Abstract
Translation initiation determines both the quantity and identity of the protein that is encoded in an mRNA by establishing the reading frame for protein synthesis. In eukaryotic cells, numerous translation initiation factors prepare ribosomes for polypeptide synthesis; however, the underlying dynamics of this process remain unclear1,2. A central question is how eukaryotic ribosomes transition from translation initiation to elongation. Here we use in vitro single-molecule fluorescence microscopy approaches in a purified yeast Saccharomyces cerevisiae translation system to monitor directly, in real time, the pathways of late translation initiation and the transition to elongation. This transition was slower in our eukaryotic system than that reported for Escherichia coli3,4,5. The slow entry to elongation was defined by a long residence time of eukaryotic initiation factor 5B (eIF5B) on the 80S ribosome after the joining of individual ribosomal subunits—a process that is catalysed by this universally conserved initiation factor. Inhibition of the GTPase activity of eIF5B after the joining of ribosomal subunits prevented the dissociation of eIF5B from the 80S complex, thereby preventing elongation. Our findings illustrate how the dissociation of eIF5B serves as a kinetic checkpoint for the transition from initiation to elongation, and how its release may be governed by a change in the conformation of the ribosome complex that triggers GTP hydrolysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Source data for figures are available as accompanying Source Data files. The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) server under the accession codes EMD-20324 (48S PIC), EMD-20325 (80S initiation complex) and EMD-20326 (80S elongation complex). Single-molecule fluorescence-trace data are available upon request.
Code availability
The MATLAB scripts used in this study are available at: https://drive.google.com/drive/u/1/folders/1-yKC0Nui7T7yoJScCuODPhCT4G52emLj
References
Hinnebusch, A. G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 83, 779–812 (2014).
Sokabe, M. & Fraser, C. S. Toward a kinetic understanding of eukaryotic translation. Cold Spring Harb. Perspect. Biol. 11, a032706 (2019).
Kaledhonkar, S. et al. Late steps in bacterial translation initiation visualized using time-resolved cryo-EM. Nature 570, 400–404 (2019).
Tsai, A. et al. Heterogeneous pathways and timing of factor departure during translation initiation. Nature 487, 390–393 (2012).
Goyal, A., Belardinelli, R., Maracci, C., Milón, P. & Rodnina, M. V. Directional transition from initiation to elongation in bacterial translation. Nucleic Acids Res. 43, 10700–10712 (2015).
Pestova, T. V. et al. The joining of ribosomal subunits in eukaryotes requires eIF5B. Nature 403, 332–335 (2000).
Lee, J. H. et al. Initiation factor eIF5B catalyzes second GTP-dependent step in eukaryotic translation initiation. Proc. Natl Acad. Sci. USA 99, 16689–16694 (2002).
Shin, B. S. et al. Uncoupling of initiation factor eIF5B/IF2 GTPase and translational activities by mutations that lower ribosome affinity. Cell 111, 1015–1025 (2002).
Acker, M. G. et al. Kinetic analysis of late steps of eukaryotic translation initiation. J. Mol. Biol. 385, 491–506 (2009).
Roll-Mecak, A., Cao, C., Dever, T. E. & Burley, S. K. X-ray structures of the universal translation initiation factor IF2/eIF5B: conformational changes on GDP and GTP binding. Cell 103, 781–792 (2000).
Kuhle, B. & Ficner, R. eIF5B employs a novel domain release mechanism to catalyze ribosomal subunit joining. EMBO J. 33, 1177–1191 (2014).
Fernández, I. S. et al. Molecular architecture of a eukaryotic translational initiation complex. Science 342, 1240585 (2013).
Acker, M. G., Kolitz, S. E., Mitchell, S. F., Nanda, J. S. & Lorsch, J. R. Reconstitution of yeast translation initiation. Methods Enzymol. 430, 111–145 (2007).
Chen, J. et al. High-throughput platform for real-time monitoring of biological processes by multicolor single-molecule fluorescence. Proc. Natl Acad. Sci. USA 111, 664–669 (2014).
Gutierrez, E. et al. eIF5A promotes translation of polyproline motifs. Mol. Cell 51, 35–45 (2013).
Llácer, J. L. et al. Conformational differences between open and closed states of the eukaryotic translation initiation complex. Mol. Cell 59, 399–412 (2015).
