Abstract
Early human embryonic development involves extensive lineage diversification, cell-fate specification and tissue patterning1. Despite its basic and clinical importance, early human embryonic development remains relatively unexplained owing to interspecies divergence2,3 and limited accessibility to human embryo samples. Here we report that human pluripotent stem cells (hPSCs) in a microfluidic device recapitulate, in a highly controllable and scalable fashion, landmarks of the development of the epiblast and amniotic ectoderm parts of the conceptus, including lumenogenesis of the epiblast and the resultant pro-amniotic cavity, formation of a bipolar embryonic sac, and specification of primordial germ cells and primitive streak cells. We further show that amniotic ectoderm-like cells function as a signalling centre to trigger the onset of gastrulation-like events in hPSCs. Given its controllability and scalability, the microfluidic model provides a powerful experimental system to advance knowledge of human embryology and reproduction. This model could assist in the rational design of differentiation protocols of hPSCs for disease modelling and cell therapy, and in high-throughput drug and toxicity screens to prevent pregnancy failure and birth defects.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the article and its Supplementary Information files and from the corresponding author upon request. scRNA-seq data are also available at the Gene Expression Omnibus under accession no. GSE134571. All Source Data for graphs included in the paper are available in the online version of the paper.
Code availability
MATLAB and RStudio scripts used in this work are available from the corresponding author upon request.
References
O’Rahilly, R. & Müller, F. Developmental Stages in Human Embryos. (Carnegie Institution of Washington, 1987).
Rossant, J. Mouse and human blastocyst-derived stem cells: vive les differences. Development 142, 9–12 (2015).
Davidson, K. C., Mason, E. A. & Pera, M. F. The pluripotent state in mouse and human. Development 142, 3090–3099 (2015).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Daley, G. Q. et al. Setting global standards for stem cell research and clinical translation: the 2016 ISSCR guidelines. Stem Cell Reports 6, 787–797 (2016).
Hyun, I., Wilkerson, A. & Johnston, J. Embryology policy: revisit the 14-day rule. Nature 533, 169–171 (2016).
ten Berge, D. et al. Wnt signaling mediates self-organization and axis formation in embryoid bodies. Cell Stem Cell 3, 508–518 (2008).
van den Brink, S. C. et al. Symmetry breaking, germ layer specification and axial organisation in aggregates of mouse embryonic stem cells. Development 141, 4231–4242 (2014).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E. D. & Brivanlou, A. H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Harrison, S. E., Sozen, B., Christodoulou, N., Kyprianou, C. & Zernicka-Goetz, M. Assembly of embryonic and extra-embryonic stem cells to mimic embryogenesis in vitro. Science 356, eaal1810 (2017).
Shao, Y. et al. Self-organized amniogenesis by human pluripotent stem cells in a biomimetic implantation-like niche. Nat. Mater. 16, 419–425 (2017).
Xue, X. et al. Mechanics-guided embryonic patterning of neuroectoderm tissue from human pluripotent stem cells. Nat. Mater. 17, 633–641 (2018).
Beccari, L. et al. Multi-axial self-organization properties of mouse embryonic stem cells into gastruloids. Nature 562, 272–276 (2018).
Shao, Y. et al. A pluripotent stem cell-based model for post-implantation human amniotic sac development. Nat. Commun. 8, 208 (2017).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Rivron, N. et al. Debate ethics of embryo models from stem cells. Nature 564, 183–185 (2018).
Shahbazi, M. N. et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos. Nature 552, 239–243 (2017).
Taniguchi, K. et al. Lumen formation is an intrinsic property of isolated human pluripotent stem cells. Stem Cell Reports 5, 954–962 (2015).
Dobreva, M. P. et al. Periostin as a biomarker of the amniotic membrane. Stem Cells Int. 2012, 987185 (2012).
Sasaki, K. et al. The germ cell fate of cynomolgus monkeys is specified in the nascent amnion. Dev. Cell 39, 169–185 (2016).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
Bernardo, A. S. et al. BRACHYURY and CDX2 mediate BMP-induced differentiation of human and mouse pluripotent stem cells into embryonic and extraembryonic lineages. Cell Stem Cell 9, 144–155 (2011).
Kobayashi, T. et al. Principles of early human development and germ cell program from conserved model systems. Nature 546, 416–420 (2017).
Irie, N. et al. SOX17 is a critical specifier of human primordial germ cell fate. Cell 160, 253–268 (2015).
Sasaki, K. et al. Robust in vitro induction of human germ cell fate from pluripotent stem cells. Cell Stem Cell 17, 178–194 (2015).
Aramaki, S. et al. A mesodermal factor, T, specifies mouse germ cell fate by directly activating germline determinants. Dev. Cell 27, 516–529 (2013).
Mendjan, S. et al. NANOG and CDX2 pattern distinct subtypes of human mesoderm during exit from pluripotency. Cell Stem Cell 15, 310–325 (2014).
