Abstract
Respiratory syncytial virus (RSV) and human metapneumovirus (HMPV) cause severe respiratory diseases in infants and elderly adults1. No vaccine or effective antiviral therapy currently exists to control RSV or HMPV infections. During viral genome replication and transcription, the tetrameric phosphoprotein P serves as a crucial adaptor between the ribonucleoprotein template and the L protein, which has RNA-dependent RNA polymerase (RdRp), GDP polyribonucleotidyltransferase and cap-specific methyltransferase activities2,3. How P interacts with L and mediates the association with the free form of N and with the ribonucleoprotein is not clear for HMPV or other major human pathogens, including the viruses that cause measles, Ebola and rabies. Here we report a cryo-electron microscopy reconstruction that shows the ring-shaped structure of the polymerase and capping domains of HMPV-L bound to a tetramer of P. The connector and methyltransferase domains of L are mobile with respect to the core. The putative priming loop that is important for the initiation of RNA synthesis is fully retracted, which leaves space in the active-site cavity for RNA elongation. P interacts extensively with the N-terminal region of L, burying more than 4,016 Å2 of the molecular surface area in the interface. Two of the four helices that form the coiled-coil tetramerization domain of P, and long C-terminal extensions projecting from these two helices, wrap around the L protein in a manner similar to tentacles. The structural versatility of the four P protomers—which are largely disordered in their free state—demonstrates an example of a ‘folding-upon-partner-binding’ mechanism for carrying out P adaptor functions. The structure shows that P has the potential to modulate multiple functions of L and these results should accelerate the design of specific antiviral drugs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
van den Hoogen, B. G. et al. A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat. Med. 7, 719–724 (2001).
Neubauer, J., Ogino, M., Green, T. J. & Ogino, T. Signature motifs of GDP polyribonucleotidyltransferase, a non-segmented negative strand RNA viral mRNA capping enzyme, domain in the L protein are required for covalent enzyme–pRNA intermediate formation. Nucleic Acids Res. 44, 330–341 (2016).
Liang, B. et al. Structure of the L protein of vesicular stomatitis virus from electron cryomicroscopy. Cell 162, 314–327 (2015).
Pflug, A., Guilligay, D., Reich, S. & Cusack, S. Structure of influenza A polymerase bound to the viral RNA promoter. Nature 516, 355–360 (2014).
Paesen, G. C. et al. X-ray structure and activities of an essential Mononegavirales L-protein domain. Nat. Commun. 6, 8749 (2015).
Qiu, S., Ogino, M., Luo, M., Ogino, T. & Green, T. J. Structure and function of the N-terminal domain of the vesicular stomatitis virus RNA polymerase. J. Virol. 90, 715–724 (2015).
Deval, J. et al. Molecular basis for the selective inhibition of respiratory syncytial virus RNA polymerase by 2′-fluoro-4′chloromethyl-cytidine triphosphate. PLoS Pathog. 11, e1004995 (2015).
Tao, Y., Farsetta, D. L., Nibert, M. L. & Harrison, S. C. RNA synthesis in a cage—structural studies of reovirus polymerase λ3. Cell 111, 733–745 (2002).
te Velthuis, A. J. W., Robb, N. C., Kapanidis, A. N. & Fodor, E. The role of the priming loop in influenza A virus RNA synthesis. Nat. Microbiol. 1, 16029 (2016).
Ogino, M., Gupta, N., Green, T. J. & Ogino, T. A dual-functional priming-capping loop of rhabdoviral RNA polymerases directs terminal de novo initiation and capping intermediate formation. Nucleic Acids Res. 47, 299–309 (2019).
Leyrat, C., Renner, M., Harlos, K. & Grimes, J. M. Solution and crystallographic structures of the central region of the phosphoprotein from human metapneumovirus. PLoS ONE 8, e80371 (2013).
Blondot, M. L. et al. Structure and functional analysis of the RNA- and viral phosphoprotein-binding domain of respiratory syncytial virus M2-1 protein. PLoS Pathog. 8, e1002734 (2012).