Hamilton, R., Watanabe, C. K. & de Boer, H. A. Compilation and comparison of the sequence context around the AUG startcodons in Saccharomyces cerevisiae mRNAs. Nucleic Acids Res. 15, 3581–3593 (1987).
Li, J. J., Chew, G. L. & Biggin, M. D. Quantitating translational control: mRNA abundance-dependent and independent contributions and the mRNA sequences that specify them. Nucleic Acids Res. 45, 11821–11836 (2017).
Walsh, R. M. & Martin, P. A. Growth of Saccharomyces cerevisiae and Saccharomyces uvarum in a temperature gradient incubator. J. Inst. Brew. 83, 169–172 (1977).
Lorsch, J. R. & Herschlag, D. Kinetic dissection of fundamental processes of eukaryotic translation initiation in vitro. EMBO J. 18, 6705–6717 (1999).
Acker, M. G., Shin, B. S., Dever, T. E. & Lorsch, J. R. Interaction between eukaryotic initiation factors 1A and 5B is required for efficient ribosomal subunit joining. J. Biol. Chem. 281, 8469–8475 (2006).
Pisareva, V. P. & Pisarev, A. V. eIF5 and eIF5B together stimulate 48S initiation complex formation during ribosomal scanning. Nucleic Acids Res. 42, 12052–12069 (2014).
Shah, P., Ding, Y., Niemczyk, M., Kudla, G. & Plotkin, J. B. Rate-limiting steps in yeast protein translation. Cell 153, 1589–1601 (2013).
Chu, D. et al. Translation elongation can control translation initiation on eukaryotic mRNAs. EMBO J. 33, 21–34 (2014).
Schuller, A. P., Wu, C. C. C., Dever, T. E., Buskirk, A. R. & Green, R. eIF5A functions globally in translation elongation and termination. Mol. Cell 66, 194–205 (2017).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Heyer, E. E. & Moore, M. J. Redefining the translational status of 80S monosomes. Cell 164, 757–769 (2016).
Ingolia, N. T. et al. Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep. 8, 1365–1379 (2014).
Juszkiewicz, S. et al. ZNF598 Is a quality control sensor of collided ribosomes. Mol. Cell 72, 469–481 (2018).
Ikeuchi, K. et al. Collided ribosomes form a unique structural interface to induce Hel2-driven quality control pathways. EMBO J. 38, e100276 (2019).
Yin, J., Lin, A. J., Golan, D. E. & Walsh, C. T. Site-specific protein labeling by Sfp phosphopantetheinyl transferase. Nat. Protoc. 1, 280–285 (2006).
Petrov, A., Grosely, R., Chen, J., O’Leary, S. E. & Puglisi, J. D. Multiple parallel pathways of translation initiation on the CrPV IRES. Mol. Cell 62, 92–103 (2016).
Bowen, A. M. et al. Ribosomal protein uS19 mutants reveal its role in coordinating ribosome structure and function. Translation 3, e1117703 (2015).
Ito, H., Fukuda, Y., Murata, K. & Kimura, A. Transformation of intact yeast cells treated with alkali cations. J. Bacteriol. 153, 163–168 (1983).
Maniatis, T., Fritsch, E. F. & Sambrook, J. Molecular Cloning: a Laboratory Manual (Cold Spring Harbor Laboratory Press, 1982).
O’Leary, S. E., Petrov, A., Chen, J. & Puglisi, J. D. Dynamic recognition of the mRNA cap by Saccharomyces cerevisiae eIF4E. Structure 21, 2197–2207 (2013).
Mitchell, S. F. et al. The 5′-7-methylguanosine cap on eukaryotic mRNAs serves both to stimulate canonical translation initiation and to block an alternative pathway. Mol. Cell 39, 950–962 (2010).
Algire, M. A. et al. Development and characterization of a reconstituted yeast translation initiation system. RNA 8, 382–397 (2002).
Goyer, C. et al. TIF4631 and TIF4632: two yeast genes encoding the high-molecular-weight subunits of the cap-binding protein complex (eukaryotic initiation factor 4F) contain an RNA recognition motif-like sequence and carry out an essential function. Mol. Cell. Biol. 13, 4860–4874 (1993).
