Abstract
Precise protein sequencing and folding are believed to generate the structure and chemical diversity of natural channels1,2, both of which are essential to synthetically achieve proton transport performance comparable to that seen in natural systems. Geometrically defined channels have been fabricated using peptides, DNAs, carbon nanotubes, sequence-defined polymers and organic frameworks3,4,5,6,7,8,9,10,11,12,13. However, none of these channels rivals the performance observed in their natural counterparts. Here we show that without forming an atomically structured channel, four-monomer-based random heteropolymers (RHPs)14 can mimic membrane proteins and exhibit selective proton transport across lipid bilayers at a rate similar to those of natural proton channels. Statistical control over the monomer distribution in an RHP leads to segmental heterogeneity in hydrophobicity, which facilitates the insertion of single RHPs into the lipid bilayers. It also results in bilayer-spanning segments containing polar monomers that promote the formation of hydrogen-bonded chains15,16 for proton transport. Our study demonstrates the importance of the adaptability that is enabled by statistical similarity among RHP chains and of the modularity provided by the chemical diversity of monomers, to achieve uniform behaviour in heterogeneous systems. Our results also validate statistical randomness as an unexplored approach to realize protein-like behaviour at the single-polymer-chain level in a predictable manner.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study, as well as descriptions of the methodologies used in the code, are available within the article and the Extended Data items. For reproduction purposes, the raw data used to generate the figures and input scripts used to produce the simulations are available from the Dryad Digital Repository (https://doi.org/10.6078/D1VX0B).
Code availability
All custom scripts (sequence analyses and HMM) are available from the corresponding author upon request.
References
Stouffer, A. L. et al. Structural basis for the function and inhibition of an influenza virus proton channel. Nature 451, 596–599 (2008); corrigendum 452, 380 (2008).
Schnell, J. R. & Chou, J. J. Structure and mechanism of the M2 proton channel of influenza A virus. Nature 451, 591–595 (2008).
Ghadiri, M. R., Granja, J. R. & Buehler, L. K. Artificial transmembrane ion channels from self-assembling peptide nanotubes. Nature 369, 301–304 (1994).
Xu, T. et al. Subnanometer porous thinfilms by the co-assembly of nanotube subunits and block copolymers. ACS Nano 5, 1376–1384 (2011).
Joh, N. H. et al. De novo design of a transmembrane Zn2+-transporting four-helix bundle. Science 346, 1520–1524 (2014).
Langecker, M. et al. Synthetic lipid membrane channels formed by designed DNA nanostructures. Science 338, 932–936 (2012).
Hinds, B. J. et al. Aligned multiwalled carbon nanotube membranes. Science 303, 62–65 (2004).
Geng, J. et al. Stochastic transport through carbon nanotubes in lipid bilayers and live cell membranes. Nature 514, 612–615 (2014).
Percec, V. et al. Self-assembly of amphiphilic dendritic dipeptides into helical pores. Nature 430, 764–768 (2004).
Weiss, L. A., Sakai, N., Ghebremariam, B., Ni, C. Y. & Matile, S. Rigid rod-shaped polyols: functional nonpeptide models for transmembrane proton channels. J. Am. Chem. Soc. 119, 12142–12149 (1997).
Vial, F., Oukhaled, A. G., Auvray, L. & Tribet, C. Long-living channels of well defined radius opened in lipid bilayers by polydisperse, hydrophobically-modified polyacrylic acids. Soft Matter 3, 75–78 (2007).
Jung, M., Kim, H., Baek, K. & Kim, K. Synthetic ion channel based on metal–organic polyhedra. Angew. Chem. 47, 5755–5757 (2008).
Si, W. et al. Selective artificial transmembrane channels for protons by formation of water wires. Angew. Chem. 50, 12564–12568 (2011).
Panganiban, B. et al. Random heteropolymers preserve protein function in foreign environments. Science 359, 1239–1243 (2018).
Nagle, J. F. & Morowitz, H. J. Molecular mechanisms for proton transport in membranes. Proc. Natl Acad. Sci. USA 75, 298–302 (1978).
Decoursey, T. E. Voltage-gated proton channels and other proton transfer pathways. Physiol. Rev. 83, 475–579 (2003).