Takacs, A. M., Barik, S., Das, T. & Banerjee, A. K. Phosphorylation of specific serine residues within the acidic domain of the phosphoprotein of vesicular stomatitis virus regulates transcription in vitro. J. Virol. 66, 5842–5848 (1992).
Tawar, R. G. et al. Crystal structure of a nucleocapsid-like nucleoprotein–RNA complex of respiratory syncytial virus. Science 326, 1279–1283 (2009).
Renner, M. et al. Nucleocapsid assembly in pneumoviruses is regulated by conformational switching of the N protein. eLife 5, e12627 (2016).
Gilman, M. S. A. et al. Structure of the respiratory syncytial virus polymerase complex. Cell 179, 193–204 (2019).
Sourimant, J. et al. Fine mapping and characterization of the L-polymerase-binding domain of the respiratory syncytial virus phosphoprotein. J. Virol. 89, 4421–4433 (2015).
Chenik, M., Schnell, M., Conzelmann, K. K. & Blondel, D. Mapping the interacting domains between the rabies virus polymerase and phosphoprotein. J. Virol. 72, 1925–1930 (1998).
Biacchesi, S. et al. Genetic diversity between human metapneumovirus subgroups. Virology 315, 1–9 (2003).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Kouba, T., Drncová, P. & Cusack, S. Structural snapshots of actively transcribing influenza polymerase. Nat. Struct. Mol. Biol. 26, 460–470 (2019).
Appleby, T. C. et al. Structural basis for RNA replication by the hepatitis C virus polymerase. Science 347, 771–775 (2015).
Kleywegt, G. J. & Jones, T. A. Detection, delineation, measurement and display of cavities in macromolecular structures. Acta Crystallogr. D 50, 178–185 (1994).
Kalocsay, M. APEX peroxidase-catalyzed proximity labeling and multiplexed quantitative proteomics. Methods Mol. Biol. 2008, 41–55 (2019).
Buchholz, U. J., Finke, S. & Conzelmann, K. K. Generation of bovine respiratory syncytial virus (BRSV) from cDNA: BRSV NS2 is not essential for virus replication in tissue culture, and the human RSV leader region acts as a functional BRSV genome promoter. J. Virol. 73, 251–259 (1999).
Bond, C. S. TopDraw: a sketchpad for protein structure topology cartoons. Bioinformatics 19, 311–312 (2003).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Poch, O., Blumberg, B. M., Bougueleret, L. & Tordo, N. Sequence comparison of five polymerases (L proteins) of unsegmented negative-strand RNA viruses: theoretical assignment of functional domains. J. Gen. Virol. 71, 1153–1162 (1990).
van den Hoogen, B. G., Bestebroer, T. M., Osterhaus, A. D. M. E. & Fouchier, R. A. M. Analysis of the genomic sequence of a human metapneumovirus. Virology 295, 119–132 (2002).
Bruenn, J. A. A structural and primary sequence comparison of the viral RNA-dependent RNA polymerases. Nucleic Acids Res. 31, 1821–1829 (2003).
Lang, D. M., Zemla, A. T. & Zhou, C. L. E. Highly similar structural frames link the template tunnel and NTP entry tunnel to the exterior surface in RNA-dependent RNA polymerases. Nucleic Acids Res. 41, 1464–1482 (2013).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Mason, S. W. et al. Interaction between human respiratory syncytial virus (RSV) M2-1 and P proteins is required for reconstitution of M2-1-dependent RSV minigenome activity. J. Virol. 77, 10670–10676 (2003).
Asenjo, A., Calvo, E. & Villanueva, N. Phosphorylation of human respiratory syncytial virus P protein at threonine 108 controls its interaction with the M2-1 protein in the viral RNA polymerase complex. J. Gen. Virol. 87, 3637–3642 (2006).
