Abstract
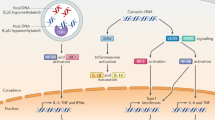
The accumulation of DNA in the cytosol serves as a key immunostimulatory signal associated with infections, cancer and genomic damage1,2. Cytosolic DNA triggers immune responses by activating the cyclic GMP–AMP synthase (cGAS)–stimulator of interferon genes (STING) pathway3. The binding of DNA to cGAS activates its enzymatic activity, leading to the synthesis of a second messenger, cyclic guanosine monophosphate–adenosine monophosphate (2′3′-cGAMP)4,5,6,7. This cyclic dinucleotide (CDN) activates STING8, which in turn activates the transcription factors interferon regulatory factor 3 (IRF3) and nuclear factor κ-light-chain-enhancer of activated B cells (NF-κB), promoting the transcription of genes encoding type I interferons and other cytokines and mediators that stimulate a broader immune response. Exogenous 2′3′-cGAMP produced by malignant cells9 and other CDNs, including those produced by bacteria10,11,12 and synthetic CDNs used in cancer immunotherapy13,14, must traverse the cell membrane to activate STING in target cells. How these charged CDNs pass through the lipid bilayer is unknown. Here we used a genome-wide CRISPR-interference screen to identify the reduced folate carrier SLC19A1, a folate–organic phosphate antiporter, as the major transporter of CDNs. Depleting SLC19A1 in human cells inhibits CDN uptake and functional responses, and overexpressing SLC19A1 increases both uptake and functional responses. In human cell lines and primary cells ex vivo, CDN uptake is inhibited by folates as well as two medications approved for treatment of inflammatory diseases, sulfasalazine and the antifolate methotrexate. The identification of SLC19A1 as the major transporter of CDNs into cells has implications for the immunotherapeutic treatment of cancer13, host responsiveness to CDN-producing pathogenic microorganisms11 and—potentially—for some inflammatory diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
The CRISPRi screen sequences were analysed using the Python-based ScreenProcessing pipeline. This custom code is available at https://github.com/mhorlbeck/ScreenProcessing.
Data availability
Raw sequencing data from the CRISPRi screens are available at NCBI Gene Expression Omnibus under accession number GSE134371.
Change history
07 March 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Ishii, K. J. et al. A Toll-like receptor-independent antiviral response induced by double-stranded B-form DNA. Nat. Immunol. 7, 40–48 (2006).
Stetson, D. B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Li, T. & Chen, Z. J. The cGAS–cGAMP–STING pathway connects DNA damage to inflammation, senescence, and cancer. J. Exp. Med. 215, 1287–1299 (2018).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP–AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP–AMP synthase. Cell 153, 1094–1107 (2013).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Diner, E. J. et al. The innate immune DNA sensor cGAS produces a noncanonical cyclic dinucleotide that activates human STING. Cell Rep. 3, 1355–1361 (2013).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Marcus, A. et al. Tumor-derived cGAMP triggers a STING-mediated interferon response in non-tumor cells to activate the NK cell response. Immunity 49, 754–763 (2018).
Woodward, J. J., Iavarone, A. T. & Portnoy, D. A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705 (2010).
McWhirter, S. M. et al. A host type I interferon response is induced by cytosolic sensing of the bacterial second messenger cyclic-di-GMP. J. Exp. Med. 206, 1899–1911 (2009).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011).
Corrales, L. et al. Direct activation of STING in the tumor microenvironment leads to potent and systemic tumor regression and immunity. Cell Rep. 11, 1018–1030 (2015).
Corrales, L., McWhirter, S. M., Dubensky, T. W., Jr & Gajewski, T. F. The host STING pathway at the interface of cancer and immunity. J. Clin. Invest. 126, 2404–2411 (2016).
Hou, Z. & Matherly, L. H. Biology of the major facilitative folate transporters SLC19A1 and SLC46A1. Curr. Top. Membr. 73, 175–204 (2014).
Zhao, R., Diop-Bove, N., Visentin, M. & Goldman, I. D. Mechanisms of membrane transport of folates into cells and across epithelia. Annu. Rev. Nutr. 31, 177–201 (2011).
Henderson, G. B. & Zevely, E. M. Structural requirements for anion substrates of the methotrexate transport system in L1210 cells. Arch. Biochem. Biophys. 221, 438–446 (1983).