Smith, A. A. A., Hall, A., Wu, V. & Xu, T. Practical prediction of heteropolymer composition and drift. ACS Macro Lett. 8, 36–40 (2019).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Hemmatian, Z. et al. Electronic control of H+ current in a bioprotonic device with Gramicidin A and Alamethicin. Nat Commun. 7, 12981 (2016)
Lin, T. I. & Schroeder, C. Definitive assignment of proton selectivity and attoampere unitary current to the M2 ion channel protein of influenza A virus. J. Virol. 75, 3647–3656 (2001).
Krishnamoorthy, G. Temperature jump as a new technique to study the kinetics of fast transport of protons across membranes. Biochemistry 25, 6666–6671 (1986).
Tunuguntla, R. H., Allen, F. I., Kim, K., Belliveau, A. & Noy, A. Ultrafast proton transport in sub-1-nm diameter carbon nanotube porins. Nat. Nanotechnol. 11, 639–644 (2016).
DeCoursey, T. E. & Cherny, V. V. Deuterium isotope effects on permeation and gating of proton channels in rat alveolar epithelium. J. Gen. Physiol. 109, 415–434 (1997).
Kučerka, N., Tristram-Nagle, S. & Nagle, J. F. Structure of fully hydrated fluid phase lipid bilayers with monounsaturated chains. J. Membr. Biol. 208, 193–202 (2006).
Golumbfskie, A. J., Pande, V. S. & Chakraborty, A. K. Simulation of biomimetic recognition between polymers and surfaces. Proc. Natl Acad. Sci. USA 96, 11707–11712 (1999).
Mowery, B. P. et al. Mimicry of antimicrobial host-defense peptides by random copolymers. J. Am. Chem. Soc. 129, 15474–15476 (2007).
Tribet, C., Audebert, R. & Popot, J. L. Amphipols: polymers that keep membrane proteins soluble in aqueous solutions. Proc. Natl Acad. Sci. USA 93, 15047–15050 (1996).
Stewart, J. C. M. Colorimetric determination of phospholipids with ammonium ferrothiocyanate. Anal. Biochem. 104, 10–14 (1980).
Lira, R. B., Steinkuhler, J., Knorr, R. L., Dimova, R. & Riske, K. A. Posing for a picture: vesicle immobilization in agarose gel. Sci. Rep. 6, 25254 (2016).
Manneville, J. B. et al. COPI coat assembly occurs on liquid-disordered domains and the associated membrane deformations are limited by membrane tension. Proc. Natl Acad. Sci. USA 105, 16946–16951 (2008).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Miyamoto, S. & Kollman, P. A. Settle: an analytical version of the shake and rattle algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
Martínez, L., Andrade, R., Birgin, E. G. & Martinez, J. M. PACKMOL: a package for building initial configurations for molecular dynamics simulations. J. Comput. Chem. 30, 2157–2164 (2009).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Darden, T., York, D. & Pedersen, L. Particle mesh ewald: an N∙log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. Model. 14, 33–38 (1996).
Zeidel, M. L., Ambudkar, S. V., Smith, B. L. & Agre, P. Reconstitution of functional water channels in liposomes containing purified red-cell CHIP28 protein. Biochemistry 31, 7436–7440 (1992).
Shen, Y. X. et al. Highly permeable artificial water channels that can self-assemble into two-dimensional arrays. Proc. Natl Acad. Sci. USA 112, 9810–9815 (2015).
Heller, W. T. et al. The suite of small-angle neutron scattering instruments at Oak Ridge National Laboratory. J. Appl. Cryst. 51, 242–248 (2018).
Rabiner, L. R. A tutorial on hidden Markov models and selected applications in speech recognition. Proc. IEEE 77, 257–286 (1989).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. A 39, 1–22 (1977).
Forney, C. D. The Viterbi algorithm. Proc. IEEE 61, 268–278 (1973).