Choi, S. K. et al. Physical and functional interaction between the eukaryotic orthologs of prokaryotic translation initiation factors IF1 and IF2. Mol. Cell. Biol. 20, 7183–7191 (2000).
Rinaldi, A. J., Suddala, K. C. & Walter, N. G. Native purification and labeling of RNA for single molecule fluorescence studies. Methods Mol. Biol. 1240, 63–95 (2015).
Marshall, R. A., Dorywalska, M. & Puglisi, J. D. Irreversible chemical steps control intersubunit dynamics during translation. Proc. Natl Acad. Sci. USA 105, 15364–15369 (2008).
Blanchard, S. C., Kim, H. D., Gonzalez, R. L. Jr, Puglisi, J. D. & Chu, S. tRNA dynamics on the ribosome during translation. Proc. Natl Acad. Sci. USA 101, 12893–12898 (2004).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Shin, B. S. et al. Structural integrity of α-helix H12 in translation initiation factor eIF5B is critical for 80S complex stability. RNA 17, 687–696 (2011).
Munro, J. B., Sanbonmatsu, K. Y., Spahn, C. M. T. & Blanchard, S. C. Navigating the ribosome’s metastable energy landscape. Trends Biochem. Sci. 34, 390–400 (2009).
Acknowledgements
We are grateful to J. Lorsch, T. Dever, J. Dinman, J. Yin and C. Aitken for sharing constructs, strains and protocols; C. Sitron and O. Brandman for sharing equipment and knowledge for use of their Freezer/Mill cryogenic mill for yeast lysis; the Stanford PAN facility for mass spectrometry analyses of proteins; and members of the Puglisi laboratory for discussion and input. This work was supported by the US National Institutes of Health (NIH) grants GM113078 and AI047365 to J.D.P.; a Knut and Alice Wallenberg Foundation postdoctoral scholarship (KAW 2015.0406) to J.W.; a National Science Foundation Graduate Research Fellowship (DGE-114747) to A.G.J.; a Damon Runyon Fellowship funded by the Damon Runyon Cancer Research Foundation (DRG-2321-18) to C.P.L.; a Stanford Bio-X fellowship to J.C.; and a Stanford Interdisciplinary Graduate Fellowship and NIH Molecular Biophysics Training Grant T32-GM008294 to A.P.
Author information
Authors and Affiliations
Contributions
J.W. performed all the biochemical and single-molecule experiments; J.W. analysed the data with help from J.C., A.P. and A.N.P.; J.W. prepared the samples, D.-H.C. and J.W. acquired and D.-H.C. processed the cryo-EM data; J.W. and J.D.P. conceived the project with input from A.G.J., C.P.L and A.N.P.; and J.W., A.G.J., C.P.L., J.C., D.-H.C. and J.D.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 A reconstituted yeast translation system with fluorescently labelled ribosomes for inter-subunit smFRET.
a, The labelling of the yeast 60S ribosomal subunit with Cy5 by means of a SNAP tag fused to the uL18 protein and the reaction with a SNAP-647 dye have previously been established32. b, In this work, we engineered a yeast strain in which all of the 40S subunits carried the N-terminal ybbR-tagged uS19 protein. After purification, the 40S was labelled by SFP synthase with CoA-547 at the serine residue (in bold and underlined) of the ybbR tag, resulting in the Cy3–40S. c, The estimated distance between the 2 ribosomal labelling sites is within 50 Å of that from published yeast 80S 3D structures (g). The ribosome model was created in PyMOL, using data from PDB 4V8Z12. d, Total internal reflection fluorescence microscopy (TIRFM) experimental setup to characterize the inter-subunit FRET signal. 80S complexes were assembled from Cy3–40S and Cy5–60S on the model mRNA (Fig. 1a) in the presence of the required factors (Methods), and were immobilized on a quartz slide used for TIRFM imaging with green laser illumination. e, f, Sample TIRFM experimental trace (e) and the inter-subunit smFRET efficiency histogram (f; fitted with a single-Gaussian distribution, with a mean FRET efficiency of 0.89 ± 0.15 s.e.m.); n = 107 molecules. g, Estimated distances between the two labelling sites on the ribosomal subunits from a few examples of published yeast 80S structures in different functional states, and the expected FRET efficiencies based on a Förster radius (R0) of 54 Å for the Cy3–Cy5 FRET pair. h, Representative SDS–PAGE analysis of the purified core eukaryotic initiation factors (blue numbering) and elongation factors (red numbering) that were used for the reconstitution of the translation system. Each component was analysed at least three times with similar results.