Acknowledgements
We thank S. C. Harrison for support, encouragement, suggestions and reading of the manuscript and acknowledge funding from NIH grant AI089618 (to S. C. Harrison). We thank Z. Li for cryo-EM support, M. Luo and M. Liao for suggestions and help with cryo-EM data acquisition, Y. Liu for suggestions on image-processing strategies, S. Jenni, S. M. Hinshaw and S. P. Lim for suggestions and critiques, and A. Shareef for insights regarding HMPV RNA synthesis assays. The J.L. laboratory was supported by Academic Research Grant Tier 3 (MOE2016-T3-1-003) from the Singapore Minister of Education (MOE) and grant NRF2016NRF-CRP001-063. X.Q. was supported by the Singapore-MIT Alliance for Research and Technology. The R.F. laboratory was supported by NIH grant AI113321.
Author information
Authors and Affiliations
Contributions
J.P., X.Q., R.F. and J.L. conceived the study. X.Q. and T.H.Y. expressed and purified HMPV-L:P. J.P. collected and processed cryo-EM data. J.P. and J.L. built and refined the structure. S.L. carried out the polymerase activity assays. M.K. performed phosphorylation studies. J.P. and A.E.S. analysed data and prepared figures. H.J. and X.Q. prepared the L mutant. T.C. and S.N. performed priming loop assays and minigenome studies. B.L. performed the primer elongation assay. J.P., X.Q., R.F. and J.L. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Tomoaki Ogino, Aartjan te Velthuis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification of HMPV-L:P and structure determination using cryo-EM.
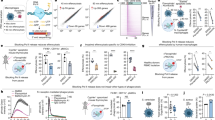
a, Representative size-exclusion chromatogram of the L:P complex (these experiments were repeated more than five times). Fractions indicated by an arrow were collected and concentrated to 0.85 mg ml−1 and used for cryo-EM analysis. Inset, SDS–PAGE followed by Coomassie blue staining of the purified samples. Free P protein separated from the L:P complex by heparin chromatography is also shown. For gel source data, see Supplementary Fig. 1. b, Raw micrograph of HMPV-L:P particles recorded in vitreous ice. Scale bar, 10 nm. c, Power spectrum of the image shown in b. We limited the high resolution for fitting to a spatial frequency of 1/5.0 Å, and 1/2.9 Å marks the highest spacing to which CTF rings were successfully fit. d, Two-dimensional classes and ‘self-consistency check’ for the cryo-EM three-dimensional reconstruction. For each pair of boxes (top and bottom) across the three rows, the top image shows one two-dimensional class average and the bottom image shows the corresponding projection from the initial three-dimensional model. e, Local resolution of the cryo-EM density map. Variations in local resolution are colour-coded from blue (3.0 Å) to red (5.9 Å), computed with Resmap44. f, FSC of the cryo-EM map as a function of the spatial frequency. The gold-standard resolution is 3.7 Å based on the FSC = 0.143 criterion, consistent with the model-to-map correlation (0.5 criterion). g, Example of the electron-density map that enabled model building. The region shown is at an interface between the RdRp and capping domain. The map is shown as a grey mesh, contoured at a level of 3σ. The atomic model is shown as sticks with residues from RdRp coloured in cyan (NTD in grey) and in green for the capping domain. h, The region shown is the three-stranded β-sheet at the interface between the RdRp (cyan sticks) and the phosphoprotein (magenta sticks). The map is shown as a grey mesh, contoured at a level of 2.5σ. We observed a nearly identical structure of the L:P complex in a reconstruction obtained by premixing the L:P complex with fully phosphorylated P, indicating that potential exchange of P affected neither the formation nor the structure of the L:P complex.
Extended Data Fig. 2 RdRp activity assay.
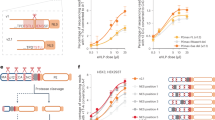
a, SDS–PAGE of HMPV wild-type L:P, L(D745A):P purified for RdRp activity assays. Proteins were purified by metal affinity, TEV cleavage of the His-tag followed by reverse His-tag affinity purification and size-exclusion chromatography. b, Analysis of the 3′ extension activity of HMPV polymerase using the le25 RNA template. Reactions were performed with rNTPs (0.5 mM each of rUTP, rGTP and rCTP), 20 μM rATP and 20 nM [α-32P]rATP. When a 3′-modified le25 (le25[SpC3], three-carbon spacer group linked to the 3′ extremity) was used as a template, synthesis of products longer than 25 nt was greatly reduced compared to le25. When only [α-32P]rATP and no other rNTP was supplied, only a product with a length longer than 25 nt was observed. This result shows that the L:P complex was capable of modifying the 3′ terminus of the template, in addition to engaging in de novo initiation at the promoter. The radiolabelled RNA products were visualized by phosphor imaging. Data are representative of three independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Flow-chart depicting structure determination using cryo-EM.