Gadue, P., Huber, T. L., Paddison, P. J. & Keller, G. M. Wnt and TGF-β signaling are required for the induction of an in vitro model of primitive streak formation using embryonic stem cells. Proc. Natl Acad. Sci. USA 103, 16806–16811 (2006).
Séguin, C. A., Draper, J. S., Nagy, A. & Rossant, J. Establishment of endoderm progenitors by SOX transcription factor expression in human embryonic stem cells. Cell Stem Cell 3, 182–195 (2008).
Villa-Diaz, L. G. et al. Synthetic polymer coatings for long-term growth of human embryonic stem cells. Nat. Biotechnol. 28, 581–583 (2010).
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 25, 681–686 (2007).
Zheng, Y. & Fu, J. Protocol for controlled modeling of human epiblast and amnion development using stem cells. Protoc. Exch. https://doi.org/10.21203/rs.2.11520/v1 (2019).
Lacoste, A., Berenshteyn, F. & Brivanlou, A. H. An efficient and reversible transposable system for gene delivery and lineage-specific differentiation in human embryonic stem cells. Cell Stem Cell 5, 332–342 (2009).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Acknowledgements
This work is supported by the University of Michigan Mechanical Engineering Faculty Support Fund (J.F.), the Michigan–Cambridge Research Initiative (J.F.), the University of Michigan Mcubed Fund (J.F.), the National Institutes of Health (R01 DK089933, D.L.G.) and the California Institute for Regenerative Medicine (RB5-07409, V.M.W.). Y.S. was partially supported by the University of Michigan Rackham Predoctoral Fellowship. The Lurie Nanofabrication Facility at the University of Michigan is acknowledged for support with microfabrication.
Author information
Authors and Affiliations
Contributions
Y.Z. and J.F. conceived and initiated the project; Y.Z. designed, performed and quantified most experiments, including scRNA-seq data analysis and interpretation; X.X. maintained cell culture, participated in experiments and independently repeated experiments; Y.S. helped to design experiments; S.W. developed the TCF/Lef:H2B-GFP reporter line and helped with scRNA-seq data analysis; Z.L. fabricated silicon moulds; S.N.E. helped with hPGCLC-related experiments; J.M.M., J.N.L. and V.M.W. provided the T-mNeonGreen reporter line; Y.Z., D.L.G. and J.F. wrote the manuscript; J.F. supervised the study. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
Y.Z., Y.S., S.N.E., D.L.G. and J.F. have filed a provisional patent related to this work (United States Provisional Patent Application No. 62/431,907).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Nicolas Rivron and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Microfluidic generation of pluripotent epiblast-like cyst.
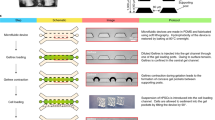
a, Photograph showing microfluidic devices in a six-well plate. Inset shows a top view of the device. Devices were filled with medium containing food colour dyes to illustrate medium reservoirs and microfluidic channels. b, Design of microfluidic device incorporating three parallel channels (80 µm in height) partitioned by trapezoid-shaped supporting posts spaced 80 µm apart. The central channel (gel channel) is preloaded with Geltrex. The other two open channels are used for cell loading (cell-loading channel) and chemical induction (induction channel), respectively. c, Protocol for generating epiblast-like cysts. After cell loading (designated as t = −18 h), a basal medium comprising Essential 6 medium and FGF2 (20 ng ml−1) is supplied to both the cell-loading and induction channels from t = 0 h onwards. d, Schematic showing cell loading, cell clustering and lumenogenesis. After injection of Geltrex into the gel channel, Geltrex gelation and contraction leads to formation of evenly spaced concave gel pockets between supporting posts. After loading human ES cells (hESC) into the cell-loading channel, the microfluidic device is tilted 90° to allow cells to settle into gel pockets (t = −18 h). At t = 0 h, nascent lumenal cavities emerge within cell clusters. At t = 36 h, epiblast-like cysts contain a single central lumenal cavity reminiscent of the pro-amniotic cavity formed in the epiblast of the blastocyst upon implantation. e, Representative bright-field images showing an array of epiblast-like cysts at t = 0 h and 36 h. Experiments were repeated five times with similar results. Scale bar, 80 µm. f, Representative confocal micrographs showing epiblast-like cysts at indicated time points stained for ezrin (top) or E-cadherin (E-cad) and laminin (Lam; bottom). Nuclei were stained with DAPI. Some epiblast-like cysts initiate lumenogenesis with multiple small ezrin+ lumenal cavities, which gradually resolve into a single central lumen, probably through cavity fusion. Scale bars, 40 