Extended Data Fig. 2 Native gel shift assays and cryo-EM analysis showing active translation initiation with our purified yeast translation system.
a, Representative gel showing initiation on the model mRNA. A merged view of Cy5 (red) and Cy3 (blue) scans of the same gel is shown. For gel source data, see Supplementary Fig. 1. The model mRNA was labelled with Cy5 and 40S was labelled with Cy3. Addition of Cy3–40S, Met-tRNAMeti:eIF2:GTP (Met-TC), eIF1 and eIF1A to the model mRNA–Cy5 resulted in the formation of a distinct 48S PIC band. Further addition of eIF5, eIF5B and 60S led to the formation of the 80S band. The experiment was repeated three times with similar results. b, Representative gel showing initiation on the cap-RPL41A mRNA. For gel source data, see Supplementary Fig. 1. The cap-RPL41A mRNA was labelled with Cy3, and other components were unlabelled. The gel was scanned for Cy3 fluorescence. Various mRNA and protein complexes were formed in the absence of 40S. After adding 40S to the mixture, a distinct 48S PIC band was formed and further addition of eIF5B and 60S shifted this band to the 80S band. The apparent electrophoretic mobility of the 48S PIC that formed with the cap-RPL41A mRNA differs from that of the 48S PIC that formed with the model mRNA; this is probably a result of the different charges or hydrodynamic radius of the complex brought about by the capped mRNA. Both 48S PIC and 80S formation were very inefficient when the cap-binding eIF4F (initiation factors 4A, 4E and 4G) and eIF4B proteins were omitted from the reaction, demonstrating the dependence of initiation on the cap when the full set of initiation factors were added. The experiment was repeated three times with similar results. c–e, Using our regime for assembly of the 48S PIC, we would expect the 48S PIC to be in the post-scanning state, with the initiation factors that are required during the scanning process potentially dissociated from the complex. A cryo-EM map at 9.9 Å resolution was obtained for the 48S PIC that formed on the cap-RPL30 mRNA (grey). This map was compared with previously reported 48S structures16 in the scanning-competent, mRNA-channel-open configuration (c) (EMD 3049; yellow), or scanning-incompetent, mRNA-channel-closed configuration (d) (EMD 3048; cyan) (Methods). The cap-RPL30–48S PIC was assembled in the same way as for our single-molecule experiments, and the comparisons showed that it resembles the post-scanning closed state, with the Met-tRNAMeti positioned in the P-site (e). The modelled Met-tRNAMeti in the EMD 3048 structure, PDB 3JAP, is shown in red.
Extended Data Fig. 3 Real-time 80S assembly on the model mRNA at the single-molecule level.
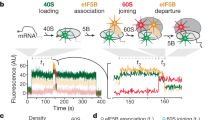
a, smFRET assay for subunit joining in ZMWs. 48S PICs were formed by incubating Cy3–40S, Met-TC, model mRNA–biotin, eIF1, eIF1A and eIF5 at 30 °C for 15 min before immobilization in the ZMWs. After washing away free components, the experiment was started with green laser illumination and delivery of Cy5–60S, eIF5 and eIF5B. The reaction was performed in the 1× Recon buffer supplemented with 1 mM GTP:Mg2+ at 20 °C. b, Example experimental trace showing real-time observation of Cy5–60S joining to immobilized Cy3–48S PIC to form the 80S complex, which is identified by the appearance of the smFRET signal. Single photobleaching events are denoted. Similar results were obtained from three independent experiments. c, The cumulative probability distribution of the dwell times for 60S joining was fitted to a double-exponential equation, resulting in a fast phase rate of about 0.22 s−1 with about 46% amplitude, and a slow phase rate of about 0.03 s−1 with about 54% amplitude (n = 178). The kinetics are comparable to previous bulk measurements of the same reaction under similar conditions9 (about 77% fast phase with a rate of about 0.076 s−1; about 23% slow phase with a rate of 0.019 s−1). d, e, Spermidine-driven initiation in the absence of eIF5B. The cumulative probability distributions of the dwell times for 60S joining (d) and the transition to elongation (e) from experiments performed in the presence or absence of 3 mM spermidine and/or 1 µM eIF5B at 20 °C with the model mRNA were fitted to a double-exponential (d) or a single-exponential (e) equation. The estimated average fast and slow phase rates (kfast and kslow) and amplitudes (Afast and Aslow) of 60S joining (from d) and Δt values (from e) with the 95% confidence intervals are shown in the inset table in e. n = 232 (for addition of spermidine and eIF5B), n = 117 (for addition of spermidine without eIF5B) and n = 164 (for addition of eIF5B without spermidine). Notably, the Δt value is small in the presence of only spermidine—probably owing to the lack of the rate-limiting eIF5B dissociation step. Consistently, this Δt value falls in the same range of the average tRNA arrival times after eIF5B departure in those experiments that were performed with labelled eIF5B (Fig. 2).
