Abstract
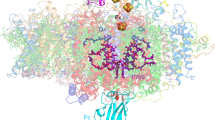
The cytochrome b6 f (cytb6 f ) complex has a central role in oxygenic photosynthesis, linking electron transfer between photosystems I and II and converting solar energy into a transmembrane proton gradient for ATP synthesis1,2,3. Electron transfer within cytb6 f occurs via the quinol (Q) cycle, which catalyses the oxidation of plastoquinol (PQH2) and the reduction of both plastocyanin (PC) and plastoquinone (PQ) at two separate sites via electron bifurcation2. In higher plants, cytb6 f also acts as a redox-sensing hub, pivotal to the regulation of light harvesting and cyclic electron transfer that protect against metabolic and environmental stresses3. Here we present a 3.6 Å resolution cryo-electron microscopy (cryo-EM) structure of the dimeric cytb6 f complex from spinach, which reveals the structural basis for operation of the Q cycle and its redox-sensing function. The complex contains up to three natively bound PQ molecules. The first, PQ1, is located in one cytb6 f monomer near the PQ oxidation site (Qp) adjacent to haem bp and chlorophyll a. Two conformations of the chlorophyll a phytyl tail were resolved, one that prevents access to the Qp site and another that permits it, supporting a gating function for the chlorophyll a involved in redox sensing. PQ2 straddles the intermonomer cavity, partially obstructing the PQ reduction site (Qn) on the PQ1 side and committing the electron transfer network to turnover at the occupied Qn site in the neighbouring monomer. A conformational switch involving the haem cn propionate promotes two-electron, two-proton reduction at the Qn site and avoids formation of the reactive intermediate semiquinone. The location of a tentatively assigned third PQ molecule is consistent with a transition between the Qp and Qn sites in opposite monomers during the Q cycle. The spinach cytb6 f structure therefore provides new insights into how the complex fulfils its catalytic and regulatory roles in photosynthesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data are available from the authors and/or are included with the manuscript or in the Supplementary Information. Atomic coordinates and the cryo-EM density map have been deposited in the Protein Data Bank under accession number 6RQF and the Electron Microscopy Data Bank (EMDB) under accession number EMD-4981.
References
Hill, R. & Bendall, F. Function of the two cytochrome components in chloroplasts: a working hypothesis. Nature 186, 136–137 (1960).
Cramer, W. A., Hasan, S. S. & Yamashita, E. The Q cycle of cytochrome bc complexes: a structure perspective. Biochim. Biophys. Acta 1807, 788–802 (2011).
Tikhonov, A. N. The cytochrome b 6f complex at the crossroad of photosynthetic electron transport pathways. Plant Physiol. Biochem. 81, 163–183 (2014).
Xia, D. et al. Crystal structure of the cytochrome bc 1 complex from bovine heart mitochondria. Science 277, 60–66 (1997).
Esser, L. et al. Inhibitor-complexed structures of the cytochrome bc 1 from the photosynthetic bacterium Rhodobacter sphaeroides. J. Biol. Chem. 283, 2846–2857 (2008).
Cape, J. L., Bowman, M. K. & Kramer, D. M. Understanding the cytochrome bc complexes by what they don’t do. The Q-cycle at 30. Trends Plant Sci. 11, 46–55 (2006).
Stroebel, D., Choquet, Y., Popot, J.-L. & Picot, D. An atypical haem in the cytochrome b 6f complex. Nature 426, 413–418 (2003).
Kurisu, G., Zhang, H., Smith, J. L. & Cramer, W. A. Structure of the cytochrome b 6f complex of oxygenic photosynthesis: tuning the cavity. Science 302, 1009–1014 (2003).
Baniulis, D. et al. Structure–function, stability, and chemical modification of the cyanobacterial cytochrome b 6f complex from Nostoc sp. PCC 7120. J. Biol. Chem. 284, 9861–9869 (2009).
Bellafiore, S., Barneche, F., Peltier, G. & Rochaix, J. D. State transitions and light adaptation require chloroplast thylakoid protein kinase STN7. Nature 433, 892–895 (2005).
