Abstract
Heterochromatin affects genome function at many levels. It enables heritable gene repression, maintains chromosome integrity and provides mechanical rigidity to the nucleus1,2. These diverse functions are proposed to arise in part from compaction of the underlying chromatin2. A major type of heterochromatin contains at its core the complex formed between HP1 proteins and chromatin that is methylated on histone H3, lysine 9 (H3K9me). HP1 is proposed to use oligomerization to compact chromatin into phase-separated condensates3,4,5,6. Yet, how HP1-mediated phase separation relates to chromatin compaction remains unclear. Here we show that chromatin compaction by the Schizosaccharomyces pombe HP1 protein Swi6 results in phase-separated liquid condensates. Unexpectedly, we find that Swi6 substantially increases the accessibility and dynamics of buried histone residues within a nucleosome. Restraining these dynamics impairs compaction of chromatin into liquid droplets by Swi6. Our results indicate that Swi6 couples its oligomerization to the phase separation of chromatin by a counterintuitive mechanism, namely the dynamic exposure of buried nucleosomal regions. We propose that such reshaping of the octamer core by Swi6 increases opportunities for multivalent interactions between nucleosomes, thereby promoting phase separation. This mechanism may more generally drive chromatin organization beyond heterochromatin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data that support the findings of this study are available within the paper and its Supplementary Information files. Annotated XLMS Spectra are available using MS-Viewer at: http://msviewer.ucsf.edu/prospector/cgi-bin/msform.cgi?form=msviewer using the search key: 9ojpmsdzk5. Any additional data are available from the corresponding author upon reasonable request.
References
Stephens, A. D. et al. Chromatin histone modifications and rigidity affect nuclear morphology independent of lamins. Mol. Biol. Cell 29, 220–233 (2018).
Allshire, R. C. & Madhani, H. D. Ten principles of heterochromatin formation and function. Nat. Rev. Mol. Cell Biol. 19, 229–244 (2018).
Canzio, D. et al. Chromodomain-mediated oligomerization of HP1 suggests a nucleosome-bridging mechanism for heterochromatin assembly. Mol. Cell 41, 67–81 (2011).
Canzio, D. et al. A conformational switch in HP1 releases auto-inhibition to drive heterochromatin assembly. Nature 496, 377–381 (2013).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Eissenberg, J. C. & Elgin, S. C. R. HP1a: a structural chromosomal protein regulating transcription. Trends Genet. 30, 103–110 (2014).
Smothers, J. F. & Henikoff, S. The HP1 chromo shadow domain binds a consensus peptide pentamer. Curr. Biol. 10, 27–30 (2000).
Isaac, R. S. et al. Biochemical basis for distinct roles of the heterochromatin proteins Swi6 and Chp2. J. Mol. Biol. 429, 3666–3677 (2017).
Dawson, M. A. et al. JAK2 phosphorylates histone H3Y41 and excludes HP1α from chromatin. Nature 461, 819–822 (2009).
Lavigne, M. et al. Interaction of HP1 and Brg1/Brm with the globular domain of histone H3 is required for HP1-mediated repression. PLoS Genet. 5, e1000769 (2009).
Liu, Y. et al. Peptide recognition by HP1 chromoshadow domains revisited: plasticity in the pseudosymmetric histone binding site of human HP1. J. Biol. Chem. 292, 5655–5664 (2017).
Hoofnagle, A. N., Resing, K. A. & Ahn, N. G. Protein analysis by hydrogen exchange mass spectrometry. Annu. Rev. Biophys. Biomol. Struct. 32, 1–25 (2003).
Rosenzweig, R. & Kay, L. E. Bringing dynamic molecular machines into focus by methyl-TROSY NMR. Annu. Rev. Biochem. 83, 291–315 (2014).
Sinha, K. K., Gross, J. D. & Narlikar, G. J. Distortion of histone octamer core promotes nucleosome mobilization by a chromatin remodeler. Science 355, eaaa3761 (2017).