Extended Data Fig. 4 Single-molecule assays demonstrate that the association of the first aa-tRNA to the A site is codon- and tRNA-specific and can lead to further elongation.
a, Experimental setup (left) and sample fluorescence trace (right) for the single-molecule assay to assess the tRNA-binding specificity of the A site. 48S PICs that contain Cy3–40S, Met-tRNAMeti and the 3′-end-biotyinlated model mRNA (encoding Met-Phe-Lys-stop) were immobilized in ZMWs in the presence of the required initiation factors. Experiments were started by illuminating ZMWs with a green laser and delivering Cy5–60S, Cy3.5–Lys-tRNALys:eEF1A:GTP ternary complex (Cy3.5–Lys-TC) and initiation factors. No Cy3.5–Lys-tRNALys-binding events were observed in the 15-min imaging window, demonstrating that the aa-tRNA association with the A site was codon- and tRNA-specific (n = 150). b–e, The elongation competence of the 80S complex is scored by smFRET between tRNAs (that is, (Cy3.5)tRNAPhe–(Cy5)tRNALys) after initiation and elongation to the second elongation codon in the model mRNA. To study the tRNA–tRNA smFRET in the context of ribosomal inter-subunit smFRET, we decided to use Cy3 and Cy5.5 to label the 40S and 60S subunits, respectively. Therefore, we engineered the yeast 60S to carry a ybbR tag at the C terminus of uL18 and labelled the 60S with Cy5.5–CoA by SFP synthase (b). c, The different tag and label did not notably affect the kinetics of the transition to elongation (red curve, Δt = 95.4 ± 2.6 s (mean ± 95% confidence interval); n = 146; model mRNA, 20 °C and 3 mM free Mg2+) compared with the original Cy5–SNAP-tagged 60S (black curve, Δt = 92.2 ± 2.5 s (mean ± 95% confidence interval); n = 164). d, Sample fluorescence trace from the experiments in which Cy5.5–60S, Cy3.5–Phe-TC, Cy5–Lys-TC, eEF2, eEF3:ATP, eIF5A and other required initiation factors were delivered to immobilized 48S PICs containing Cy3–40S, Met-tRNAMeti and the 3′-end-biotyinlated model mRNA, and illuminated with a green laser at 20 °C. Out of n = 152 molecules that showed the sequential 60S and Cy3.5–Phe-tRNA association events, n = 113 molecules showed the subsequent Cy3.5–tRNAPhe to Cy5–tRNALys FRET signal. e, The distribution of the dwell times between the appearance of the Cy3.5 and Cy5 signals was fitted to a single-exponential equation, and the average time was 142 ± 8 s (mean ± 95% confidence interval; n = 113). Thus, 74% of the first aa-tRNA associations with the A site yielded the elongation to the next codon.