Goldman, I. D. The characteristics of the membrane transport of amethopterin and the naturally occurring folates. Ann. NY Acad. Sci. 186, 400–422 (1971).
Yang, C. H., Sirotnak, F. M. & Dembo, M. Interaction between anions and the reduced folate/methotrexate transport system in L1210 cell plasma membrane vesicles: directional symmetry and anion specificity for differential mobility of loaded and unloaded carrier. J. Membr. Biol. 79, 285–292 (1984).
Goldman, I. D., Lichtenstein, N. S. & Oliverio, V. T. Carrier-mediated transport of the folic acid analogue, methotrexate, in the L1210 leukemia cell. J. Biol. Chem. 243, 5007–5017 (1968).
Lin, R., Heylbroeck, C., Genin, P., Pitha, P. M. & Hiscott, J. Essential role of interferon regulatory factor 3 in direct activation of RANTES chemokine transcription. Mol. Cell. Biol. 19, 959–966 (1999).
Brownell, J. et al. Direct, interferon-independent activation of the CXCL10 promoter by NF-κB and interferon regulatory factor 3 during hepatitis C virus infection. J. Virol. 88, 1582–1590 (2014).
Jansen, G. et al. Sulfasalazine is a potent inhibitor of the reduced folate carrier: implications for combination therapies with methotrexate in rheumatoid arthritis. Arthritis Rheum. 50, 2130–2139 (2004).
Kato, K. et al. Structural insights into cGAMP degradation by ecto-nucleotide pyrophosphatase phosphodiesterase 1. Nat. Commun. 9, 4424 (2018).
Goldman, I. D. A model system for the study of heteroexchange diffusion: methotrexate–folate interactions in L1210 leukemia and Ehrlich ascites tumor cells. Biochim. Biophys. Acta 233, 624–634 (1971).
Henderson, G. B., Grzelakowska-Sztabert, B., Zevely, E. M. & Huennekens, F. M. Binding properties of the 5-methyltetrahydrofolate/methotrexate transport system in L1210 cells. Arch. Biochem. Biophys. 202, 144–149 (1980).
Plosker, G. L. & Croom, K. F. Sulfasalazine: a review of its use in the management of rheumatoid arthritis. Drugs 65, 1825–1849 (2005).
Kozuch, P. L. & Hanauer, S. B. Treatment of inflammatory bowel disease: a review of medical therapy. World J. Gastroenterol. 14, 354–377 (2008).
Rajitha, P., Biswas, R., Sabitha, M. & Jayakumar, R. Methotrexate in the treatment of psoriasis and rheumatoid arthritis: mechanistic insights, current issues and novel delivery approaches. Curr. Pharm. Des. 23, 3550–3566 (2017).
Ahn, J., Son, S., Oliveira, S. C. & Barber, G. N. STING-dependent signaling underlies IL-10 controlled inflammatory colitis. Cell Rep. 21, 3873–3884 (2017).
van de Weijer, M. L. et al. A high-coverage shRNA screen identifies TMEM129 as an E3 ligase involved in ER-associated protein degradation. Nat. Commun. 5, 3832 (2014).
Zhao, R. et al. Impact of the reduced folate carrier on the accumulation of active thiamin metabolites in murine leukemia cells. J. Biol. Chem. 276, 1114–1118 (2001).
Zhao, R., Gao, F. & Goldman, I. D. Reduced folate carrier transports thiamine monophosphate: an alternative route for thiamine delivery into mammalian cells. Am. J. Physiol. Cell Physiol. 282, C1512–C1517 (2002).
Visentin, M., Zhao, R. & Goldman, I. D. Augmentation of reduced folate carrier-mediated folate/antifolate transport through an antiport mechanism with 5-aminoimidazole-4-carboxamide riboside monophosphate. Mol. Pharmacol. 82, 209–216 (2012).
Henderson, G. B. & Zevely, E. M. Anion exchange mechanism for transport of methotrexate in L1210 cells. Biochem. Biophys. Res. Commun. 99, 163–169 (1981).
Hamblett, K. J. et al. SLC46A3 is required to transport catabolites of noncleavable antibody maytansine conjugates from the lysosome to the cytoplasm. Cancer Res. 75, 5329–5340 (2015).
van Diemen, F. R. et al. CRISPR/Cas9-mediated genome editing of herpesviruses limits productive and latent infections. PLoS Pathog. 12, e1005701 (2016).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife 5, 1–20 (2016).