Zhou, B.-R. et al. Structural insights into the histone H1-nucleosome complex. Proc. Natl Acad. Sci. USA 110, 19390–19395 (2013).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Stoddard, C. I. et al. A nucleosome bridging mechanism for activation of a maintenance DNA methyltransferase. Mol. Cell 73, 73–83.e6 (2019).
Azzaz, A. M. et al. Human heterochromatin protein 1α promotes nucleosome associations that drive chromatin condensation. J. Biol. Chem. 289, 6850–6861 (2014).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Maeshima, K. et al. Nucleosomal arrays self-assemble into supramolecular globular structures lacking 30-nm fibers. EMBO J. 35, 1115–1132 (2016).
Cheutin, T., Gorski, S. A., May, K. M., Singh, P. B. & Misteli, T. In vivo dynamics of Swi6 in yeast: evidence for a stochastic model of heterochromatin. Mol. Cell. Biol. 24, 3157–3167 (2004).
Cheutin, T. et al. Maintenance of stable heterochromatin domains by dynamic HP1 binding. Science 299, 721–725 (2003).
Festenstein, R. et al. Modulation of heterochromatin protein 1 dynamics in primary Mammalian cells. Science 299, 719–721 (2003).
Haldar, S., Saini, A., Nanda, J. S., Saini, S. & Singh, J. Role of Swi6/HP1 self-association-mediated recruitment of Clr4/Suv39 in establishment and maintenance of heterochromatin in fission yeast. J. Biol. Chem. 286, 9308–9320 (2011).
Kilic, S., Bachmann, A. L., Bryan, L. C. & Fierz, B. Multivalency governs HP1α association dynamics with the silent chromatin state. Nat. Commun. 6, 7313 (2015).
Kalashnikova, A. A., Porter-Goff, M. E., Muthurajan, U. M., Luger, K. & Hansen, J. C. The role of the nucleosome acidic patch in modulating higher order chromatin structure. J. R. Soc. Interface 10, 20121022–20121022 (2013).
Aygün, O., Mehta, S. & Grewal, S. I. S. HDAC-mediated suppression of histone turnover promotes epigenetic stability of heterochromatin. Nat. Struct. Mol. Biol. 20, 547–554 (2013).
Fussner, E., Ching, R. W. & Bazett-Jones, D. P. Living without 30nm chromatin fibers. Trends Biochem. Sci. 36, 1–6 (2011).
Cai, S. et al. Cryo-ET reveals the macromolecular reorganization of S. pombe mitotic chromosomes in vivo. Proc. Natl Acad. Sci. USA 115, 10977–10982 (2018).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2003).
Tugarinov, V., Kanelis, V. & Kay, L. E. Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat. Protoc. 1, 749–754 (2006).
Hamel, D. J. & Dahlquist, F. W. The contact interface of a 120 kD CheA−CheW complex by methyl TROSY interaction spectroscopy. J. Am. Chem. Soc. 127, 9676–9677 (2005).
Simon, M. D. et al. The site-specific installation of methyl-lysine analogs into recombinant histones. Cell 128, 1003–1012 (2007).
Zhang, X. et al. Structure of the Neurospora SET domain protein DIM-5, a histone H3 lysine methyltransferase. Cell 111, 117–127 (2002).
Mishima, Y. et al. Hinge and chromoshadow of HP1α participate in recognition of K9 methylated histone H3 in nucleosomes. J. Mol. Biol. 425, 54–70 (2013).
Luger, K., Rechsteiner, T. J. & Richmond, T. J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999).
Dorigo, B. et al. Nucleosome arrays reveal the two-start organization of the chromatin fiber. Science 306, 1571–1573 (2004).
Rogge, R. A. et al. Assembly of nucleosomal arrays from recombinant core histones and nucleosome positioning DNA. J. Vis. Exp. 79, 50354 (2013).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Kato, H. et al. Architecture of the high mobility group nucleosomal protein 2-nucleosome complex as revealed by methyl-based NMR. Proc. Natl Acad. Sci. USA 108, 12283–12288 (2011).