Extended Data Fig. 5 Comparisons of the kinetics of the transition from initiation to elongation under various conditions.
a–c, Transition dwell times (open circles) were fitted to single-exponential distributions to estimate the average dwell times (that is, Δt values) (grey bars; error bars in black represent the 95% confidence intervals). a, b, Comparison of Δt values in experiments performed with the model mRNA at 20 °C when no extra factors were added (data taken from Fig. 1d; n = 164), or in the presence of eIF3 and eEF3 (n = 130) or with addition of eIF5A (n = 143) (a); or when the concentration of Cy3.5–Phe-TC was 50 nM (n = 221) or 100 nM (data taken from Fig. 1d; n = 164) (b). c, Similar comparison for experiments performed with the cap-RPL30 mRNA at 20 °C when the concentration of Cy3.5–Phe-TC was 100 nM (n = 118) or 200 nM (n = 132). d, The Δt values compared across all assayed mRNAs at 20 °C and 30 °C (Fig. 1a, d). In the model mRNA-Kozak_-3U, the A at the −3 position of the optimal Kozak sequence was mutated to U, which largely abolished the Kozak sequence effect. From bottom to top for each group: n = 118, n = 130, n = 130, n = 121, n = 149, n = 161, n = 189, n = 159, n = 164 and n = 136.
Extended Data Fig. 6 Neither truncation nor fluorescent labelling of eIF5B perturbs its function.
a, Fluorescent labelling of eIF5B with Cy5.5 by means of a ybbR tag at the N-terminal end, which is distal from the ribosomal subunit labels and hence not expected to interfere with the inter-subunit smFRET. The ribosome model was created in PyMOL, using data from PDB 4V8Z12. b, A N-terminally truncated version of eIF5B (eIF5B-Trunc) was used in most of our assays, as in other reported reconstituted, purified yeast translation assays8,9,11,12,13,14,15,21,25. Previous reports failed to purify the full-length protein and have demonstrated that the truncated protein supported initiation in vitro and in vivo8,9. c, We tagged eIF5B-Trunc at the N terminus with a ybbR tag and labelled the protein with a CoA–Cy5.5 dye using SFP synthase. A representative gel is shown, which was first scanned for Cy5.5 fluorescence (right) and subsequently stained with Coomassie blue (left) following SDS–PAGE analysis of ybbR–eIF5B-Trunc after labelling with and without SFP synthase. The experiment was repeated three times with similar results. d, The GTPase activity of Cy5.5–eIF5B-Trunc was not perturbed by the labelling. Multiple-turnover GTP hydrolysis was performed in 50 mM HEPES-KOH pH 7.5, 10 mM Mg(OAc)2 and 100 mM KOAc at 30 °C 30 min before quenching with malachite green assay solution. Where applicable, concentrations were: GTP, 100 µM; eIF5B-Trunc, 2.5 µM; Cy5.5–eIF5B-Trunc, 2.5 µM; 40S and 60S, 0.2 µM each. The GTP-only group was used as a negative control and the values were normalized to 0. Bars represent mean, and error bars indicate standard deviations of three biological replicates (individual data points are indicated with open circles). e, The dwell times (open circles) between 60S arrival and the arrival of Phe-TC to the A site were fitted to single-exponential distributions for experiments performed with Cy5.5–eIF5B (related to Fig. 2c) versus those with unlabelled eIF5B (related to Fig. 1d), and at 20 °C or 30 °C. Error bars (in black) represent the 95% confidence intervals of the average dwell times (Δt values). From bottom to top for each group: n = 141, n = 159, n = 164, n = 118, n = 133, n = 130, n = 131, n = 189, n = 134 and n = 164. f, Despite it being reported previously that purification of recombinant yeast full-length eIF5B (eIF5B-FL) cannot be achieved49, we were able to recombinantly express and purify it, as shown by a 12% SDS–PAGE gel analysis. The experiment was repeated three times with similar results. g, Use of full-length eIF5B in our assay did not lead to faster transition to elongation in experiments performed with the cap-RPL30 mRNA at 3 mM free Mg2+ and 20 °C. Error bars (in black) represent the 95% confidence intervals of the average dwell times (Δt values, grey bars) from fitting of the dwell times (open circles) to single-exponential distributions. n = 118 (related to Fig. 1d) (left) and n = 205 (right).