Kampmann, M., Bassik, M. C. & Weissman, J. S. Functional genomics platform for pooled screening and generation of mammalian genetic interaction maps. Nat. Protoc. 9, 1825–1847 (2014).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Kampmann, M., Bassik, M. C. & Weissman, J. S. Integrated platform for genome-wide screening and construction of high-density genetic interaction maps in mammalian cells. Proc. Natl Acad. Sci. USA 110, E2317–E2326 (2013).
Sadlish, H., Williams, F. M. R. & Flintoff, W. F. Functional role of arginine 373 in substrate translocation by the reduced folate carrier. J. Biol. Chem. 277, 42105–42112 (2002).
Huynh, T. N. et al. An HD-domain phosphodiesterase mediates cooperative hydrolysis of c-di-AMP to affect bacterial growth and virulence. Proc. Natl Acad. Sci. USA 112, E747–E756 (2015).
Sureka, K. et al. The cyclic dinucleotide c-di-AMP is an allosteric regulator of metabolic enzyme function. Cell 158, 1389–1401 (2014).
Henderson, G. B. & Zevely, E. M. Affinity labeling of the 5-methyltetrahydrofolate/methotrexate transport protein of L1210 cells by treatment with an N-hydroxysuccinimide ester of [3H]methotrexate. J. Biol. Chem. 259, 4558–4562 (1984).
McFarland, A. P. et al. Sensing of bacterial cyclic dinucleotides by the oxidoreductase RECON promotes NF-κB activation and shapes a proinflammatory antibacterial state. Immunity 46, 433–445 (2017).
Acknowledgements
We thank L. Zhang and E. Seidel for laboratory and technical assistance, H. Nolla and A. Valeros for assistance with cell sorting, the UC Berkeley High Throughput Screening Facility for preparation of gRNA lentivirus, A. P. McFarland for assistance in the isolation of primary cells from mice, S. L. McDevitt for assistance with deep sequencing, J. An and K. Elkon for assistance in the collection and isolation of primary peripheral blood leukocytes from healthy, human volunteers, and Raulet laboratory members, R. Vance, M. DuPage, M. van Gent, J. Thorner and A. van Elsas for helpful discussions. R.D.L. is a Cancer Research Institute Irvington Fellow supported by the Cancer Research Institute. D.H.R. is supported by NIH grants R01-AI113041 and R01-CA093678. B.G.G. is supported by the IGI-AstraZeneca Postdoctoral Fellowship, J.J.W. is supported by the Pew Scholars Program in the Biomedical Sciences, the Lupus Research Alliance and NIH grant 1R21AI137758-01. S.A.Z. is supported by grants from the University of Washington/Fred Hutchinson Cancer Research Center Viral Pathogenesis Training Program (AI083203), the University of Washington Medical Scientist Training Program (GM007266) and the Seattle ARCS foundation. J.E.C. is supported by the National Institute of Health New Innovator Awards (DP2 HL141006), the Li Ka Shing Foundation and the Heritage Medical Research Institute. This work employed the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley, supported by NIH S10 Instrumentation Grants S10 OD018174, S10RR029668 and S10RR027303.
Author information
Authors and Affiliations
Contributions
R.D.L., S.A.Z. and N.E.G. performed and analysed the experiments; L.O., S.M.M. and G.E.K. assisted with the experiments; S.K.W. and B.G.G. analysed the deep-sequencing data and advised on the screen design; R.D.L., S.A.Z., B.G.G., J.E.C., J.J.W. and D.H.R. designed the experiments and R.D.L., S.A.Z., J.J.W. and D.H.R. prepared the manuscript. All authors critically read the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.H.R. is a co-founder of Dragonfly Therapeutics and served or serves on the scientific advisory boards of Dragonfly, Aduro Biotech, and Ignite Immunotherapy; he has a financial interest in all four companies and could benefit from commercialization of the results of this research. S.M.M. is, and G.E.K. was, an employee of Aduro Biotech.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Structures of CDNs used in this study and gating strategy for the genome-wide CRISPRi screens.
a, Structures of the CDNs used in this study. b, Representative gating strategy for flow-cytometry-based sorting of the CRISPRi library of reporter-expressing THP-1 cells stimulated with CDNs. Cells were gated on the basis of their forward scatter (FSC) and side scatter (SSC) using gate P1. The P1 cells were subsequently selected for co-expression of BFP (fluorescent marker for the CRISPRi gRNAs) and GFP (marker for the expression of the reporter construct) (gate P2). Gate P3 excluded cell doublets present among P2 cells. Gate P4 selected for the lowest 25% of cells with respect to tdTomato expression and gate P5 selected for the highest 25%. c, Representative flow cytometry dot plots showing tdTomato expression in unstimulated cells or in cells stimulated for 20 h with cells for 20 h with CDN (2′3′-RR CDA). Data are representative of n = 3 biological replicates.