Acknowledgements
This work was supported by the US Department of Defense, Army Research Office, under contract W911NF-13-1-0232 and the National Science Foundation under contract DMR-183696. M.O.d.l.C. and B.Q. acknowledge support through grant DE-FG02-08ER46539 from the Department of Energy (DOE) Basic Energy Science Office and the Center for Computation and Theory of Soft Materials, as well as computational support by the Sherman Fairchild Foundation. Z. H. and M. R. acknowledge support from the Air Force Office of Sponsored Research Award FA9550-15-1-0273. A part of this research used resources at the High Flux Isotope Laboratory, a DOE Office of Science User Facility operated by the Oak Ridge National Laboratory. We acknowledge the support of the National Institute of Standards and Technology, US Department of Commerce, in providing the neutron research facilities used in this work. We thank L. He and Y. Liu for help in the SANS studies. Scattering studies at the Advanced Light Source and RHP characterization at the Molecular Foundry were supported by the Office of Science, Office of Basic Energy Sciences of the US DOE under contract DE-AC02-05CH11231. We thank A. A. A. Smith for help in polymer syntheses; Y. W. Qian for characterizing the liposome sizes; A. Martin and E. M. López-Alfonzo for help in stopped-flow experiments. We thank CoC-NMR for help with RHP characterization; instruments at CoC-NMR are supported in part by NIH S10OD024998.
Author information
Authors and Affiliations
Contributions
T.J. made the first experimental observation of RHP-based proton transport. T.X. directed the project development. T.J. and T.X. designed the experiments and data analysis. T.J. performed imaging, FRAP and transport studies on liposomes. A.H. synthesized and characterized RHPs. M.E. and Z.R. performed the RHP segmental analysis. Z.H. and M.R. performed transport studies on SLBs. B.Q. and M.O.d.l.C. performed all-atom simulations and data analysis. Y.Z. and H.H performed HMM studies. T.J. and A.D.C. prepared and characterized liposomes. T.J. and W.T.H. collected and analysed SANS data with T.X. All authors participated in the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Junli Hou, Kenichi Kuroda and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Interactions between RHPs and liposomes.
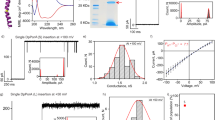
a, DSC profiles of LUVs and RHP1- and RHP1,100-containing LUVs (POPE:POPG molar ratio, 3:1). LUVs were grown with RHP1 or RHP1,100 with a lipid-to-RHP molar ratio of 50:1 and a lipid concentration of 1 mg mL−1. The background signals of the buffer and the polymers were subtracted. No phase transition was observed for RHP itself. The plots are shifted vertically for clarity by +0.0001 for RHP1,100 and +0.0002 for RHP1. b, Full-width at half-maximum of the lipid phase transition peaks shown in a. Error bars are 1 s.d. (n = 3). c, d, Representative confocal fluorescence image of POPC GUVs incubated with Texas Red-labelled RHP1 (c) and Texas Red-labelled RHP1,100 (d).
Extended Data Fig. 2 All-atom molecular dynamics simulations.
a, Sequences of the nine RHP1 chains used in the simulations. b, Snapshot of the 2nd–5th RHP1 chains in the POPC lipid bilayer in the simulations. MMA, EHMA, OEGMA and SPMA are coloured red, pink, blue and purple, respectively. Lipids are shown in grey colour. c, Representative HBCs in the transmembrane regions of RHP1s. The oxygen and hydrogen atoms of water molecules are coloured red and white, respectively. Hydrogen bonds are shown with red dashed lines. Lipids are omitted for clarity. d, Snapshots of the initial (left) and final (right) RHP1 conformations in the simulations. The RHP1 chain is highlighted, along with K+ (orange) and Cl− (green) ions. e, Convergence of the simulations. Top, area per lipid as a function of the simulation time. Bottom, Coulomb and Lennard–Jones interactions between POPC lipids and the 1st–5th RHP1s as a function of simulation time. Different colours denote the results from five parallel molecular dynamics simulations.
Extended Data Fig. 3 Transmembrane proton transport.
a, pH gradient-driven proton flux through RHP1 in the LUVs of POPE/POPG (molar ratio 3:1). b, Inner pH changes in POPC LUVs incubated with RHP1 and RHP variants over 200 s after adding Vln. ‘RHPwoM’, ‘RHPwoSP’ and ‘RHP7/0’ represent the RHP without MMA, the RHP without SPMA and the RHP without EHMA, respectively. c, pH gradient-driven proton flux in the influx and outflux directions. RHP1 was added to the solutions of preformed LUVs. d, pH gradient-driven proton flux through RHP1 in DMPC SUVs at 37 °C. RHP1-containing SUVs were prepared as indicated in the Methods section ‘SANS investigation’ using H2O-based buffers, with an RHP1:DMPC ratio of 0.001.