Extended Data Fig. 7 Use of a non-hydrolysable GTP analogue, GDPNP, traps eIF5B on the 80S and prevents the transition to elongation.
a, Sample trace from experiments performed with GTP (n = 134). The grey highlighted part of the trace is shown in Fig. 2a as a magnified view. b, Sample trace from experiments performed with non-hydrolysable GDPNP (n = 105), in which eIF5B was trapped on the 80S after 60S joining and no tRNA binding to the A site was observed. Black dashed boxes show Cy5.5 events that are transient eIF5B sampling events to the 48S PIC before 60S joining. c, The cumulative probability distributions of the lifetimes of Cy5.5–eIF5B on 80S in experiments performed with the model mRNA at 3 mM Mg2+ and 20 °C in the presence of GTP (n = 134; related to Fig. 2d) or GDPNP (n = 105). In the presence of GTP, the mean lifetime was 51.9 s ( ± 1.8 s, 95% confidence interval from fitting to a single-exponential equation); in the presence of GDPNP, the mean lifetime was 848 s ( ± 19 s, 95% confidence interval from fitting to a linear equation).
Extended Data Fig. 8 The effects of eIF5B mutations on the kinetics of 60S joining and the transition to elongation.
a, b, The cumulative probability distributions of the dwell times for 60S joining (a) and the transition to elongation (b) from experiments performed with unlabelled wild-type eIF5B (n = 164), the eIF5B(T439A) mutant (n = 131) or the eIF5B(H505Y) mutant (n = 119) and the model mRNA at 20 °C in the presence of 3 mM Mg2+ and 1 mM GTP. c, The estimated average fast and slow phase rates (kfast and kslow) and amplitudes (Afast and Aslow) of 60S joining (from a) and Δt values (from b) with the 95% confidence intervals from fitting the distributions to a double-exponential (wild-type eIF5B and eIF5B(H505Y)) or a single-exponential (eIF5B(T439A)) equation for the 60S joining; or to a single-exponential equation for the transition dwell times. ∗In the case of eIF5B(T439A), out of n = 131 molecules showing 60S joining signal, only n = 17 molecules showed the subsequent signal of aa-tRNA association with the A site. Thus, the eIF5B(T439A) mutant is defective in catalysing subunit joining (consistent with bulk measurement9), and inhibits tRNA association with the A site.
Extended Data Fig. 9 Free Mg2+ concentration modulates 60S joining and the transition to elongation.
a, The dwell times for the transition from initiation to elongation (open circles) were fitted to single-exponential distributions to estimate the average transition dwell times (Δt values, with 95% confidence intervals, red) from experiments performed with unlabelled eIF5B and cap-RPL30 mRNA or model mRNA-Kozak at 20 °C in the presence of 1–10 mM Mg2+ (data for 3 mM Mg2+ were taken from Fig. 1d). For cap-RPL30, unstable 80S formation was observed at 1 mM Mg2+ (f) and thus no Δt values were obtained. From bottom to top for each group: n = 108, n = 189, n = 124, n = 150, n = 144, n = 118 and n = 195. b, The dwell times for the transition from initiation to elongation (open circles) were fitted to single-exponential distributions to estimate the average transition dwell times (Δt values, with 95% confidence intervals, red) from experiments performed with unlabelled eIF5B and cap-RPL30 or model mRNA-Kozak, under the indicated conditions. From left to right for each group: n = 195, n = 152, n = 118, n = 130, n = 150, n = 132, n = 189 and n = 159. c, The dwell times (open circles) for the transition from initiation to elongation, eIF5B lifetimes on 80S, and tRNA arrival times after eIF5B departure were fitted to single-exponential distributions to estimate the average dwell times (with 95% confidence intervals, black). Experiments were performed with cap-RPL30 mRNA at 3 mM free Mg2+ and 20 °C in the presence of unlabelled eIF5B (n = 195; data taken from Fig. 1d) or Cy5.5–eIF5B (n = 164; data taken from Fig. 2c); or at 2 mM free Mg2+ and 30 °C in the presence of unlabelled eIF5B (n = 152; data taken from b) or Cy5.5–eIF5B (n = 150). d, The dwell times for 60S joining (open circles) from the same experiments as in a were fitted to single-exponential distributions (for model mRNA-Kozak) or double- exponential distributions (for cap-RPL30; the average times for the fast phase were plotted here) to estimate the average dwell time (with 95% confidence intervals, red). e, The cumulative probability distribution of the 60S joining times, showing the compromised rate of 60S joining in experiments that were performed with the model mRNA-Kozak at 1 mM Mg2+ and 20 °C as in a (n = 150). However, we still observed that the arrival