Extended Data Fig. 2 Results of genome-wide CRISPRi screens for host factors crucial for CDN stimulation.
a, b, Volcano plots of the gRNA-targeted genes enriched or depleted in the tdTomato reporter-low versus reporter-high groups after stimulation with 2′3′-RR CDA (a), or 2′3′-cGAMP (b). Each panel represents the combined results of two independent screens. Calculations of phenotypes and Mann–Whitney P values were performed as described in Methods.
Extended Data Fig. 3 SLC19A1 is critical for CDN-induced gene expression.
a, b, mRNA expression levels of SLC19A1 (a) or IRF3 (b) in THP-1 cells expressing a CRISPRi vector and a control non-targeting gRNA or gRNAs targeting IRF3 or SLC19A1 (two gRNAs each). c, THP-1 cells described in a and b were exposed to [3H]-methotrexate. After 1 h, radioactivity (in counts per minute, cpm) in lysates of cell pellets was measured. Counts were normalized to protein concentrations in the lysate. d, THP-1 cells expressing a control gRNA or SLC19A1-targeting gRNA were exposed to indicated concentrations of 2′3′-RR CDA or 2′3′-cGAMP. After 20 h, the mean fluorescence intensity (MFI) of tdTomato was quantified by flow cytometry. e, THP-1 cells expressing the indicated CRISPRi gRNAs or non-targeting gRNA (control), were stimulated with IFN-β (100 ng ml−1). After 18–22 h, tdTomato expression was quantified as in Fig. 2a. f, U937 cells expressing the indicated CRISPRi gRNAs or non-targeting gRNA (control), were stimulated with 2′3′-RR CDA (1.67 μg ml−1), 2′3′-cGAMP (15 μg ml−1) or IFN-β (100 ng ml−1). After 18–22h, tdTomato expression was quantified as in Fig. 2a. g, h, Induction of CXCL10 mRNA (g) or CCL5 mRNA (h) in control (non-targeting gRNA) THP-1 cells or THP-1 cells expressing the indicated CRISPRi gRNAs after 5 h stimulation with 5 μg ml−1 2′3′-RR CDA. i, CXCL10 protein expression in the supernatant of indicated gRNA-expressing THP-1 cells after exposure to 2 μg ml−1 2′3′-RR CDA for 20 h. j, Various cell lines expressing a control vector or an SLC19A1 expression vector were stimulated with IFN-β (100 ng ml−1). After 18–22 h, tdTomato expression was quantified as in Fig. 2a. k, THP-1 cells were incubated with increasing concentrations of the non-competitive inhibitor sulfasalazine or DMSO as vehicle control, before stimulation with 2′3′-RR CDA (1.25 μg ml−1), 2′3′-cGAMP (15 μg ml−1) or IFN-β (100 ng ml−1). After 18–22 h, tdTomato expression was quantified as in Fig. 2a. The data were normalized to the DMSO controls. In a–c and e–f, mean of n = 2 biological replicates are shown. In d, g–k, mean ± s.e.m. of n = 3 biological replicates are shown. Statistical analyses were performed to compare each cell line to the control using a one-way ANOVA followed by post hoc Dunnett’s test (d, g–i) or two-way ANOVA followed by uncorrected Fisher’s least significant difference tests (j). ****P ≤ 0.0001; n.s., not significant.
Extended Data Fig. 4 SLC19A1 overexpression increases the response to CDNs in various cell lines.
Various cell lines expressing a control vector or an SLC19A1 expression vector (SLC19A1 OE) stimulated with 2′3′-cGAMP (10 μg ml−1) (e) or IFN-β (100 ng ml−1) (or 100 ng ml−1 mouse IFN-β in the case of RAW cells). After 20 h, reporter expression was quantified by flow cytometry. Representative flow plots of n = 3 biological replicates shown in Fig. 2f and Extended Data Fig. 3j.