Chalmers, M. J. et al. Probing protein ligand interactions by automated hydrogen/deuterium exchange mass spectrometry. Anal. Chem. 78, 1005–1014 (2006).
Keppel, T. R. & Weis, D. D. Mapping residual structure in intrinsically disordered proteins at residue resolution using millisecond hydrogen/deuterium exchange and residue averaging. J. Am. Soc. Mass Spectrom. 26, 547–554 (2015).
Pascal, B. D. et al. HDX workbench: software for the analysis of H/D exchange MS data. J. Am. Soc. Mass Spectrom. 23, 1512–1521 (2012).
Zhang, Z. & Smith, D. L. Determination of amide hydrogen exchange by mass spectrometry: a new tool for protein structure elucidation. Protein Sci. 2, 522–531 (1993).
Gamarra, N., Johnson, S. L., Trnka, M. J., Burlingame, A. L. & Narlikar, G. J. The nucleosomal acidic patch relieves auto-inhibition by the ISWI remodeler SNF2h. eLife 7, 34270 (2018).
Trnka, M. J., Baker, P. R., Robinson, P. J. J., Burlingame, A. L. & Chalkley, R. J. Matching cross-linked peptide spectra: only as good as the worse identification. Mol. Cell. Proteomics 13, 420–434 (2014).
Brown, P. H. & Schuck, P. Macromolecular size-and-shape distributions by sedimentation velocity analytical ultracentrifugation. Biophys. J. 90, 4651–4661 (2006).
Brautigam, C. A. Calculations and publication-quality illustrations for analytical ultracentrifugation data. Methods Enzymol. 562, 109–133 (2015).
Acknowledgements
We thank M. Rosen for sharing data before publication. We thank J. Tretyakova for help and training in histone purification; R. S. Isaac for providing initial nucleosomal array DNA; J. Pelton and the QB3 NMR Facility at the University of California Berkeley for help with collecting and processing NMR data; M. Keenen for providing Peg-silane-coated slides and protocols; C. Stoddard for providing ZMET2; S. Catania and H. Madhani for guidance in S. pombe experiments; D. Canzio for help with AUC; the Nikon Imaging Center at UCSF and K. Herrington for training and help with microscope data acquisition. We thank L. Hsieh, N. Gamarra, D. Canzio, A. Larson, E. Nora and H. Madhani for comments on the manuscript and members of the Gross and Narlikar laboratories for discussion. This work was supported by grant NIH NCATS UL1 TR000004 to S.S., J.D.G. and G.J.N.; Sandler Family Foundation Program for Breakthrough Research Post-doctoral Fellowship to S.S.; NIH NIGMS R01 GM121962 to J.D.G.; NIH R01GM108455 and R35 GM127020 to G.J.N.; PBBR New Frontier Research Award to G.J.N.; Dr. Miriam and Sheldon G. Adelson Medical Research Foundation to A.L.B., NIH NIGMS P41GM103481 to A.L.B.; Instrumentation Grants- Qexactive Plus (Thermo): NIH S10D016229 to A.L.B., Orbitrap Fusion Lumos (Thermo): University of California, San Francisco (Program for Breakthrough Biomedical Research (PBBR).
Author information
Authors and Affiliations
Contributions
S.S., J.D.G. and G.J.N. identified and developed the core questions. S.S. performed the bulk of the experiments. V.D. performed HDX-MS experiments and exported the data with the help of B.D.P. M.J.T. performed and analysed XLMS. S.S. and M.J.T. processed HDX-MS raw data. R.W.T. helped with the processing and collection of NMR data. S.S., J.D.G. and G.J.N. wrote the manuscript with contributions from the other authors. G.J.N. and J.D.G. oversaw the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Swi6 interaction with chromatin.
