Abstract
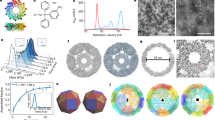
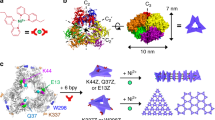
Many proteins exist naturally as symmetrical homooligomers or homopolymers1. The emergent structural and functional properties of such protein assemblies have inspired extensive efforts in biomolecular design2,3,4,5. As synthesized by ribosomes, proteins are inherently asymmetric. Thus, they must acquire multiple surface patches that selectively associate to generate the different symmetry elements needed to form higher-order architectures1,6—a daunting task for protein design. Here we address this problem using an inorganic chemical approach, whereby multiple modes of protein–protein interactions and symmetry are simultaneously achieved by selective, ‘one-pot’ coordination of soft and hard metal ions. We show that a monomeric protein (protomer) appropriately modified with biologically inspired hydroxamate groups and zinc-binding motifs assembles through concurrent Fe3+ and Zn2+ coordination into discrete dodecameric and hexameric cages. Our cages closely resemble natural polyhedral protein architectures7,8 and are, to our knowledge, unique among designed systems9,10,11,12,13 in that they possess tightly packed shells devoid of large apertures. At the same time, they can assemble and disassemble in response to diverse stimuli, owing to their heterobimetallic construction on minimal interprotein-bonding footprints. With stoichiometries ranging from [2 Fe:9 Zn:6 protomers] to [8 Fe:21 Zn:12 protomers], these protein cages represent some of the compositionally most complex protein assemblies—or inorganic coordination complexes—obtained by design.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The principal data supporting the findings of this work are available within the figures and the Supplementary Information. Additional data that support the findings of this study are available from the corresponding author on request. Structural data obtained by X-ray crystallography and cryo-EM have been deposited into the RCSB PDB and EMDB data banks with the following accession codes: 6OT4 (BMC2), 6OT7 (BMC3), 6OT8 (BMC4), 6OT9 (BMC1) and 6OVH (BMC3 cryo-EM) in the PDB or EMD-20212 at The Electron Microscopy Data Bank.
References
Marsh, J. A. & Teichmann, S. A. Structure, dynamics, assembly, and evolution of protein complexes. Annu. Rev. Biochem. 84, 551–575 (2015).
Padilla, J. E., Colovos, C. & Yeates, T. O. Nanohedra: using symmetry to design self assembling protein cages, layers, crystals, and filaments. Proc. Natl Acad. Sci. USA 98, 2217–2221 (2001).
Bai, Y., Luo, Q. & Liu, J. Protein self-assembly via supramolecular strategies. Chem. Soc. Rev. 45, 2756–2767 (2016).
Hamley, I. W. Protein assemblies: nature-inspired and designed nanostructures. Biomacromolecules 20, 1829–1848 (2019).
Churchfield, L. A. & Tezcan, F. A. Design and construction of functional supramolecular metalloprotein assemblies. Acc. Chem. Res. 52, 345–355 (2019).
Yeates, T. O. Geometric principles for designing highly symmetric self-assembling protein nanomaterials. Annu. Rev. Biophys. 46, 23–42 (2017).
Johnson, J. E. & Speir, J. A. Quasi-equivalent viruses: a paradigm for protein assemblies. J. Mol. Biol. 269, 665–675 (1997).
Lawson, D. M. et al. Solving the structure of human H ferritin by genetically engineering intermolecular crystal contacts. Nature 349, 541–544 (1991).
Lai, Y.-T., Cascio, D. & Yeates, T. O. Structure of a 16-nm cage designed by using protein oligomers. Science 336, 1129 (2012).
Bale, J. B. et al. Accurate design of megadalton-scale two-component icosahedral protein complexes. Science 353, 389–394 (2016).
Cristie-David, A. S. & Marsh, E. N. G. Metal-dependent assembly of a protein nano-cage. Protein Sci. 28, 1620–1629 (2019).
Fletcher, J. M. et al. Self-assembling cages from coiled-coil peptide modules. Science 340, 595–599 (2013).
Malay, A. D. et al. An ultra-stable gold-coordinated protein cage displaying reversible assembly. Nature 569, 438–442 (2019).
Pluth, M. D., Bergman, R. G. & Raymond, K. N. Acid catalysis in basic solution: a supramolecular host promotes orthoformate hydrolysis. Science 316, 85–88 (2007).
Chakrabarty, R., Mukherjee, P. S. & Stang, P. J. Supramolecular coordination: self-assembly of finite two- and three-dimensional ensembles. Chem. Rev. 111, 6810–6918 (2011).