Extended Data Fig. 4 Proton and water transport rates.
a, Representative stopped-flow fluorescence traces of RHP1-containing POPC LUVs after rapid exposure to a buffer of a higher pH value. The initial and final equilibrated pH values are 6.85 and 7.35, respectively. Vln was added before the measurements. b, Calculated permeability values of RHP1-containing LUVs at various RHP1-to-lipid ratios. Error bars are 1 s.d. (n = 3). c, d, Conductance (c) and initial proton transport rate (d) per RHP1 chain at various RHP1-to-lipid ratios. Error bars are 1 s.d. (n = 3). e, Stopped-flow fluorescence traces of RHP1-containing LUVs after rapid exposure to a buffer of a higher pH or pD value. The initial and final equilibrated pH values are 6.85 and 7.35, respectively, for the H2O-based assay. The initial and final pD values are 7.25 and 7.75, respectively, for the D2O-based assay. Vln was added to the RHP1-containing LUV solutions, except for the GramA-containing LUVs. f, Osmotic water permeability of Vln, GramA and RHP1+Vln in POPC LUVs at 7 °C. Vln (K+ channel) and GramA (proton/cations/water channel) are the negative and positive controls, respectively. GramA, Vln and RHP1 were added to the solutions of preformed LUVs. ‘Premixed’ denotes premixing of RHP1 and the lipid before the preparation of LUVs. Error bars are 1 s.d. (n = 7).
Extended Data Fig. 5 SANS investigations.
a, SANS profiles of free RHP1 in 86% D2O. The solid line denotes a fit using a polydisperse sphere model for RHP1. The signal of the solvent has been subtracted in the presented curves. b, Schematic illustration of the three-shell vesicle model for the SUV, showing the regions of the liposome core (core), inner bilayer headgroup (shell 1), lipid bilayer core (shell 2) and outer bilayer headgroup (shell 3). c, Structural parameters obtained by fitting the SANS data with the three-shell vesicle model for the liposome and the polydisperse sphere model for RHP1. Parameters that were fixed during the fitting are denoted with ‘f’; RMS represents root mean square. SLDs that were fixed were determined from the known sample composition, including when RHP1 was present in the bilayer, because the concentration was too low to impact the SLD value of shells significantly. The SLD values used for the liposome core, for shells 1, 2, 3 and for the solvent are 5.41, 1.88, 6.84, 1.88 and 5.41 × 10−6 Å−2, respectively. The SLD value for RHP1 is 1.4 × 10−6 Å−2 in the sphere model.
Extended Data Fig. 6 Sequence analyses of RHPs.
a, Statistical distribution of number of OEGMA monomers per RHP1 chains in the 4,500 simulated RHP1 sequences (DP = 130). This distribution reflects the compositional dispersity in RHP1 chains. b, Statistical size distribution of hydrophobic segments with one OEGMA in RHP1s of various chain lengths ranging from DP = 40 to 200. The number of chains (4,500) was kept constant for each DP analysis. c, Normalized profiles of the size distribution in b. d, Number of hydrophobic segments per chain at various chain lengths. The counted hydrophobic segments here contain one OEGMA with segment size ≥11. e, Statistical size distributions of blocks of EHMA (pink) and MMA (red) within hydrophobic segments containing one OEGMA in RHP variants. Results are shown for hydrophobic segments with segment size ≥11. f, Schematic illustration of five EHMA monomers distributed at the end of an inserted RHP1 chain in a molecular dynamics simulation. EHMA, MMA and OEGMA are coloured pink, red and blue, respectively. Lipids are omitted for clarity.
