Abstract
The αβ T cell receptor (TCR), in association with the CD3γε–CD3δε–CD3ζζ signalling hexamer, is the primary determinant of T cell development and activation, and of immune responses to foreign antigens. The mechanism of assembly of the TCR–CD3 complex remains unknown. Here we report a cryo-electron microscopy structure of human TCRαβ in complex with the CD3 hexamer at 3.7 Å resolution. The structure contains the complete extracellular domains and all the transmembrane helices of TCR–CD3. The octameric TCR–CD3 complex is assembled with 1:1:1:1 stoichiometry of TCRαβ:CD3γε:CD3δε:CD3ζζ. Assembly of the extracellular domains of TCR–CD3 is mediated by the constant domains and connecting peptides of TCRαβ that pack against CD3γε–CD3δε, forming a trimer-like structure proximal to the plasma membrane. The transmembrane segment of the CD3 complex adopts a barrel-like structure formed by interaction of the two transmembrane helices of CD3ζζ with those of CD3γε and CD3δε. Insertion of the transmembrane helices of TCRαβ into the barrel-like structure via both hydrophobic and ionic interactions results in transmembrane assembly of the TCR–CD3 complex. Together, our data reveal the structural basis for TCR–CD3 complex assembly, providing clues to TCR triggering and a foundation for rational design of immunotherapies that target the complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The atomic coordinates of TCR–CD3 have been deposited in the Protein Data Bank with the accession code 6JXR. The corresponding maps have been deposited in the Electron Microscopy Data Bank with the accession code EMD-9895. The datasets generated and analysed during the current study are available from the corresponding authors upon reasonable request.
Change history
01 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-03245-4
References
Germain, R. N. & Stefanová, I. The dynamics of T cell receptor signaling: complex orchestration and the key roles of tempo and cooperation. Annu. Rev. Immunol. 17, 467–522 (1999).
Davis, M. M. & Bjorkman, P. J. T-cell antigen receptor genes and T-cell recognition. Nature 334, 395–402 (1988).
Samelson, L. E. & Klausner, R. D. The T-cell antigen receptor. Structure and mechanism of activation. Ann. NY Acad. Sci. 540, 1–3 (1988).
Rossjohn, J. et al. T cell antigen receptor recognition of antigen-presenting molecules. Annu. Rev. Immunol. 33, 169–200 (2015).
Meuer, S. C. et al. Evidence for the T3-associated 90K heterodimer as the T-cell antigen receptor. Nature 303, 808–810 (1983).
Wucherpfennig, K. W., Gagnon, E., Call, M. J., Huseby, E. S. & Call, M. E. Structural biology of the T-cell receptor: insights into receptor assembly, ligand recognition, and initiation of signaling. Cold Spring Harb. Perspect. Biol. 2, a005140 (2010).
Gaud, G., Lesourne, R. & Love, P. E. Regulatory mechanisms in T cell receptor signalling. Nat. Rev. Immunol. 18, 485–497 (2018).
Rudolph, M. G., Stanfield, R. L. & Wilson, I. A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006).
Wang, J. H. & Reinherz, E. L. The structural basis of αβ T-lineage immune recognition: TCR docking topologies, mechanotransduction, and co-receptor function. Immunol. Rev. 250, 102–119 (2012).
Call, M. E., Pyrdol, J. & Wucherpfennig, K. W. Stoichiometry of the T-cell receptor–CD3 complex and key intermediates assembled in the endoplasmic reticulum. EMBO J. 23, 2348–2357 (2004).
Garboczi, D. N. et al. Structure of the complex between human T-cell receptor, viral peptide and HLA-A2. Nature 384, 134–141 (1996).
Bäckström, B. T. et al. A motif within the T cell receptor α chain constant region connecting peptide domain controls antigen responsiveness. Immunity 5, 437–447 (1996).
Wang, Y. et al. A conserved CXXC motif in CD3epsilon is critical for T cell development and TCR signaling. PLoS Biol. 7, e1000253 (2009).
Love, P. E. & Hayes, S. M. ITAM-mediated signaling by the T-cell antigen receptor. Cold Spring Harb. Perspect. Biol. 2, a002485 (2010).
Kane, L. P., Lin, J. & Weiss, A. Signal transduction by the TCR for antigen. Curr. Opin. Immunol. 12, 242–249 (2000).
van der Merwe, P. A. & Dushek, O. Mechanisms for T cell receptor triggering. Nat. Rev. Immunol. 11, 47–55 (2011).