Yoshizawa, M., Klosterman, J. K. & Fujita, M. Functional molecular flasks: new properties and reactions within discrete, self-assembled hosts. Angew. Chem. Int. Ed. 48, 3418–3438 (2009).
Mal, P., Breiner, B., Rissanen, K. & Nitschke, J. R. White phosphorus is air-stable within a self-assembled tetrahedral capsule. Science 324, 1697–1699 (2009).
Liu, Y., Hu, C., Comotti, A. & Ward, M. D. Supramolecular Archimedean cages assembled with 72 hydrogen bonds. Science 333, 436–440 (2011).
Mateu, M. G. Assembly, stability and dynamics of virus capsids. Arch. Biochem. Biophys. 531, 65–79 (2013).
Sun, X., Johnson, D. W., Caulder, D. L., Raymond, K. N. & Wong, E. H. Rational design and assembly of M2M′3L6 supramolecular clusters with C3h symmetry by exploiting incommensurate symmetry numbers. J. Am. Chem. Soc. 123, 2752–2763 (2001).
Smulders, M. M. J., Jiménez, A. & Nitschke, J. R. Integrative self-sorting synthesis of a Fe8Pt6L24 cubic cage. Angew. Chem. Int. Ed. 51, 6681–6685 (2012).
Brodin, J. D. et al. Metal-directed, chemically tunable assembly of one-, two- and three-dimensional crystalline protein arrays. Nat. Chem. 4, 375–382 (2012).
Suzuki, Y. et al. Self-assembly of coherently dynamic, auxetic, two-dimensional protein crystals. Nature 533, 369–373 (2016).
Radford, R. J., Nguyen, P. C. & Tezcan, F. A. Modular and versatile hybrid coordination motifs on α-helical protein surfaces. Inorg. Chem. 49, 7106–7115 (2010).
Pearson, R. G. Hard and soft acids and bases. J. Am. Chem. Soc. 85, 3533–3539 (1963).
Wong, G. B., Kappel, M. J., Raymond, K. N., Matzanke, B. & Winkelmann, G. Coordination chemistry of microbial iron transport compounds. 24. Characterization of coprogen and ferricrocin, two ferric hydroxamate siderophores. J. Am. Chem. Soc. 105, 810–815 (1983).
Crumbliss, A. L. Iron bioavailability and the coordination chemistry of hydroxamic acids. Coord. Chem. Rev. 105, 155–179 (1990).
Ni, T. W. & Tezcan, F. A. Structural characterization of a microperoxidase inside a metal-directed protein cage. Angew. Chem. Int. Ed. 49, 7014–7018 (2010).
Failes, T. W. & Hambley, T. W. Crystal structures of tris(hydroxamato) complexes of iron(III). Aust. J. Chem. 53, 879–881 (2000).
Mayhew, S. G. The redox potential of dithionite and SO−2 from equilibrium reactions with flavodoxins, methyl viologen and hydrogen plus hydrogenase. Eur. J. Biochem. 85, 535–547 (1978).
Borsook, H. & Keighley, G. Oxidation-reduction potential of ascorbic acid (vitamin C). Proc. Natl Acad. Sci. USA 19, 875–878 (1933).
Michalak, K., Wicha, J. & Wójcik, J. Studies towards dynamic kinetic resolution of 4-hydroxy-2-methylcyclopent-2-en-1-one and its E-O-trityloxime. Tetrahedron 72, 4813–4820 (2016).
Liu, H. & Naismith, J. H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 8, 91 (2008).
Faraone-Mennella, J., Tezcan, F. A., Gray, H. B. & Winkler, J. R. Stability and folding kinetics of structurally characterized cytochrome c-b562. Biochemistry 45, 10504–10511 (2006).
Bailey, J. B., Subramanian, R. H., Churchfield, L. A. & Tezcan, F. A. in Methods in Enzymology Vol. 580 (ed. Pecoraro, V. L.) 223–250 (Academic, 2016).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. D 66, 133–144 (2010).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Terwilliger, T. C. et al. Phenix—a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Schrodinger, LLC. The PyMOL Molecular Graphics System. version 1.3 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Lindner, H. J. & Gottlicher, S. Die Kristall- und Molekulstruktur des Eisen(III)-benzhydroxamat-Trihydrates. Acta Crystallogr. B 25, 832–842 (1969).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Schuck, P. A model for sedimentation in inhomogeneous media. I. Dynamic density gradients from sedimenting co-solutes. Biophys. Chem. 108, 187–200 (2004).
Kleywegt, G. J. & Jones, T. A. Detection, delineation, measurement and display of cavities in macromolecular structures. Acta Crystallogr. D 50, 178–185 (1994).