Yamori, W. & Shikanai, T. Physiological functions of cyclic electron transport around photosystem I in sustaining photosynthesis and plant growth. Annu. Rev. Plant Biol. 67, 81–106 (2016).
Horton, P. & Black, M. T. Activation of adenosine 5′-triphosphate induced quenching of chlorophyll fluorescence by reduced plastoquinone. The basis of state I–state II transitions in chloroplasts. FEBS Lett. 119, 141–144 (1980).
Vener, A. V., van Kan, P. J. M., Rich, P. R., Ohad, I. & Andersson, B. Plastoquinol at the quinol oxidation site of reduced cytochrome bf mediates signal transduction between light and protein phosphorylation: thylakoid protein kinase deactivation by a single-turnover flash. Proc. Natl Acad. Sci. USA 94, 1585–1590 (1997).
Gal, A., Hauska, G., Herrmann, R. & Ohad, I. Interaction between light harvesting chlorophyll-a/b protein (LHCII) kinase and cytochrome b 6/f complex. In vitro control of kinase activity. J. Biol. Chem. 265, 19742–19749 (1990).
Allen, J. F., Bennett, J., Steinback, K. E. & Arntzen, C. J. Chloroplast protein phosphorylation couples plastoquinone redox state to distribution of excitation energy between photosystems. Nature 291, 25–29 (1981).
Wood, W. H. J. et al. Dynamic thylakoid stacking regulates the balance between linear and cyclic photosynthetic electron transfer. Nat. Plants 4, 116–127 (2018).
Zhang, H., Whitelegge, J. P. & Cramer, W. A. Ferredoxin:NADP+ oxidoreductase is a subunit of the chloroplast cytochrome b 6f complex. J. Biol. Chem. 276, 38159–38165 (2001).
Zhu, X. G., Long, S. P. & Ort, D. R. Improving photosynthetic efficiency for greater yield. Annu. Rev. Plant Biol. 61, 235–261 (2010).
Simkin, A. J., McAusland, L., Lawson, T. & Raines, C. A. Overexpression of the RieskeFeS protein increases electron transport rates and biomass yield. Plant Physiol. 175, 134–145 (2017).
Zhang, Z. et al. Electron transfer by domain movement in cytochrome bc 1. Nature 392, 677–684 (1998).
Yan, J., Kurisu, G. & Cramer, W. A. Intraprotein transfer of the quinone analogue inhibitor 2,5-dibromo-3-methyl-6-isopropyl-p-benzoquinone in the cytochrome b6 f complex. Proc. Natl Acad. Sci. USA 103, 69–74 (2006).
Yamashita, E., Zhang, H. & Cramer, W. A. Structure of the cytochrome b 6f complex: quinone analogue inhibitors as ligands of heme c n. J. Mol. Biol. 370, 39–52 (2007).
Hasan, S. S., Yamashita, E., Baniulis, D. & Cramer, W. A. Quinone-dependent proton transfer pathways in the photosynthetic cytochrome b 6 f complex. Proc. Natl Acad. Sci. USA 110, 4297–4302 (2013).
Sarewicz, M. et al. Metastable radical state, nonreactive with oxygen, is inherent to catalysis by respiratory and photosynthetic cytochromes bc 1/b 6f. Proc. Natl Acad. Sci. USA 114, 1323–1328 (2017).
Singh, S. K. et al. Trans-membrane signalling in photosynthetic state transitions: redox- and structure-dependent interaction in vitro between Stt7 kinase and the cytochrome b6f complex. J. Biol. Chem. 291, 21740–21750 (2016).
Zito, F., Vinh, J., Popot, J. L. & Finazzi, G. Chimeric fusions of subunit IV and PetL in the b 6f complex of Chlamydomonas reinhardtii: structural implications and consequences on state transitions. J. Biol. Chem. 277, 12446–12455 (2002).
Saif Hasan, S., Yamashita, E. & Cramer, W. A. Transmembrane signaling and assembly of the cytochrome b 6f-lipidic charge transfer complex. Biochim. Biophys. Acta 1827, 1295–1308 (2013).
Nawrocki, W. J. et al. The mechanism of cyclic electron flow. Biochim. Biophys. Acta 1860, 433–438 (2019).