Extended Data Fig. 5 Covalent inhibition of SLC19A1 by NHS–methotrexate blocks STING activation.
a, Schematic overview of CDN-induced phosphorylation (P) of STING and downstream effectors TBK1 and IRF3. b, THP-1 monocytes pre-treated with DMSO or NHS–methotrexate (5 µM) were treated with varying concentrations of 2′3′-cGAMP for 4 h, and the amounts of IFNB1 or ISG15 transcripts were measured by RT–qPCR. c, Semi-native PAGE and immunoblot analysis of STING dimerization and phosphorylation in DMSO and NHS–methotrexate (5 µM) pre-treated THP-1 monocytes stimulated with 100 µM 2′3′-cGAMP for 4 h. For gel source data, see Supplementary Fig. 1. d, DMSO and NHS–methotrexate (5 µM)-treated THP-1 monocytes were treated with 100 µM 2′3′-cGAMP in the presence and absence of digitonin (5 µg ml−1) for 4 h and the induction of IFNB1 mRNA was measured by RT–qPCR. e, f, DMSO and NHS–methotrexate (5 µM) pre-treated THP-1 monocytes (e) or K562 cells (f) were stimulated for 4 h with 100 µM 2′3′-cGAMP, or not, in the presence or absence of digitonin (5 µg ml), and the induction of OASL and ISG15 mRNA was measured by RT–qPCR. In b, d–f, data are mean of n = 2 technical replicates and are representative of 3 independent experiments with similar results. In c, data are representative of three independent experiments with similar results.
Extended Data Fig. 6 SLC19A1 is critical for CDN uptake in human cell lines and primary cells.
a, TLC analysis of [32P]ATP standard (STD) and enzymatically synthesized [32P]2′3′-cGAMP (2′3′-cGAMP was purified on STING resin). Unbound nucleotides flowed through the resin (STING FT). Following four washes, the bound [32P]2′3′-cGAMP was eluted over three fractions. b, DRaCALA binding analysis of [32P]2′3′-cGAMP to STING C-CTD in the presence of competing unlabelled nucleotides (200 μM). c, TLC analysis of [32P]ATP and enzymatically synthesized [32P]2′3′-cGAMP and [32P]-c-di-AMP. d, Binding titration of [32P]2′3′-cGAMP or [32P]c-di-AMP to mSTING CTD, determined with DRaCALA assays. Red lines represent the 95% confidence interval for the nonlinear regression. e, Time course of [32P]2′3′-cGAMP (left) or [32P]3′3′-CDA (right) uptake in THP-1 monocytes. f, TLC analysis (left) and STING-binding (DRaCALA) (right) of [32P]ATP standard, or [32P]2′3′-cGAMP recovered from supernatants of THP-1 monocytes at the indicated time points. g, h, Effect of cell culture medium pH on [32P]2′3′-cGAMP uptake in THP-1 monocytes (g) or U937 monocytes (h). i, Time course of [32P]2′3′-cGAMP uptake by CIR cells transduced with empty vector or SLC19A1 expression vector. j, mRNA expression levels of SLC19A1 (SLC.) in K562 cells expressing control shRNAs (sh1 and sh2) or an SLC19A1-targeting shRNA (sh9). k, mRNA expression levels of CXCL10 in K562 cells described in j, stimulated with 5 μg ml−1 2′3′-RR CDA (RR CDA) for 5 h. l, [3H]-methotrexate uptake in K562 cells described in j, 1 h after exposure to [3H]-methotrexate. m, Time course of [32P]2′3′-cGAMP uptake in K562 cells described in j. n, Time course of [32P]2′3′-cGAMP uptake in U937 monocytes in the presence or absence of 500 µM competing, unlabelled (anti-)folates and sulfasalazine. o, Time course of [32P]2′3′-cGAMP uptake in K562 cells in the presence or absence of 500 µM competing, unlabelled (anti-)folates or sulfasalazine. p, Competition uptake assay of [32P]2′3′-cGAMP uptake in THP-1 cells in the presence of varying concentrations of competing, unlabelled 5-me-THF (IC50 = 4.10 ± 0.16 nM), methotrexate (IC50 = 54.83 ± 5.08 nM), 2′3′-cGAMP (IC50 = 1.89 ± 0.11 µM), sulfasalazine ((IC50 = 2.06 ± 0.17 µM), and folic acid (IC50 = 4.79 ± 0.08 µM). q, Trans-stimulation of [32P]2′3′-cGAMP influx in THP-1 cells by 5-me-THF. Cells were preloaded with indicated concentrations of 5-me-THF for 30 min. Cells were washed and incubated with [32P]2′3′-cGAMP for one hour. r, Normalized [32P]2′3′-cGAMP uptake after 1 h in DMSO or NHS–methotrexate (5 µM)-treated human PBMCs from four healthy donors. In a, c, data are representative of three independent experiments with similar results. In b, data are mean of n = 2 technical replicates and are representative of 3 independent experiments. In d, f, data are mean of n = 2 technical replicates and are representative of 2 independent experiments. In e, m, data are mean ± s.d. of n = 3 technical replicates and are representative of 3 independent experiments. In g–i and n–r, data are means ± s.d. of n = 3 technical replicates and are representative of 2 independent experiments. In j, k, data are mean ± s.e.m. of n = 3 biologically independent experiments. In l, data are mean of n = 2 biologically independent experiments.