Extended Data Fig. 7 HMM prediction.
a, Comparison between state-3 segments (size ≥7; top) assigned using the method described in Methods section ‘Hydrophobic segments’ and those predicted by HMM (bottom). ‘1’, ‘2’, ‘3’ and ‘4’ denote MMA, OEGMA, EHMA and SPMA, respectively. b, Transition diagram with a state-3 segment of size ≥7. States 1 and 2 are denoted as S1 and S2, respectively. State 3 here is broken into 10 states to comply with the rules outlined in Methods section ‘HMM prediction’. S5 and S9 are the special states in a state-3 segment that must emit OEGMA with probability 1 (denoted as ‘prob. = 1’). c, Statistical size distribution of the state-3 segments predicted by HMM in 4,500 RHP1 sequences. d, Number of the predicted state-3 segments per polymer chain. e, Statistical size distribution of predicted state-3 segments containing one OEGMA. f, Number of predicted state-3 segments containing one OEGMA per polymer chain. 4,375 out of 4,500 RHP1 chains contain at least one state-3 segment.
Extended Data Fig. 8 Proton transport on SLBs.
a, Liquid atomic force microscopy images of a Pd surface covered with SLBs, showing a surface roughness of about 0.74 nm. b, Image for SLB+RHP1 (RHP1-to-lipid molar ratio, 0.001), showing a roughness of about 2.07 nm. c, Surface covered with SLB+RHP1 (RHP1-to-lipid molar ratio, 0.01), showing a roughness of about 5.61 nm. d, Current–voltage sweep of the protonic device. The Pd/SLB contact has a current of <0.01 nA across the entire voltage range, confirming the high polarization resistance of the SLB (black trace). With RHP1 incorporated, an oxidation peak is observed at about 48 mV, indicating H+ flow across the SLB; a reduction peak is observed at about −45 mV, indicating conduction of H across the SLB.
Supplementary information
Video 1
All-atom MD simulation trajectory showing the dynamic feature of a RHP1 chains in the POPC lipid bilayer. The RHP1 sequence is 1111411421123112111121213111132332323322, where 1, 2, 3, and 4 denote MMA, OEGMA, EHMA, and SPMA, respectively. Lipids are colored grey, with the phosphorous atoms in green. Hydrogen atoms and all other molecules are omitted for clarity. The frequency is 100 ps per frame for a total duration of 100 ns.
Video 2
All-atom MD simulation trajectory showing the dynamic feature of a RHP1 chains in the POPC lipid bilayer. The RHP1 sequence is 3141221233112332221312121331111111112114, where 1, 2, 3, and 4 denote MMA, OEGMA, EHMA, and SPMA, respectively. Lipids are colored grey, with the phosphorous atoms in green. Hydrogen atoms and all other molecules are omitted for clarity. The frequency is 100 ps per frame for a total duration of 100 ns.
Video 3
All-atom MD simulation trajectory demonstrating the dynamic hydrogen-bonded chains (HBCs) between RHP1 and water molecules. The RHP1 sequence is 1111411421123112111121213111132332323322, where 1, 2, 3, and 4 denote MMA, OEGMA, EHMA, and SPMA, respectively. Hydrogen bonds are plotted using blue dashed lines. The phosphorus atoms of the POPC lipid are shown at green beads. Water molecules are shown as red (O) + white (H) beads. The frequency is 10 ps per frame for a total duration of 1 ns.
Video 4
All-atom MD simulation trajectory demonstrating the dynamic hydrogen-bonded chains (HBCs) between RHP1 and water molecules. The RHP1 sequence is 3141221233112332221312121331111111112114, where 1, 2, 3, and 4 denote MMA, OEGMA, EHMA, and SPMA, respectively. Hydrogen bonds are plotted using blue dashed lines. The phosphorus atoms of the POPC lipid are shown at green beads. Water molecules are shown as red (O) + white (H) beads. The frequency is 10 ps per frame for a total duration of 1 ns.
Rights and permissions
About this article
Cite this article
Jiang, T., Hall, A., Eres, M. et al. Single-chain heteropolymers transport protons selectively and rapidly. Nature 577, 216–220 (2020). https://doi.org/10.1038/s41586-019-1881-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1881-0
This article is cited by
-
In vivo therapy of osteosarcoma using anion transporters-based supramolecular drugs
Journal of Nanobiotechnology (2024)
-
DNA nanopores as artificial membrane channels for bioprotonics
Nature Communications (2023)
-
Transient water wires mediate selective proton transport in designed channel proteins
Nature Chemistry (2023)
-
A high-throughput platform for efficient exploration of functional polypeptide chemical space
Nature Synthesis (2023)
-
Polymeric protagonists for biological processes
Nature Chemistry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.