a, Model for HP1-mediated heterochromatin assembly: the bridging model relies on the ability of HP1 to oligomerize across nucleosomes; the phase separation model relies on the ability of HP1 to form phase-separated assemblies that can sequester chromatin. The molecular link between these two models is unclear. b, Sedimentation velocity-analytical ultracentrifugation (SV-AUC) analysis on H3Kc9me3 dinucleosomes. The continuous sedimentation coefficient distribution (c(s)) is shown as a two-dimensional distribution. The measured masses using a continuous function c(s) distribution versus theoretically predicted masses are shown. c, SV-AUC on H3Kc9me3 dinucleosomes with Swi6. Data are as in b. Analysis is performed using a continuous function c(s) with a bimodal f/f0 distribution. The peak at s = 21.24 corresponds to the Swi6–dinucleosome complex. The peak at s = 4.5 corresponds to free Swi6 dimers, as previously shown4. d, e, Analysis of raw SV-AUC data for dinucleosome alone (d) and dinucleosome plus Swi6 complex (e) showing that: (i) the fit lines describe the boundary data well; (ii) the r.m.s.d. value is below 0.01; and (iii) the residual bitmap has very few diagonal features indicating the good quality of the fit. For b–d, measurements are representative of two independent experiments.
Extended Data Fig. 2 Swi6–nucleosome cross-linking analysis.
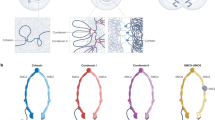
a, EDC cross-linking network between histones and Swi6. Histones tails and core regions are indicated, as well as the different Swi6 domains. b, Interactions between Swi6 domains, histone tails and core domains mapped by cross-link spectral counts. The number of cross-linked mass spectra matched to a given domain pair is indicated by the colour of the tile as well as the numbers given. c, d, Analysis as in a and b but performed only on the structured domains of Swi6 (CSD and chromodomain) and on the folded core of histones. e, The Swi6 CSD binds to H2B core. Superposition of 1H–15N HSQC spectra of Swi6 CSD with (black) and without (red) H2B peptide. Measurements are representative of three independent experiments. The sequence of the H2B peptide is shown. f, Chemical shift perturbation for assigned resonances between Swi6 CSD alone and with the addition of the H2B peptide shown in e. g, Intra-nucleosome cross-links are altered by Swi6 binding. Each axis maps residue position within a histone, and the red dashed lines indicate the boundaries between histone tail and core domains. A cross-link is represented by a filled circle between two residues, with area proportional to the number of MS2 spectra identifying a given cross-link. Cross-links found in the free nucleosome state are in green, and cross-links found in the Swi6-bound nucleosome are in blue. The red squares highlight changes in the H3–H3 and H4–H4 cross-linking patterns between the free and bound nucleosome states. h, New histone–histone cross-links detected only with Swi6 binding involve buried regions of the nucleosome. The plots report the solvent-accessible surface area (SASA; y axis) versus residues number (x axis) for each histone. Cross-linkable residues (E, D and K) are indicated by a circle. Empty circles represent residues that do not show any cross-linking, and filled circles represent cross-linked residues. The area of the circle reports the number of cross-linked spectra with one peptide mapped to a particular residue. The red ovals highlight buried residues that were observed to cross-link only in the presence of Swi6. Analysis in a–d, g and h is performed on one of two XLMS datasets. Similar results were obtained with the other dataset that used the BS3 cross-linker.
Extended Data Fig. 3 HDX-MS with nucleosome–Swi6 complex and nucleosomes alone.