Arnett, K. L., Harrison, S. C. & Wiley, D. C. Crystal structure of a human CD3-ε/δ dimer in complex with a UCHT1 single-chain antibody fragment. Proc. Natl Acad. Sci. USA 101, 16268–16273 (2004).
Kjer-Nielsen, L. et al. Crystal structure of the human T cell receptor CD3εγ heterodimer complexed to the therapeutic mAb OKT3. Proc. Natl Acad. Sci. USA 101, 7675–7680 (2004).
Le Nours, J. et al. Atypical natural killer T-cell receptor recognition of CD1d-lipid antigens. Nat. Commun. 7, 10570–10584 (2016).
Call, M. E. et al. The structure of the ζζ transmembrane dimer reveals features essential for its assembly with the T cell receptor. Cell 127, 355–368 (2006).
Borg, N. A. et al. The CDR3 regions of an immunodominant T cell receptor dictate the ‘energetic landscape’ of peptide–MHC recognition. Nat. Immunol. 6, 171–180 (2005).
Hogquist, K. A.OKT3 and H57-597: from discovery, to commercialization, to the clinic. J. Immunol. 197, 3429–3430 (2016).
Sadelain, M., Rivière, I. & Riddell, S. Therapeutic T cell engineering. Nature 545, 423–431 (2017).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Yan, C. et al. Structure of a yeast spliceosome at 3.6-angstrom resolution. Science 349, 1182–1191 (2015).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Beddoe, T. et al. Antigen ligation triggers a conformational change within the constant domain of the αβ T cell receptor. Immunity 30, 777–788 (2009).
Birnbaum, M. E. et al. Molecular architecture of the αβ T cell receptor–CD3 complex. Proc. Natl Acad. Sci. USA 111, 17576–17581 (2014).
Natarajan, A. et al. Structural model of the extracellular assembly of the TCR–CD3 complex. Cell Rep. 14, 2833–2845 (2016).
Call, M. E., Pyrdol, J., Wiedmann, M. & Wucherpfennig, K. W. The organizing principle in the formation of the T cell receptor–CD3 complex. Cell 111, 967–979 (2002).
Fernandes, R. A. et al. T cell receptors are structures capable of initiating signaling in the absence of large conformational rearrangements. J. Biol. Chem. 287, 13324–13335 (2012).
Baker, B. M., Gagnon, S. J., Biddison, W. E. & Wiley, D. C. Conversion of a T cell antagonist into an agonist by repairing a defect in the TCR/peptide/MHC interface: implications for TCR signaling. Immunity 13, 475–484 (2000).
Yin, Y., Wang, X. X. & Mariuzza, R. A. Crystal structure of a complete ternary complex of T-cell receptor, peptide–MHC, and CD4. Proc. Natl Acad. Sci. USA 109, 5405–5410 (2012).
Sasada, T. et al. Involvement of the TCR Cβ FG loop in thymic selection and T cell function. J. Exp. Med. 195, 1419–1431 (2002).
Natarajan, K. et al. An allosteric site in the T-cell receptor Cβ domain plays a critical signalling role. Nat. Commun. 8, 15260–15274 (2017).
Chakraborty, A. K. & Weiss, A. Insights into the initiation of TCR signaling. Nat. Immunol. 15, 798–807 (2014).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED Protein Analysis Workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Hovmöller, S., Zhou, T. & Ohlson, T. Conformations of amino acids in proteins. Acta Crystallogr. D 58, 768–776 (2002).
Acknowledgements
We thank the Core Facilities at School of Life Science and Technology of Harbin Institute of Technology and the Core Facilities of Peking University School of Life Sciences for assistance with negative-staining electron microscopy, and the cryo-EM platform of Peking University for help with cryo-EM data collection. The computation was supported by the High-performance Computing Platform of Peking University. We thank J. Chai for critical reading of the manuscript. This research was funded by the National Natural Science Foundation of China grant no. 31825008 and 31422014 to Z.H.; 31725007 and 31630087 to N.G.; 31800630 to Y.Z.; 31700655 to N.L. and the Ministry of Science and Technology of China (2016YFA0500700 to N.G.).