Mazurenko, S. et al. CalFitter: a web server for analysis of protein thermal denaturation data. Nucleic Acids Res. 46, W344–W349 (2018).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Acknowledgements
This work was supported by the US Department of Energy (Division of Materials Sciences, Office of Basic Energy Sciences; DE-SC0003844; for the design strategy, EM imaging and analysis, and biochemical analysis) and by the National Science Foundation (Division of Materials Research; DMR-1602537; for crystallographic analysis). E.G. acknowledges support by an EMBO Long-Term Postdoctoral Fellowship (ALTF 1336-2015). J.E. acknowledges support by a DFG Research Fellowship (DFG 393131496). R.H.S. was supported by the National Institute of Health Chemical Biology Interfaces Training Grant UC San Diego (T32GM112584). We acknowledge the use of the UCSD Cryo-EM Facility, which is supported by NIH grants to T.S.B. and a gift from the Agouron Institute to UCSD. Crystallographic data were collected either at Stanford Synchrotron Radiation Lightsource (SSRL) or at the Lawrence Berkeley Natural Laboratory on behalf of the Department of Energy.
Author information
Authors and Affiliations
Contributions
E.G. conceived the project, and designed and performed most experiments. R.H.S. and R.G.A. performed and processed the ns-TEM data and performed structural modelling and analysis. R.H.S. conducted encapsulation experiments. J.E. performed crystallographic analysis. J.B.B. and J.A.C. synthesized the IHA ligand. X.Y., T.B. and T.S.B. performed the cryo-EM data collection and processing. F.A.T. conceived and directed the project and wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Jack Johnson, Todd Yeates and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of the IHA ligand and the BMC constructs.
a, b, NMR spectra of N-hydroxy-2-iodoacetamide in DMSO-d6 for 1H (a) and 13C (b). c–f, ESI-MS of as-isolated and HA-functionalized BMC constructs, and AUC profiles of HA-functionalized protomers for BMC1 (c), BMC2 (d), BMC3 (e) and BMC4 (f). The calculated masses for each unlabelled protein are determined by summing the mass of the polypeptide sequence and the c-type haem (618 Da) covalently linked to the cytochrome.
Extended Data Fig. 2 Structural comparison of CFMC1 and BMC1 cages.
a, The symmetric substructures of the CFMC1 dodecameric unit and its per-protomer SASA and BSA values. Associative surfaces on the protomers are coloured red for homologous interactions and red/orange or blue/cyan for heterologous interactions (right). b, Summary of engineered metal-coordination motifs for BMC constructs (see Supplementary Table 1 for all mutations). c, d, Comparison of C2 and C3 symmetric interfaces and corresponding metal binding sites for CFMC1 (c) and BMC1 (d). Full cages are shown as surfaces; insets show details of each interface. Fe and Zn ions are represented as orange and teal spheres, respectively. e, Cartoon representation of a full-size BMC1 cage with all metal ions shown as spheres. PDB ID: 3M4B (CFMC1), 6OT9 (BMC1).
Extended Data Fig. 3 ns-TEM characterization of BMC constructs.
a, b, Dissolved Fe:Zn:BMC1 (a) and Fe:Zn:BMC2 crystals (b) in a buffer containing 100 mM HEPES (pH 7.5), 200 mM MgCl2 and 800 μM ZnCl2. c, Self-assembled Fe:Zn:BMC3 cages in a buffer containing 20 mM Tris (pH 8.5), 20 μM FeSO4 and 60 μM ZnCl2. Histograms in b, c reflect the size distributions of Fe:Zn:BMC2 and Fe:Zn:BMC3 cage diameters as measured from ns-TEM images. Gaussian fits to both distributions are drawn as solid lines along with their centres and standard deviations reported. Scale bars, 50 nm.
Extended Data Fig. 4 Cavity volumes of BMC cages.
Solvent-accessible cavity volumes within BMC cages as calculated by a 1.4 Å rolling probe are shown visually as blue meshes and reported numerically below. Spherical cavities, shown as yellow spheres in Figs. 2, 4, are reproduced for comparison to the calculated volumes. BMC proteins are represented as transparent cylinders.