of tRNA to the A site occurred readily after 80S formation. The kinetics of 60S joining under these reaction conditions were not fitted well by single- or double-exponential distributions and therefore no average time was deduced for the bar plot in d. f, Sample trace from experiments performed with cap-RPL30 mRNA at 1 mM Mg2+ and 20 °C as described in a (n = 200). g, h, Dot plots for Fig. 2c, e, showing the single-exponential distributions of the dwell times, with the average dwell times (with 95% confidence interval) in black. Notes: (1) increasing free Mg2+ concentrations increased the Δt values (that is, higher Mg2+ concentration favours the eIF5B-bound semi-rotated 80S conformation (Extended Data Fig. 10)). This seems contrary to the known effects of free Mg2+ concentrations on the conformation of the bacterial pre-translocation 70S elongation complex50: lower Mg2+ concentration favours the rotated state and higher Mg2+ concentration favours the non-rotated state. The apparent discrepancy might be a result of the different compositions of the complexes: in our case, the semi-rotated state of the 80S contains an acylated Met-tRNAMeti in the P site and a protein factor eIF5B bound in the A site; whereas in the pre-translocation 70S complex, the P-site tRNA is deacylated and a peptidyl-tRNA is bound in the A site. The differences in the energy landscape between these two types of complexes can also be explained by the observation that there are no apparent fluctuations between semi-rotated and non-rotated 80S conformations with eIF5B bound, whereas there are frequent fluctuations between rotated and non-rotated 70S conformations in the pre-translocation complexes. (2) The different magnitudes of dependence of Δt values on temperature (Fig. 1d) and Mg2+ concentration for different mRNAs suggest that there are mRNA sequence-context differences in the thermodynamics that govern the transition from initiation to elongation. Similarly, the minimal Mg2+ concentrations required for stable 80S assembly on different mRNAs are different.
Extended Data Fig. 10 Cryo-EM analyses of the on-pathway 80S complexes during initiation and the transition to elongation.
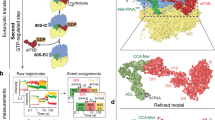
a, Cumulative probability distributions of the dwell times for 60S joining to 48S PIC (that is, the formation of the 80S complex, which is shown in black and designated as ‘total 80S’) and the subsequent association of aa-tRNA with the A site (corresponding to the Δt value, which is shown in blue and designated as ‘80S EC’) were from the single-molecule experiment with the model mRNA and wild-type eIF5B at 20 °C and 3 mM free Mg2+ (n = 164). The red curve corresponds to the evolution of the fraction of the eIF5B-bound 80S complex (‘80S IC’) over time, simulated with the kinetics of the other two curves. The dashed lines indicate that at time point 45 s, around 64.5% of the 80S complexes should be in the 80S initiation complex state under these conditions. b, The population sizes of the two classes of 80S complexes that were observed by cryo-EM (80S IC, 17,602 particles, 70%; 80S EC, 7,542 particles, 30%) were in agreement with predictions from our single-molecule kinetics at time point 45 s. c–f, The compositional and conformational differences between the 80S initiation and elongation complexes. g. Summary of cryo-EM data collection and processing for the 48S PIC (Extended Data Fig. 2c–e), 80S initiation complex (c) and 80S elongation complex (d) maps.
Supplementary information
Supplementary Information
This document contains the Supplementary Note and Supplementary Figure 1.
Rights and permissions
About this article
Cite this article
Wang, J., Johnson, A.G., Lapointe, C.P. et al. eIF5B gates the transition from translation initiation to elongation. Nature 573, 605–608 (2019). https://doi.org/10.1038/s41586-019-1561-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1561-0
This article is cited by
-
The molecular basis of translation initiation and its regulation in eukaryotes
Nature Reviews Molecular Cell Biology (2024)
-
Start codon-associated ribosomal frameshifting mediates nutrient stress adaptation
Nature Structural & Molecular Biology (2023)
-
Single-molecule visualization of mRNA circularization during translation
Experimental & Molecular Medicine (2023)
-
Principles, challenges, and advances in ribosome profiling: from bulk to low-input and single-cell analysis
Advanced Biotechnology (2023)
-
eIF5B and eIF1A reorient initiator tRNA to allow ribosomal subunit joining
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.