Author information
Authors and Affiliations
Contributions
D.D., J.L., B.Z., L.Z., S.X. and Y.W. prepared the protein samples for negative staining and cryo-EM. D.D., J.L. and L.Z. performed negative staining. L.Z. and N.L. performed cryo-EM data acquisition and data processing. Y.Z. and N.G. built and refined the model. N.G. oversaw the cryo-EM and contributed to the manuscript preparation. Z.H. directed the project, oversaw biochemistry studies, cryo-EM sample preparation and model building and wrote the manuscript with input and support from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Ellis L. Reinherz and Nikolaos G. Sgourakis for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 TCR–CD3 protein complex purification.
a, Shown here are the results of western blot analyses using antibodies against the subunits of TCR–CD3. The peak fractions from gel filtration were immunoblotted with the indicated antibodies. b, The native or cross-linked TCR–CD3 protein bands were separated by reducing SDS–PAGE and visualized by staining with Coomassie blue. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Binding affinities between conformation-specific antibodies and the native or glutaraldehyde-treated TCR–CD3 complex.
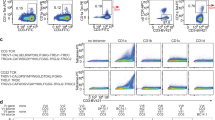
a–e, Binding affinities of native or glutaraldehyde-treated TCR–CD3 protein complex with activation antibodies OKT3 and UCHT1 and three antibodies targeting CD3γ, CD3δ and CD3ζ, measured by bio-layer interferometry. Green or blue curves are the experimental trace obtained from bio-layer interferometry experiments, and red curves are the best global fits to the data used to calculate the equilibrium-dissociation constant (Kd) values. Data are representative of three independent experiments. f, Negative staining of TCR–CD3 particles before (top) and after (bottom) cross-linking.
Extended Data Fig. 3 Cryo-EM image processing procedure.
a, A representative raw cryo-EM image. b, Two-dimensional class averages of the TCR–CD3 particles. c, Image-processing workflow of the TCR–CD3 particles. d, Gold-standard FSC curve of the final density map. e, Angular distribution of the TCR–CD3 particles in the final round of 3D refinement. f, Final local resolution estimation of the cryo-EM map.
Extended Data Fig. 4 Cryo-EM density map of the TCR-CD3 complex.
a–h, Local density of eight well-resolved TCR–CD3 subunits. The last four residues (D309–G312) in the cytoplasmic tail of TCRβ are not well defined by cryoEM density. In a–h, density maps are displayed with a similar threshold. i–n, Local density of all N-linked glycosylation regions in the final map. o, p, Local density of two representative extracellular regions of TCR–CD3 complex in the final map. q, r, Local density of two representative transmembrane regions of TCR–CD3 complex in the final map.
Extended Data Fig. 5 Structural comparison of free and pMHC-bound TCRαβ with that from the TCR–CD3 complex.
a, Structural comparison of free (PDB: 4X6B, green) and pMHC-bound (PDB: 4WWK, cyan) TCRαβ with that from TCR–CD3 complex. b, Structural comparison of the pMHC-bound TCRα and TCRβ chains (PDB: 3T0E) with cryo-EM structure of TCR–CD3. Per-residue backbone atom root mean square deviation (RMSD) values for the TCRα and TCRβ chains are listed.
Extended Data Fig. 6 Structure of CD3ζζ′.
a, The interface of the CD3ζζ′ homodimer. b, Structural comparison of the CD3ζζ′ homodimer (in TCR–CD3) with free CD3ζζ′ (PDB: 2HAC) shown in red.
Extended Data Fig. 8 Density maps of the conserved regions in the Cβ domain and interactions between the CP segments of Cα and Cβ domains.
a, Density map of the region near helix 3 (residues E153–K159) of the Cβ domain. b, Density map of the region near helix 4 (residues S216–N225) of the Cβ domain. c–f, Density map of the interaction sites between the CP segment of Cα and the H3 and H4 helices of the Cβ domain.
Supplementary information
Supplementary Figure
This file contains the uncropped gel blots for Extended Data Fig. 1a, b.
Rights and permissions
About this article
Cite this article
Dong, D., Zheng, L., Lin, J. et al. Structural basis of assembly of the human T cell receptor–CD3 complex. Nature 573, 546–552 (2019). https://doi.org/10.1038/s41586-019-1537-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1537-0
This article is cited by
-
Tumor heterogeneity and immune-evasive T follicular cell lymphoma phenotypes at single-cell resolution
Leukemia (2024)
-
Structure-guided engineering of immunotherapies targeting TRBC1 and TRBC2 in T cell malignancies
Nature Communications (2024)
-
The screening, identification, design and clinical application of tumor-specific neoantigens for TCR-T cells
Molecular Cancer (2023)
-
Catch bond models may explain how force amplifies TCR signaling and antigen discrimination
Nature Communications (2023)
-
Structural analysis of cancer-relevant TCR-CD3 and peptide-MHC complexes by cryoEM
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.