Extended Data Fig. 5 Anomalous densities of engineered metal binding sites and conformational flexibility of Cys82–HA site.
a–j, Cartoon and stick representations of the symmetric interfaces of BMC1 (a, b), BMC2 (c–e), BMC3 (f–h) and BMC4 (i, j) showing the engineered metal binding sites with the C63–HA ligands (a, c, f), C82–HA ligands (d, g, i) and Zn binding sites (b, e, h, j). The difference in the anomalous signal between pairs of datasets above and below the K-shell energy of Zn and Fe, respectively, are depicted as blue or orange meshes. A strong signal illustrates a strong change in anomalous signal across the respective edge, in turn suggesting the presence of the respective metal. The top right corner of each panel indicates the energies of the datasets used for the map of the respective colour. All anomalous difference maps are contoured at 3σ. As datasets around the Fe-edge were not available for BMC1 and BMC3 (necessitating calculations using anomalous difference density of singular datasets), the calculated f″ values for Zn at 7.3 and 9.3 keV are 0.82 and 0.52 (that is, non-zero) and thus some residual anomalous signal of the lower energy maps around the Zn atoms is expected to result even from strictly selective Zn loading. For a more quantitative analysis of the nature of the bound metal, ratios of the anomalous signal to the expected values (bottom left corner of each panel) were calculated as described in the Methods. k, Stick representation of the BMC2 Cys82–HA binding site in both alternative conformations with the anomalous difference density over the Fe-edge shown as orange mesh and a simulated annealing omit map (omitting all C82–HA atoms and Fe) of the normal electron density as light blue mesh contoured at 2σ. For all Cys–HA binding sites, arrows indicate the handedness of the binding site as Δ (right handed) or Λ (left handed). The reversion of handedness in k with the respective view angle is indicated by arrows. Colour code for atoms in all panels: Fe in orange, Zn in blue, S in yellow, O in red and N in dark blue.
Extended Data Fig. 6 Solution characterization of self-assembled BMC3 and BMC4 cages.
a–c, The oligomerization state of BMC3 cages as monitored by AUC measurements following incubation with various first-row transition metal ions (a), incubation with Zn2+ and Fe3+ (Fe(acetylacetonate)3) (b) and disassembly via sequestration of metal ions by EDTA (c). d, AUC profiles of BMC variants after equilibration for 2 h at the indicated temperatures (top). Thermal unfolding of BMC variants as measured by circular dichroism spectroscopy at 222 nm (bottom). e, AUC profiles of BMC following treatment with chemical reductants of different reduction potentials (left). ns-TEM micrographs (middle and right) are shown for cage samples incubated with the corresponding chemical reductants.
Extended Data Fig. 7 Cryo-EM analysis of BMC3 cages.
a, Representative cryo-EM micrograph and 2D class averages. b, Flowchart detailing image processing from collected movie stacks to final map. Additional details can be found in the Methods. c, FSC curves calculated between the half-maps (black line), atomic model to the unmasked full map (purple line) and atomic model to the masked full map (blue line). Resolution values are indicated at the gold-standard FSC 0.143 criterion. d, Local resolution estimates of the final reconstruction calculated using ResMap. e, Electron density shown at BMC3 C3 interfaces highlighting poorly resolved density (reflecting high flexibility) at hydroxamate sites and multiple conformations of W66.
Extended Data Fig. 8 Encapsulation of rhodamine inside BMC3 cages.
a, Fluorescence characterization of BMC3 samples incubated with rhodamine. Cages encapsulating rhodamine were treated with EDTA and washed before measuring fluorescence intensity. b, AUC profiles of cages encapsulating rhodamine monitored at the haem Soret absorption maximum (λmax = 415 nm) and rhodamine absorption maximum (λmax = 555 nm). c, UV-vis characterization of BMC3 samples incubated with rhodamine. d, Difference spectra of BMC3 samples and BMC3 protomer shown in c. Free rhodamine dissolved in buffer is shown as dark-red dashes. e, Repeated fluorescence characterization of a solution containing BMC3 cages encapsulating rhodamine over several days. The sample was washed three times before each fluorescence measurement.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Supplementary Tables 1-6. Supplementary Tables include the sequences, crystallization conditions and crystallographic quantification of metal content for the different proteins employed in this research.
Rights and permissions
About this article
Cite this article
Golub, E., Subramanian, R.H., Esselborn, J. et al. Constructing protein polyhedra via orthogonal chemical interactions. Nature 578, 172–176 (2020). https://doi.org/10.1038/s41586-019-1928-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1928-2
This article is cited by
-
DNA-origami-directed virus capsid polymorphism
Nature Nanotechnology (2023)
-
Supramolecular assembling systems of hemoproteins using chemical modifications
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2023)
-
Constructing artificial mimic-enzyme catalysts for carbon dioxide electroreduction
Science China Chemistry (2022)
-
Design of metal-mediated protein assemblies via hydroxamic acid functionalities
Nature Protocols (2021)
-
De novo metalloprotein design
Nature Reviews Chemistry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.