Extended Data Fig. 7 SLC19A1 expression or inhibition has no effect on CDN uptake and signalling in mouse cells.
a, mRNA expression levels of Slc19a1 in mouse L1210 cells expressing control shRNAs (sh1 and sh2) or Slc19a1-targeting shRNA (sh4 and sh6). b, mRNA expression levels of Cxcl10 in L1210 cells described in a stimulated with 5 μg ml−1 2′3′-RR CDA (RR CDA) for 5 h. c, [3H]-methotrexate uptake in L1210 cells described in a 1 h after exposure to [3H]-methotrexate. d, Time course of [32P]2′3′-cGAMP uptake in L1210 cells described in a. e, mRNA expression levels of Slc19a1 in mouse C1498 cells expressing control shRNAs (sh1 and sh2) or Slc19a1-targeting shRNA (sh6). f, mRNA expression levels of Cxcl10 in the C1498 cells described in e, stimulated with 5 μg ml−1 2′3′-RR CDA (CDN) for 5 h. g, [3H]-methotrexate uptake in C1498 cells described in e 1 h after exposure to [3H]-methotrexate. h, Time course of [32P]2′3′-cGAMP uptake in C1498 cells transduced with a non-targeting control shRNA or Slc19a1 shRNA. i, j, mRNA expression levels of Slc19a1 in mouse BMMs (i) or BMDCs (j) not transduced or transduced with control shRNAs (sh1 and 2) or an shRNA targeting Slc19a1. k, l, mRNA expression of the Cxcl10 in cells described in i, j stimulated with 5 μg ml−1 2′3′-RR CDA (CDN) for 5 h. m, Time course of [32P]2′3′-cGAMP uptake in primary mouse splenocytes in the presence and absence of 500 µM competing, unlabelled (anti-) folates and sulfasalazine. n, Time course of [32P]2′3′-cGAMP uptake in primary mouse splenocytes pretreated or not with NHS–methotrexate (5 µM). o, Time course of [32P]2′3′-cGAMP uptake in L1210 cells pretreated or not with NHS–methotrexate (5 µM). In a–c, e–g, j, l, data are mean of n = 2 biologically independent experiments. In d, h, i, k, n, o, data are mean ± s.d. of n = 3 technical replicates and are representative of 2 independent experiments. In m, data are mean of n = 2 technical replicates and are representative of 2 independent experiments. In time-course experiments (d, h, m–o), data are presented as counts per minute normalized to cell count.
Extended Data Fig. 8 2′3′-cGAMP binds to SLC19A1.
a, Left, SDS–PAGE analysis followed by Coomassie blue staining of His-tagged human SLC19A1 (purified on Ni-NTA) pull-downs with Sepharose beads coupled with 2′3′-cGAMP (+) or control Sepharose beads (−). Input is shown on the right. b, Western blots of the samples in a with SLC19A1 antibody. c, Pull-downs of SLC19A1 competed with CDNs. His-tagged SLC19A1 was incubated with no CDN or with the indicated competing CDNs (250 µM) before pull-downs with 2′3′-cGAMP-Sepharose, followed by SDS–PAGE and western blotting with SLC19A1 antibody. A pulldown with control Sepharose is shown for comparison. For gel source data, see Supplementary Fig. 1. d, SDS–PAGE analysis followed by Coomassie blue staining of pull-downs of mSTING CTD with 2′3′-cGAMP (+) or control (−) Sepharose. In all panels, data are representative of two independent experiments with similar results.