Further information is provided in the Methods.
Extended Data Fig. 4 Phosphoprotein tetramer in complex with L.
a, The L protein (cyan) is represented as a molecular surface and the tetrameric P protein subunits are represented as ribbons, following the colour codes in Figs. 1–3. b, Structures adopted by the four individual P subunits bound to L, coloured as a blue-to-red rainbow from the N- to the C-terminal end. Secondary structures boundaries are noted for each subunit. c, d, Superposition of the tetramerization helices in the context of the L:P complex and the free P protein. Structures are represented as coloured ribbons with the free phosphoprotein coiled-coil (PDB 4BXT) coloured in grey and the four P subunits reported in this study coloured according to Fig. 1. The r.m.s.d. of the superimposition is 1.13 Å over 88 α-carbon atoms. e, View of the complex in which L and P have been pulled apart to display electrostatic surfaces. f, Overall view of the L:P complex with P shown as ribbons and L as electrostatic surface. The P tetramer consists of subunits P1 (magenta), P2 (hot pink), P3 (salmon) and P4 (pink).
Extended Data Fig. 5 Topology of the L:P complex.
Topological depiction of the secondary structure elements of L and P. Helices are depicted as tubes and strands as arrows. The colour code is the same as in Fig. 1. The RdRp domain and its subdomains and the capping domain are coloured as in Figs. 1, 2: NTD in grey, fingers in blue, palm in red, thumb in dark green and CAP in green. The four subunits of the phosphoprotein P1, P2, P3 and P4 are coloured as in Fig. 1. Secondary structure boundaries are indicated.
Extended Data Fig. 6 View of the NTD.
NTD is displayed as grey ribbons (colours are as in Fig. 2a). The evolutionary conserved residues clustered near the rNTP entry tunnel play a part in transcription and are represented as sticks and are labelled. The equivalent region of the VSV-L superimposed on HMPV-L is coloured in lighter grey.
Extended Data Fig. 7 Model for an elongation complex stalled by the addition of ALS-8176 5′-triphosphate.
ALS-8176 5′-triphosphate is a nucleoside triphosphate analogue against RSV and HMPV currently in phase 2 clinical trials. The 744GDNQ747 catalytic motif and the positions (A789V, L795I and I796V) for which mutations conferred resistance (identified by passaging RSV) are mapped onto the HMPV-L structure (respectively corresponding to A723, V729 and V730) and displayed as sticks. These conservative mutations probably affect inhibitor binding by inducing a slight repositioning of the helix, owing to altered hydrophobic contacts with neighbouring helices. The sequence alignment is shown in Supplementary Fig. 2. the protein is coloured according to Fig. 2a, and the template and nascent RNA strands according to Fig. 3a.
Extended Data Fig. 8 MS2 spectrum of the Ser148 P phosphopeptide.
a, One MS2 spectrum used for identification of the phosphorylated P peptide 142DALDLLS#DNEEEDAESSILTFEER is displayed. Tandem mass spectrum (top) and deviation (bottom) enabled detection of phosphorylation (indicated by a hash symbol) at site Ser148. Peptides fragmented from the N terminus (b-fragments) and C terminus (y-fragments) are coloured in blue and red, respectively. b, The y and b ion series m/z identified in the spectrum shown in a and their deviation from theoretical m/z are shown in the table. The present pattern of phosphorylation agrees with observations showing that phosphorylation of the peptide comprising residues 100–120 of RSV-P45—in particular, phosphorylation of Thr10846, which corresponds to Ser148 of HMPV-P (Extended Data Fig. 9)—controls its interaction with the M2-1 protein.