a, Experimental scheme for HDX-MS. b, Changes in deuterium incorporation are reported for every histone at five different time points. Each horizontal bar represents an individual peptide, and peptides are placed beneath the schematic of secondary structure elements of the histones. Peptides are coloured as specified, showing the mean of deprotection derived from multiple peptides obtained from one of two independent experiments with similar results. Specifically, more than a 35% increase in deuterium incorporation compared to nucleosomes alone is observed in: (i) residues 74–90 of H3, corresponding to a portion of helix α1 and α2, and the connecting loop 1; (ii) residues 48–61 of H3, corresponding to helix αN of H3; and (iii) residues 85–102 of H4, corresponding to helix α3. Other regions with a marked increase in deuterium uptake are the C-terminal region and helix α2 of H2A (residues 113–129 and 40–51), the C-terminal of H2B (helix αC, residues 104–122), and H4 helix α2 and loop 2 (residues 50–60 and 72–84). These increases in the rates of deuterium incorporation caused by Swi6 binding indicate extensive changes in the backbone hydrogen bonding of histones within the nucleosome and partial unfolding of the helices in the histones. c, Kinetics of deuterium uptake of example histone peptides (residue numbers indicated) over the time course. Data are mean and s.d. of multiple peptides obtained from one of two independent experiments with similar results. Error bars not shown for points when shorter than the height of the symbol.
Extended Data Fig. 4 Nucleosome methyl-TROSY NMR.
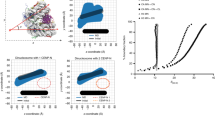
a, The methyl-TROSY spectra of H3 and H2B are well resolved and the chemical shift differences between the cross-peaks of our spectra and of the published spectra of Drosophila nucleosomes are small (less than 0.1 ppm)41. Owing to these small changes and the high protein conservation between the two species, we were able to transfer the assignments to all the cross-peaks of H3 and most of the cross-peaks of H2B. The chemical shift change (Δδ) plot reports the differences in chemical shift of the cross-peaks for conserved Ile, Leu and Val (ILV) residues in Drosophila and Xenopus histones. Δδ values were calculated from the equation:\(\sqrt{0.5({({\delta {\rm{H}}}_{Drosophila}-{\delta {\rm{H}}}_{Xenopus})}^{2}+{(0.2({\delta {\rm{C}}}_{Drosophila}-{\delta {\rm{C}}}_{Xenopus}))}^{2})}\) in which the factor of 0.2 is used as a scaling factor for the carbon spectral width. The average Δδ value is shown as a red dashed line. Experiments were performed twice to optimize conditions; data from one experiment are reported. b, Amino acid sequence alignment of Drosophila and Xenopus H2B and H3 histones. ILV residues conserved between the two species are coloured in blue, and non-conserved ILV residues are in red. Ile residues are fully conserved between H3 histones. Leu and Val are conserved between H2B histones, except for V15, V38 and V66, which are present only in the Xenopus histone. V15, V38 and V66 are not included in our NMR analysis. c, Quantification of the SASA of the residues indicated (x axis) in histone H2B (left) and H3 (right). The SASA was calculated from the PDB structure file 1KX5 using the program POPS. The SASA values are for the entire residue and represent fraction of exposed surface area. Bars corresponding to ILV residues are coloured in red with shading as specified, reflecting changes reported from the methyl-TROSY experiments due to Swi6 binding. ILV residues not assigned or not included in the analysis are in blue. The schematic of secondary structure elements is shown below the x axis.
Extended Data Fig. 5 Swi6 disorganizes the nucleosome core.
