transfection. Left: western blot of KIF18A protein expression levels in HPT1 and HPT2 before and after KIF18A overexpression. Right: quantification of KIF18A expression levels (normalized to α-Tubulin). *P = 0.013, **P = 0.005; one-tailed t-test; n = 2 biological replicates. In all bar plots and line plots, data represent the mean ± s.d. unless otherwise noted; n = 3 biological replicates unless otherwise noted. r, Proliferation curves of HPT1 cells before and after overexpression of KIF18A (KIF18A-OE), in the absence or presence of MPI-0479605 (250nM).
Extended Data Fig. 10 Additional validation of the increased sensitivity of aneuploid cells to KIF18A inhibition.
a, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen across multiple cell lineages. *P = 0.022; two-tailed t-test. b, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen, after accounting for lineage-specific differences in gene dependency scores using linear regression. *P = 0.012; two-tailed t-test. c, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen, across TP53 mutation classes. * P = 0.026; two-tailed t-test. d, The correlations between AS and the dependency on KIF18A in the DRIVE RNAi screen, for cell lines that have not undergone whole-genome duplication (that is, cell lines with basal ploidy of n = 2). Spearman’s ρ = -0.27 (P = 7e-04). e, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen, after removing the effect of doubling time on gene dependency scores using linear regression. *P = 0.022; two-tailed t-test. f, The sensitivity of near-euploid and highly-aneuploid cancer cell lines without microsatellite instability (MSS lines only) to the knockdown of KIF18A in the DRIVE RNAi screen. ***P = 3e-04; two-tailed t-test. g, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen, in cell lines that are WT for the 4 genes most selectively mutated in aneuploid human tumours (after TP53)12. *P = 0.021 and P = 0.02, for CTCF and ARID1A, respectively; **P = 0.004; two-tailed t-test. h, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen, after removing the effect of lineage subtype on gene dependency scores using linear regression. *P = 0.024; two-tailed t-test. i, The sensitivity of near-euploid and highly-aneuploid cancer cell lines to the knockdown of KIF18A in the DRIVE RNAi screen, after removing the effect of HET70 scores on gene dependency scores using linear regression. **P = 0.003; two-tailed t-test. j, Left: western blot of KIF18A protein expression levels in RPE1 and RPT cell lines. Right: Quantification of KIF18A expression levels (normalized to GAPDH). *P = 0.023; one-tailed t-test. Data represent the mean ± s.d.; n = 3 biological replicates. k, Relative protein expression levels of KIF18A, confirming successful KIF18A knockdown in the RPE1 and RPT cell lines 72h post-transfection. Left: western blot of KIF18A protein expression levels in RPE1, RPT1 and RPT3 before and after siRNA-mediated KIF18A knockdown. Right: Quantification of KIF18A expression levels (normalized to α-Tubulin). *P = 0.034, **P = 0.004; one-tailed t-test. Data represent the mean ± s.d.; n = 3 biological replicates. l, Time-lapse imaging-based quantification of the time from nuclear envelope breakdown (NEBD) to anaphase onset in RPE1 and RPT cell lines exposed to non-targeting or KIF18A-targeting siRNAs for 72h. ** P < 0.01; ****P < 1e-04; two-tailed t-test. m, The prevalence of micronuclei formation in HCT116 and HPT cells exposed to non-targeting or KIF18A-targeting siRNAs for 72h. n.s., P > 0.05; **P < 0.01; ***P < 0.001; two-tailed Fisher’s exact test. n, Imaging-based quantification of the prevalence of cell divisions with multipolar spindles in RPE1 and RPT cell lines treated with non-targeting control or KIF18A-targeting siRNAs for 72h. *P < 0.05; **P < 0.01; ***P < 0.001; two-tailed t-test; Error bars, s.d.; n = 3 biological replicates.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-15, Supplementary Figures 1-4 and Supplementary References.
Supplementary Table 1
Chromosome-arm copy number calls and aneuploidy scores for 997 human cancer cell lines. For each cell line, the copy number status of each chromosome arm was determined by comparing the weighted median log2 copy number (wmed_CN) value of the arm to the basal ploidy of the cell line. AS were determined as the number of chromosome arms that were gained or lost in each cell line.
Supplementary Table 2
Genetic dependencies of highly-aneuploid cancer cells. The lists include all genes on which aneuploid cancer cell lines were found to be more dependent than euploid cancer cell lines (effect size<-0.1, q<0.25) in the RNAi-Achilles and RNAi-DRIVE genetic dependency screens. P-values were derived from two-sided empirical-Bayes-moderated t-statistics. Q values were computed using the Benjamini–Hochberg method.
Supplementary Table 3
Functional annotation enrichment analysis of aneuploidy-associated genetic dependencies. DAVID functional enrichment analysis was performed on the genes that came up as differentially essential between the near-euploid and highly-aneuploid cell lines, focusing on the GO_BP gene sets. Gene sets with q<0.1 are listed. P-values were derived from the modified one-sided Fisher’s exact test applied by DAVID59. Q values were computed using the Benjamini–Hochberg method.
Supplementary Table 4
mRNA expression differences between near-euploid and highly-aneuploid cancer cells. A comparison of mRNA expression between the near-euploid and highly-aneuploid cancer cell lines, using the CCLE gene expression profiles2. P-values were derived from two-sided empirical-Bayes-moderated t-statistics. Q values were computed using the Benjamini–Hochberg method.
Supplementary Table 5
Chemical sensitivities of highly-aneuploid cancer cells. The lists present the differential dug sensitivities between near-euploid and highly-aneuploid cancer cell lines, for all of the compounds tested in the CTD2, GDSC and PRISM drug screens. P-values were derived from two-sided empirical-Bayes-moderated t-statistics. Q values were computed using the Benjamini–Hochberg method.
Supplementary Table 6
Cancer cell line sensitivity to the SAC inhibitor reversine. Results of a PRISM screen of reversine (at 8 doses) across 530 human cancer cell lines. Shown are the Area Under the ROC Curve (AUC) values.
Supplementary Table 7
Gene expression profiles of HCT116 and HPT cells exposed to SAC inhibitors. L1000-based expression values of 10,174 genes63 in HCT116-WT, HCT116-GFP, HPT1 and HPT2 cells treated with reversine (250nM or 500nM), MPI-0479605 (250nM), positive controls (reversine at 10µM and Mitoxantrone at 10µM) or negative controls (DMSO).
Supplementary Table 8
Association between aneuploidy levels and KIF18A mRNA and protein expression levels. Comparison of CCLE mRNA and protein expression values between the near-euploid and highly-aneuploid cell line groups, across multiple tissue types. P-values are derived from a two-tailed t-test.
Rights and permissions
About this article
Cite this article
Cohen-Sharir, Y., McFarland, J.M., Abdusamad, M. et al. Aneuploidy renders cancer cells vulnerable to mitotic checkpoint inhibition. Nature 590, 486–491 (2021). https://doi.org/10.1038/s41586-020-03114-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-03114-6
This article is cited by
-
Aneuploid serves as a prognostic marker and favors immunosuppressive microenvironment in ovarian cancer
Journal of Ovarian Research (2024)
-
Machine-learning analysis reveals an important role for negative selection in shaping cancer aneuploidy landscapes
Genome Biology (2024)
-
Therapeutic targeting of the KIF18A motor protein in cancers with chromosomal instability
Nature Cancer (2024)
-
CKAP5 stabilizes CENP-E at kinetochores by regulating microtubule-chromosome attachments
EMBO Reports (2024)
-
The two sides of chromosomal instability: drivers and brakes in cancer
Signal Transduction and Targeted Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.