Osyczka, A., Moser, C. C., Daldal, F. & Dutton, P. L. Reversible redox energy coupling in electron transfer chains. Nature 427, 607–612 (2004).
Świerczek, M. et al. An electronic bus bar lies in the core of cytochrome bc 1. Science 329, 451–454 (2010).
Alric, J., Pierre, Y., Picot, D., Lavergne, J. & Rappaport, F. Spectral and redox characterization of the heme c i of the cytochrome b 6f complex. Proc. Natl Acad. Sci. USA 102, 15860–15865 (2005).
Dietrich, J. & Kühlbrandt, W. Purification and two-dimensional crystallization of highly active cytochrome b 6 f complex from spinach. FEBS Lett. 463, 97–102 (1999).
Porra, R. J., Thompson, W. A. & Kriedemann, P. E. Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim. Biophys. Acta 975, 384–394 (1989).
Cramer, W. A. & Whitmarsh, J. Photosynthetic cytochromes. Annu. Rev. Plant Physiol. 28, 133–172 (1977).
Dawson, R. M. C., Elliot, D. C., Elliot, W. H. & Jones, K. M. Data for Biochemical Research (Clarendon, 1986).
Tan, S. & Ho, K. K. Purification of an acidic plastocyanin from Microcystis aeruginosa. Biochim. Biophys. Acta 973, 111–117 (1989).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D 73, 496–502 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Hasan, S. S. & Cramer, W. A. Internal lipid architecture of the hetero-oligomeric cytochrome b 6 f complex. Structure 22, 1008–1015 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Proctor, M. S. et al. Plant and algal chlorophyll synthases function in Synechocystis and interact with the YidC/Alb3 membrane insertase. FEBS Lett. 592, 3062–3073 (2018).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand and contrast loss in single particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Acknowledgements
M.P.J. acknowledges funding from the Leverhulme Trust grant RPG-2016-161. C.N.H., P.Q., A.H., D.J.K.S. and M.P.J. also acknowledge financial support from the Biotechnology and Biological Sciences Research Council (BBSRC UK) award numbers BB/M000265/1 and BB/P002005/1. L.A.M. was supported by a White Rose doctoral studentship, G.E.M. was supported by a doctoral studentship from The Grantham Foundation and D.A.F. was supported by a University of Sheffield doctoral scholarship. Cryo-EM data was collected at the Astbury Biostructure Laboratory funded by the University of Leeds (ABSL award) and the Wellcome Trust (108466/Z/15/Z). We thank S. Tzokov, J. Bergeron, J. Wilson and D. Mann for their helpful advice and assistance with the EM and data processing.
Author information
Authors and Affiliations
Contributions
P.Q., C.N.H., N.A.R. and M.P.J. supervised the project. L.A.M., G.E.M., P.Q., C.N.H., R.F.T. and M.P.J. designed the experiments. L.A.M. and G.E.M. purified the cytb6 f complex, L.A.M., G.E.M., A.H. and D.J.K.S. characterized the cytb6 f complex. L.A.M., P.Q., D.A.F. and R.F.T. collected, processed and/or analysed the cryo-EM data. L.A.M., C.N.H. and M.P.J. wrote the manuscript. All authors proofread and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Zhenfeng Liu, Alexander Tikhonov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Purification of cytb6 f from spinach.