Extended Data Fig. 9 Structure-based sequence alignment of the phosphoprotein from HMPV (labelled HMPV-A, strain CAN97-83) and other known pneumoviral P proteins.
HMPV-B, human metapneumovirus subgroup B; HRSV-A and -B, human respiratory syncytial virus subgroup A and B, respectively; BRSV, bovine respiratory syncytial virus; PVM, pneumonia virus of mice; AMPV-A and AMPV-C, avian metapneumovirus subgroup A and C, respectively. The following sequence accession codes from GenBank were used for the alignment: HMPV-A, AAQ67693.1 (used in this study); HMPV-B, AAQ67684.1; HRSV-A, AAX23990.1; HRSV-B, AAR14262.1; BRSV, AAL49395.1; PVM, AAW79177.1; AMPV-A, AAT68644.1; and AMPV-C, AAT86110.1. The secondary structure of HMPV-P subunit P1 (this study) is displayed above the alignment. Phosphorylation sites are highlighted in brown. Positively charged residues of HMPV_P are shaded in blue, negatively charged residues in magenta and hydrophobic residues 29–135 in yellow. The conserved region containing hydrophobic residues critical for L:P interactions is highlighted in green. Structural alignment of P from HMPV and RSV16 showed similar overall tetramer organization. However, differences are observed in subunit P1 with an r.m.s.d. of 2.24 Å over 82 residues. Although P is generally more mobile with weaker densities and higher B-factors compared to L, the region following the β-hairpin (residues 175–215 in HMPV) does adopt a slightly different conformation compared to RSV P1. Subunit P3 has an r.m.s.d. of 1.94 Å over 45 residues, owing to a slightly tilted C-terminal helix compared to RSV. Subunit P2 is most similar with an r.m.s.d. of 0.92 Å over 56 residues. Subunit P4 has an r.m.s.d. of 1.33 Å over 47 residues. The eight residues of HRSV-P, which are crucial for the interaction with HRSV-L and substitutions of which impair viral replication, are shaded in dark green (data from a previous study16). With the exception of Asn189 (HRSV-P), for which a deletion is present in HMPV-P, these residues are conserved in HMPV-P and other known pneumoviral P proteins.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-2.
Video 1: Overall view of the HMPV L:P complex
Overall structure and organization of the L:P complex. L:P is presented as coloured ribbons and then as coloured molecular surfaces, with the NTD in grey, the RdRp domain in cyan and the capping domain in green. This is followed by L being represented as a molecular surface coloured by electrostatic potential (blue: positive; red: negative). The phosphoprotein asymmetric tetramer comprised of subunits P1, P2, P3 and P4 are represented as coloured ribbons: P1 in magenta, P2: hotpink, P3: salmon and P4: pink. The entry tunnel for rNTPs, exit site of nascent RNA as well as the entry and exit tunnel for the RNA template are indicated.
Video 2: Close-up views of residues of P interacting with L
L is represented by its molecular surface colored in cyan. P is represented as colored ribbons: in magenta for P1, hotpink for P2, salmon for P3 and pink for P4. The N terminal and C terminal termini of each phosphoprotein subunit are shown. The residues in contacts between P1 and L are displayed and represented as sticks.
Source data
Rights and permissions
About this article
Cite this article
Pan, J., Qian, X., Lattmann, S. et al. Structure of the human metapneumovirus polymerase phosphoprotein complex. Nature 577, 275–279 (2020). https://doi.org/10.1038/s41586-019-1759-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1759-1
This article is cited by
-
Structures of the promoter-bound respiratory syncytial virus polymerase
Nature (2024)
-
Structural basis for dimerization of a paramyxovirus polymerase complex
Nature Communications (2024)
-
An intermediate state allows influenza polymerase to switch smoothly between transcription and replication cycles
Nature Structural & Molecular Biology (2023)
-
Structure of the Newcastle Disease Virus L protein in complex with tetrameric phosphoprotein
Nature Communications (2023)
-
Conserved allosteric inhibitory site on the respiratory syncytial virus and human metapneumovirus RNA-dependent RNA polymerases
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.