Extended Data Fig. 9 RNA sequencing data of STING and SLC19A1 mRNA expression in 934 human cancer cell lines available at the Cancer Cell Line Encyclopedia website.
Expression is presented as transcripts per kilobase million (TPM). Data are downloaded from the European Bioinformatics Institute Gene expression Atlas (https://www.ebi.ac.uk/gxa/home). The dataset included three of the cell lines we examined, as shown.
Extended Data Fig. 10 The effect of SLC46A1 and SLC46A3 expression on CDN-induced reporter activation.
a, b, Enforced expression of SLC46A1 and SLC46A3 affects the responses of THP-1 cells to CDNs. Control THP-1 cells (transduced with empty expression vector) or SLC46A1-transduced THP-1 cells (a), or control THP-1 cells or SLC46A3-transduced cells (b) were stimulated with 2′3′-RR CDA (1.25 μg ml−1), 2′3′-cGAMP (15 μg ml−1) or IFN-β (100 ng ml−1). tdTomato reporter expression was measured by flow cytometry 18–22 h after stimulation. c, d, SLC46A1 or SLC46A3 depletions had little or no effects on cellular responses to CDNs, and combining depletion of SLC46A1 or SLC46A3 with SLC19A1 depletion had no additional effect compared to SLC19A1 depletion alone. THP-1 cells were transduced with non-targeting control CRISPRi gRNAs or SLC19A1-targeting CRISPRi gRNA in combination with a second control CRISPRi gRNA or SLC46A1-targeting CRISPRi gRNA in (c) or SLC46A3-targeting gRNA in (d). Cells were stimulated with 2′3′-RR CDA (1.67 μg ml−1), 2′3′-cGAMP (10 μg ml−1) or IFN-β (100 ng ml−1). tdTomato reporter expression was measured by flow cytometry 18–22 h after stimulation. Combined data of three independent experiments. Statistical analysis was performed using unpaired two-tailed Student’s t-tests (a, b) or one-way ANOVA followed by post hoc Tukey’s test when comparing only the effects of depleting SLC46A1 (c) or SLC46A3 (d). Data are mean ± s.e.m. of n = 3 independent replicates.
Supplementary information
Supplementary Figure 1
Uncropped original source images related to Figures 3a, 3c, 3d, 4g, 4h, and Extended Data Figures 5c, and 8c.
Supplementary Table 1
Ranking of target genes based on the ratio between individual gRNAs present in the populations that were hyper-responsive (hyper) or hypo-responsive to CDN treatment of n=2 biological replicates.
Supplementary Table 2
Ranking of targeted genes present in the populations hyper-responsive (hyper) or hypo-responsive (hypo) to CDN treatment. RRA ranking is based on the score computed by the MaGeCK program, and phenotypes and p-value calculated by the ScreenProcessing pipeline of n=2 biological replicates (see Methods). (included as Excel file).
Supplementary Table 3
Guide RNAs (gRNAs) used in this study.
Supplementary Table 4
RT-qPCR primers used in this study.
Supplementary Table 5
Primers and conditions used for the CDN screens.
Source data
Rights and permissions
About this article
Cite this article
Luteijn, R.D., Zaver, S.A., Gowen, B.G. et al. SLC19A1 transports immunoreactive cyclic dinucleotides. Nature 573, 434–438 (2019). https://doi.org/10.1038/s41586-019-1553-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1553-0
This article is cited by
-
A new physiological medium uncovers biochemical and cellular alterations in Lesch-Nyhan disease fibroblasts
Molecular Medicine (2024)
-
Harnessing innate immune pathways for therapeutic advancement in cancer
Signal Transduction and Targeted Therapy (2024)
-
CD4+ T cells produce IFN-I to license cDC1s for induction of cytotoxic T-cell activity in human tumors
Cellular & Molecular Immunology (2024)
-
Targeting serine-glycine-one-carbon metabolism as a vulnerability in cancers
Biomarker Research (2023)
-
Targeting cGAS/STING signaling-mediated myeloid immune cell dysfunction in TIME
Journal of Biomedical Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.