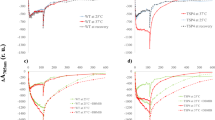
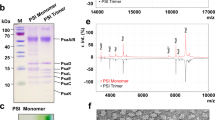
a, Absorption spectrum of ascorbate-reduced purified b6 f complex. The peak at 421 nm corresponds to the Soret band of bound pigments (chlorophyll a and haems). The peaks at 554 and 668 nm correspond to c-type haems of cytf and chlorophyll a, respectively. The inset panel shows redox difference spectra of ascorbate-reduced minus ferricyanide-oxidized b6 f (dashed line) and dithionite-reduced minus ascorbate-reduced (dotted line) cytb6 f. Redox difference spectra show haem f absorption peaks at 523 and 554 nm as well as absorption peaks at 534 and 563 nm corresponding to the b-type haems of cytb6. The calculated ratio of cytb6 b-type haems to the c-type haem of cytf was ~2 using extinction coefficients of 25 mM cm−1 ( f ) and 21 mM cm−1 (b6)34. The spectra exhibit the absorption properties characteristic of intact cytb6 f. Spectra were recorded at room temperature. b, SDS–PAGE analysis of purified cytb6 f indicates that the sample is highly pure, with the four large subunits of the complex (cytf, cytb6, the Rieske ISP and subunit IV) running at ~31 kDa, ~24 kDa, ~20 kDa and ~17 kDa, respectively and the four small subunits (PetG, PetL, PetM and PetN) running at around 4 kDa (not shown). c, d, Negative-stain and BN-PAGE analysis of purified cytb6 f demonstrates the sample is dimeric and highly homogenous, with a single band corresponding to dimeric cytb6 f shown in lane 1. Lane 2 shows a sample that has been deliberately monomerized following incubation with 1% Triton X-100 for 1 h. For gel source data see Supplementary Fig. 1. e, The catalytic rate of plastocyanin reduction by the purified dimeric cytb6 f complex as determined by stopped-flow absorbance spectroscopy. A rate of 200 e− s−1 was determined by taking the initial linear region from the enzyme-catalysed reaction (solid line) and subtracting the background rate measured in the absence of enzyme (long-dashed line). Plastocyanin reduction was not observed in the absence of decylplastoquinol (short-dashed line). Reactions were initiated upon addition of decylplastoquinol to the solution containing plastocyanin and b6 f while monitoring the loss of absorbance at 597 nm. Final concentrations were 50 µM plastocyanin, 185 nm b6 f and 250 µM decylplastoquinol. All experiments were performed in triplicate and controls were performed in the absence of b6 f or decylplastoquinol.
Extended Data Fig. 2 Cryo-EM micrographs of the spinach cytb6 f complex and calculation of the cryo-EM map global and local resolution.
a, Cytb6 f particles covered by a thin layer of vitreous ice on a supported carbon film. b, Examples of dimeric cytb6 f particles are circled in green. We recorded 6,035 cryo-EM movies, from which 422,660 particles were picked manually for reference-free 2D classification. The final density map was calculated from 108,560 particles. c, Gold-standard refinement was used for estimation of the final map resolution (solid black line). The global resolution of 3.58 Å was calculated using a FSC cut-off at 0.143. A model-to-map FSC curve (solid grey line) was also calculated. d, e, A C1 density map of the cytb6 f complex both with (d) and without (e) the detergent shell. The map is coloured according to local resolution estimated by RELION and viewed from within the plane of the membrane. The colour key on the right shows the local structural resolution in angstroms (Å).
Extended Data Fig. 3 Cryo-EM densities and structural models of polypeptides in the cytb6 f complex.
Polypeptides are coloured as in Fig. 1. The contour levels of the density maps were adjusted to 0.0144.
Extended Data Fig. 4 Cryo-EM densities and structural models of prosthetic groups, lipids and plastoquinone molecules in the cytb6 f complex.
c-type haems ( f, cn; dark blue), b-type haems (bp, bn; red), 9-cis β-carotene (orange), chlorophyll a (major conformation, dark green; minor conformation, light green), 2Fe-2S (burnt orange and yellow), plastoquinones (yellow), monogalactosyl diacylglycerol (light pink), phosphatidylcholine (light cyan), sulfoquinovosyl diacylglycerol (light green) and phosphatidylglycerol (light purple). The contour levels of the density maps were adjusted to 0.0068.
Extended Data Fig. 5 Alternative interpretation of the region assigned as PQ2.
a, b, The density map showing two possible alternative conformations for PQ2, the major conformation (a) and the alternative conformation (b). Cofactors are coloured as in Extended Data Fig. 4 with b-type haems (bp and bn) coloured red, c-type haems (cn) coloured dark blue, chlorophyll a (major conformation) coloured dark green, plastoquinones coloured yellow and the cytb6 subunit coloured light green. The contour level of the density map was adjusted to 0.0089.