a, Mapping of the Swi6-induced changes in the nucleosome structure (PDB code 1KX5), as detected by XLMS (Fig. 1), HDX-MS (Fig. 2), and methyl-TROSY NMR (Fig. 3). H3 residues I51 and I62 are shown as spheres. The figure highlights how the three different techniques consistently reported alterations in the same regions of the octamer. b, In the nucleosome crystal structure, H2B is coloured in light orange, H2B peptide used in the CSD 1H–15N HSQC NMR experiment is in cyan and Leu and Val residues analysed in the nucleosome methyl-TROSY NMR are represented as spheres and coloured according to the extent of broadening determined by Ibound /Ifree. The magnified panel shows the location of the H2B peptide tested for binding to the CSD. c, Model depicting the effect of Swi6 on nucleosome conformations. Nucleosomes in solution can sample an ensemble of conformational states, of which the canonical conformation observed in the crystal structure is the most populated and lowest energy state. Swi6 increases nucleosome dynamics and promotes nucleosomes alternative states. Top, in the absence of Swi6, the equilibrium is pushed towards the nucleosome in the canonical state. Bottom, Swi6 increases nucleosome dynamics and promotes formation of a larger ensemble of nucleosomal conformations. d, Model of how disulfide cross-links in the nucleosome core prevent nucleosome core dynamics. e, This thermodynamic cycle integrates the different states of the nucleosome and nucleosome–Swi6 complexes implied by our work. In this thermodynamic cycle, the overall process involves a thermodynamic coupling between Swi6 binding and increased nucleosome dynamics. Path 1 shows how the intrinsic equilibrium between static and dynamic nucleosome states can be driven towards a dynamic nucleosomal state by Swi6 binding, and path 2 shows how binding by Swi6 to a static nucleosome can drive the equilibrium of the Swi6–nucleosome complex towards a more dynamic nucleosome state. Both paths lead to the same final state where the conformation of the Swi6-bound nucleosome is different from the one observed in the crystal structure (PDB code 1KX5). Importantly, both paths are thermodynamically equivalent.
Extended Data Fig. 6 Nucleosome octamer dynamics are important for Swi6 nucleosome binding.
a, Representative non-reducing SDS–PAGE showing nucleosome with H3–H4 histone disulfide-linked (H3•H4 S-S) or reduced (H3•H4 S-H). Around 90% of H3 and H4 are disulfide-linked in both the methylated and unmethylated nucleosomes. These are the cross-linking sites used in Fig. 4. For gel source data, see Supplementary Fig. 1. b, Nucleosome binding assays by fluorescence anisotropy showing reduced binding of Swi6 to H3K9me3 nucleosomes containing H3–H4 disulfide-linked octamer (H3•H4 S-S versus H3•H4 S-H). c, Fluorescence anisotropy measuring Swi6 binding to unmethylated nucleosome, H3•H4 S-H and S-S, showing that disulfide linkages have no effect on Swi6 binding to unmethylated nucleosome. d, Additional residues in the H2B and H4 mutated to Cys for generating dynamically restrained octamers are represented with spheres. e, Representative non-reducing SDS–PAGE showing nucleosome with H2B-H4 histone disulfide-linked (H2B•H4 S-S) or reduced (H2B•H4 S-H). Around 50% of H2B and H4 are disulfide-linked in both H3Kc9me3 and unmethylated nucleosomes. For gel source data, see Supplementary Fig. 1. f, Nucleosome binding assays by fluorescence anisotropy showing reduced binding of Swi6 to H2B-H4 disulfide-linked octamer (H2B•H4 S-S). g, Fluorescence anisotropy experiments showing comparable Swi6 binding to H3cK9me3 non-oxidized, H3cK9me3 oxidized and H2B•H4 S-H mononucleosomes. These results show that the oxidation process does not alter Swi6 binding to Cys-devoid nucleosomes and that the presence of reduced Cys does not affect Swi6 binding either. h, Fluorescence anisotropy measuring Swi6 binding to unmethylated nucleosome, H2B•H4 S-H and S-S, showing that disulfide linkages have no effect on Swi6 binding to unmethylated nucleosomes. i, Nucleosome binding assays by fluorescence anisotropy showing that Zmet2 binding to H2B–H4 disulfide-linked octamer (H2B•H4 S-S) is not significantly affected. FP, fluorescence polarization units. Measurements entailed at least three independent experiments and error bars reflect s.d.
Extended Data Fig. 7 Characterization of H3 enzymatic methylation by Dim5.