Extended Data Fig. 6 Alternative interpretations of the density map in the region assigned as PQ3.
a, b, The density map modelled with a plastoquinone molecule (a) and a phosphatidylcholine molecule (b). Top, the protein-free density map; bottom, the map including cytb6 (green). The 2.9 Å distance indicates a close contact between the PQ3 head group and the conserved Lys208. Cofactors are coloured as in Extended Data Fig. 4 with b-type haems (bp and bn) coloured red, chlorophyll a (major conformation) in dark green, plastoquinones in yellow, phosphatidylcholine in light cyan, sulfoquinovosyl diacylglycerol in mint green and the cytb6 subunit in light green. The contour level of the density map was adjusted to 0.0127.
Extended Data Fig. 7 Multiple sequence alignment of cytb6 f subunits cytf and cytb6.
a, b, Sequences of cytf (a) and cytb6 (b) from cyanobacterial (M. laminosus and Nostoc sp. PCC7120), algal (C. reinhardtii) and plant (S. oleracea) subunits were aligned in Clustal Omega v.1.2.4. Conserved identities are indicated by asterisks, and similarities by double or single dots. Polar residues are coloured in green, positively charged residues are pink, hydrophobic residues are red and negatively charged residues are blue. The sequences omit signal peptides.
Extended Data Fig. 8 Multiple sequence alignment of the Rieske ISP, subunit IV, PetG, PetL, PetM and PetN.
a–f, Sequences of Rieske ISP (a), subunit IV (b), PetG (c), PetL (d), PetM (e) and PetN (f) from cyanobacterial (M. laminosus and Nostoc sp. PCC7120), algal (C. reinhardtii) and plant (S. oleracea) subunits were aligned in Clustal Omega v.1.2.4. Conserved identities are indicated by asterisks, and similarities by double or single dots. Polar residues are coloured in green, positively charged residues are pink, hydrophobic residues are red and negatively charged residues are blue. The sequences omit signal peptides.
Supplementary information
Supplementary Figure 1
.Raw gel images of SDS-PAGE and Native-PAGE gels shown in Extended Data Fig. 1 b and c. a, SDS-PAGE (NuPAGE 12% Bis-Tris Gel; Invitrogen) analysis of the purified cyt b6f dimer. Lane 1, 10 µl Precision Plus Protein 10-250 kDa Unstained Standard (BioRad); lane 2, 1 µl of purified b6f dimer (~17 µM); lane 3, 2 µl of purified b6f dimer (~17 µM); lane 4, 3 µl of purified b6f dimer (~17 µM). The positions of cyt b6f subunits are marked. Empty lanes are indicated by a (-). Image cropped to include only lanes 1 and 2 as indicated by the black dashed box. b, Native-PAGE (Native-PAGE 3-12% Bis-Tris Gel; Invitrogen) analysis of the purified cyt b6f. Lane 1, 2 µl dimeric cyt b6f (~17 µM); lane 2, 1 µl of dimeric cyt b6f (~17µM) incubated with 1% triton for 1 hour to monomerise sample. The positions of cyt b6f monomer and dimer are marked. Image cropped to include only lanes 2 and 3 as indicated by the black dashed box.
Rights and permissions
About this article
Cite this article
Malone, L.A., Qian, P., Mayneord, G.E. et al. Cryo-EM structure of the spinach cytochrome b6 f complex at 3.6 Å resolution. Nature 575, 535–539 (2019). https://doi.org/10.1038/s41586-019-1746-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1746-6
This article is cited by
-
The cytochrome b6f complex: plastoquinol oxidation and regulation of electron transport in chloroplasts
Photosynthesis Research (2024)
-
Peculiarities of DNP-INT and DBMIB as inhibitors of the photosynthetic electron transport
Photosynthesis Research (2023)
-
Absolute quantification of cellular levels of photosynthesis-related proteins in Synechocystis sp. PCC 6803
Photosynthesis Research (2023)
-
Combination of long-term 13CO2 labeling and isotopolog profiling allows turnover analysis of photosynthetic pigments in Arabidopsis leaves
Plant Methods (2022)
-
De-etiolation-induced protein 1 (DEIP1) mediates assembly of the cytochrome b6f complex in Arabidopsis
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.