Methylation of H3 was followed by EST-based LC–MS/MS analysis of the intact proteins. a, Charge state envelope of untreated (left) and methylated (right) H3. The bottom panel, focused on a single charge state, shows disappearance of the starting material and the formation of higher mass species, spaced 14 Da apart. b, ETD-based mass spectrometry of precursor ions corresponding to unmethylated H3 (top), H3K9me3 (middle) and H3K9me3-K18me3 (bottom). The rationale for these assignments is based on the precursor mass values and by the product ions. The z-ions (purple) do not change between the three spectra, whereas the pattern of c-ions (blue) are mass shifted by 42 Da and charge-state shifted by 1+ at C9 between the top and middle or bottom panels, and again at C18 between the middle and bottom panels. These assignments are further validated by bottom-up proteomics analysis of Lys-C-digested samples (not shown). c, The precursor ion spectra in a were deconvoluted using Xtract, which models both the charge states and the isotope distributions. Deconvoluted MH+ values are consistent with multiple methylation states (owing to the difficulty of modelling isotope distributions from large proteins, particularly as there is some underlying oxidation, Xtract sometimes picks the wrong monoisotope). Deconvoluted intensities show that the enzymatically treated sample (right) contains no notable unmethylated H3 or mono- and di-methylated H3K9. Although 100% of analysed sample is tri-methylated at H3K9, additional methylations occur at H3K18 as noted above. The sample analysed here was used to carry out all the experiments to minimize variability.
Extended Data Fig. 8 Nucleosome disorganization and chromatin phase separation are specifically driven by Swi6.
a, Schematic of chromatin self-association assay. b, c, Restraining octamer dynamics do not affect the condensation of unmethylated chromatin in liquid droplets. Self-association assays in b are performed with unmethylated nucleosome arrays containing H3•H4 S-S (oxidized) and H3•H4 S-H (reduced) octamers as a function of increasing concentration of Swi6. In c, bright-field representative images of the array–Swi6 complex analysed in b show that the formation of phase-separated droplets is not affected by the disulfide linkages in absence of H3K9me3. d, Swi6 induces chromatin condensation in liquid droplets at physiological conditions—150 mM KCl, 30 °C, 2 μM Swi6. Bright-field representative images. e, Swi6 alone does not form phase-separated liquid droplets. Bright-field representative images of the Swi6 alone. f, Left, self-association assay performed with naked DNA (601 × 12, approximately 2 kb) in presence of increasing concentrations of Swi6. Right:,bright-field representative images of the DNA–Swi6 complex analysed in the left panel, showing the formation of phase-separated droplets. g, Zmet2 does not form phase-separated assemblies alone and it does not promote chromatin condensation in liquid droplets. Experiments are done at 40 nM nucleosome arrays, with 2 and 5 μM of Zmet2. Bright-field images of buffer only and Swi6–chromatin droplets are shown as controls. h, Titration of the denaturant guanidinium-HCl into nucleosome arrays does not promote formation of phase-separated droplets, which indicates that Swi6 specifically disorganizes the nucleosome core. i, Disulfide cross-links between H3 and H4 impair Mg2+-driven self-association of nucleosome arrays, indicating the role of nucleosome core dynamic for inter-nucleosome interactions. j, Left, Mg2+-driven self-association of unmethylated and H3K9cme3 arrays are comparable. Right, representative bright-field images of the H3K9cme3 array sedimentation analysed in the left panel, showing the formation of rounded phase-separated condensates at 2 and 3 mM Mg2+. The assay is performed in TE 0.1 (10 mM Tris pH 7.8, 0.1 mM EDTA) and 75 mM KCl. k, Nucleosome arrays quality controls, showing comparable octamer saturation and absence of over- and under-assemblies. Arrays are digested with HpaI, run on a native acrylamide gel, and DNA is stained. For source gel data, see Supplementary Fig. 1. All arrays used in this study conformed to the quality standard shown in k. In b–i and j, measurements and images entailed three independent experiments; error bars reflect s.d.
Extended Data Fig. 9 Swi6–chromatin assemblies present liquid-like properties.
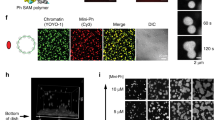
a, Chromatin–Swi6 condensates are reversible after dilution. b, Chromatin–Swi6 droplets fuse within seconds after incubation. The time course depicted is 30 s. c, Chromatin is included in Swi6-induced condensates. Nucleosome arrays are assembled with H2A–Cy3 or H2A–Cy5 and H3Kc9me3 octamers. Images showing Cy3- or Cy5-labelled chromatin mixed with Swi6 separately, before being incubated together. d, Images show condensates 30 min after mixing Cy3 and Cy5 pre-formed droplets. By around 1 h, most of the droplets contain homogeneous labelling with both dyes. Doubly labelled droplets indicate chromatin exchanges between the pre-formed condensates. e, Quantification of fluorescence intensity done in ImageJ using the Plot Profile function that creates a plot of intensity values across the yellow line shown in the left panel. Higher octamer concentration within the droplets as compare to the bulk of the solution is detected. Images are representative images from at least three independent experiments.
Extended Data Fig. 10 Swi6 dimerX and loopX mutants are defective in promoting octamer distortion and forming heterochromatin foci in S. pombe.
a, Nucleosome binding assays by fluorescence anisotropy showing that binding of Swi6 loopX mutant to H3K9cme3 nucleosomes is not affected by H2B–H4 histone disulfide linkage. H2B•H4 S-S are oxidized nucleosome, H2B•H4 S-H are reduced nucleosomes. Kd values are shown. b, Nucleosome binding assays by fluorescence anisotropy showing that binding of the Swi6 loopX and dimerX mutants to H3K9me3 nucleosomes is not affected by H3–H4 histone disulfide linkage. H3•H4 S-S are oxidized nucleosome, H3•H4 S-H are reduced nucleosomes. Kd values are shown. c, Wild-type Swi6 and loopX and dimerX mutant Swi6 proteins do not form liquid droplets in absence of chromatin (Swi6 concentration is 4 μM). d, Bright-field images showing droplet wetting over time. Swi6–chromatin condensates are incubated at room temperature and images are collected at the indicated times. Condensates are imaged on a bottom glass plate coated with Peg-silane (see Methods). e, Swi6 immunofluorescence on dcr1Δ swi6Δ S. pombe cells shows absence of Swi6 staining, confirming the specificity of the anti-Swi6 antibody. Images in c–e are representative of at least three independent experiments. In a and b, measurements entailed three independent experiments; error bars denote s.d.
Supplementary information
Supplementary Figure 1
This file contains gel source data. For Extended Data Figs 6a and 6e, Coomassie stainings of the non-reducing SDS gels show the formation of disulfide-linked H3-H4 (for Extended Data Fig. 6a) and H2B-H4 (for Extended Data Fig. 6e). For Extended Data Fig. 8k, the native acrylamide gel shows mono-nucleosomes generated upon HpaI-digestion of arrays, as arrays quality control.
Supplementary Tables
This file contains Supplementary Table 1 and 2. Supplementary Table 1: Swi6 affinity (Kd) for the indicated nucleosomes. Each value is a mean from at least three independent experiments; errors are standard deviation from the mean (SD). Supplementary Table 2: List of nucleosome mutants and corresponding amino acid substitution used in this study.
Rights and permissions
About this article
Cite this article
Sanulli, S., Trnka, M.J., Dharmarajan, V. et al. HP1 reshapes nucleosome core to promote phase separation of heterochromatin. Nature 575, 390–394 (2019). https://doi.org/10.1038/s41586-019-1669-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1669-2
This article is cited by
-
Phase separation-mediated biomolecular condensates and their relationship to tumor
Cell Communication and Signaling (2024)
-
Physiology and pharmacological targeting of phase separation
Journal of Biomedical Science (2024)
-
Liquid–liquid phase separation of H3K27me3 reader BP1 regulates transcriptional repression
Genome Biology (2024)
-
Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Nature Communications (2024)
-
Phosphorylation regulates tau’s phase separation behavior and interactions with chromatin
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.