µm. Experiments were repeated three times with similar results. g, Representative confocal micrographs showing X–Y, X–Z and Y–Z sections of epiblast-like cysts obtained at t = 36 h, stained for E-cadherin and ezrin. Nuclei were stained with DAPI. Scale bars, 40 µm. Experiments were repeated twice with similar results. h, Percentage of epiblast-like cysts with single, multiple or no lumenal cavities at indicated time points. n = 59, 57, 60, 89 and 110 cysts for t = 0 h, 6 h, 12 h, 24 h and 36 h, respectively. Data were pooled from n = 2 (0 h, 6 h, 12 h) or 3 (24 h and 36 h) independent experiments. Data represent the mean ± s.e.m. i, Cell number in each epiblast-like cyst as a function of time. n = 26 cysts for each time point. Data were pooled from n = 2 independent experiments. Red lines represent the median. j, Equivalent epiblast-like cyst diameter as a function of time. n = 56, 45, 50, 115 and 88 cysts for t = 0 h, 6 h, 12 h, 24 h and 36 h, respectively. Data were pooled from n = 3 independent experiments. Red lines represent the median. k, Embedded epiblast-like cyst perimeter percentage as a function of time. n = 33, 35, 26, 28 and 21 cysts for t = 0 h, 6 h, 12 h, 24 h and 36 h, respectively. Data were pooled from n = 2 independent experiments. Red lines represent the median. l, Representative confocal micrographs showing epiblast-like cysts at t = 36 h stained for ZO1 and actin (top) or GM130 and actin (bottom). Nuclei were stained with DAPI. Far right, magnified views of outlined regions. Scale bars, 40 µm. Experiments were repeated twice with similar results. m, Representative confocal micrographs showing epiblast-like cysts obtained at t = 36 h with indicated drugs supplemented into basal medium from t = 0–36 h. IWP2, 5 µM; BMP inhibitor LDN 193189 (LDN), 0.5 µM; TGF-β inhibitor SB 431542 (SB), 10 µM; Caspase 3 inhibitor Z-DEVD-FMK (Cas 3), 10 µM. Cysts were stained for NANOG and E-cadherin. Fluorescently labelled WGA was used to stain plasma membrane. Nuclei were stained with DAPI. Note that assays were also conducted with different concentrations of Z-DEVD-FMK (5 µM, 20 µM and 40 µM), with results compatible with those obtained from 10 µM Z-DEVD-FMK. Scale bars, 40 µm. Experiments were repeated twice with similar results. n, Equivalent epiblast-like cyst diameter at t = 36 h under indicated conditions. n = 142, 37, 44, 40 and 36 cysts for control, IWP2, LDN, SB and Cas 3 conditions, respectively. Data were pooled from n = 2 independent experiments. Red lines represent the median.
Extended Data Fig. 2 Progressive development of posteriorized embryonic-like sac.
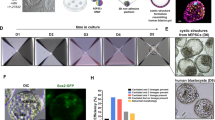
a, Protocol for generating P-ELS. After initial seeding and clustering of human ES cells (t = −18 to 0 h), lumenogenesis leads to the formation of the epiblast-like cyst containing a single central lumen. BMP4 stimulation (50 ng ml−1) from the induction channel from t = 0 h onwards leads to patterning of the epiblast-like cyst and formation of the asymmetric embryonic-like sac, with specification of the amniotic ectoderm-like fate to cells directly exposed to BMP4 induction. Development of the P-ELS triggers the onset of gastrulation-like events and expression of T in the epiblast-like compartment of the P-ELS. Bright-field image shows an array of P-ELS at t = 36 h. Scale bar, 80 µm. Experiments were repeated five times with similar results. b, Percentage of P-ELS and uniformly squamous amniotic ectoderm-like cysts at t = 36 h. Representative phase-contrast micrographs are shown. Scale bar, 40 µm. n = 396 cysts from n = 5 independent experiments. Each dot represents data obtained from an independent experiment. Red lines represent the median. c, Cell number in each cyst as a function of time. n = 26 cysts for each time point. Data were pooled from n = 2 independent experiments. Red lines represent the median. d, Representative confocal micrographs showing X–Y, X–Z and Y–Z sections of P-ELS at t = 36 h stained for E-cadherin and ezrin. Nuclei were stained with DAPI. Scale bars, 40 µm. Experiments were repeated twice with similar results. e, Representative confocal micrographs showing P-ELS at t = 36 h stained for pSMAD1/5 and TFAP2A. Nuclei were stained with DAPI. Scale bars, 40 µm. Experiments were repeated three times with similar results. f, Representative confocal micrographs showing P-ELS at indicated time points co-stained for NANOG and CDX2 (left) or T and E-cadherin (right). Nuclei were stained with DAPI. Scale bars, 40 µm. Experiments were repeated three times with similar results. g, Representative confocal micrographs showing amniotic ectoderm-like tissues at t = 72 h stained for E-cadherin, GATA3 and OCT4, TFAP2A and T, or TFAP2C and NANOG as indicated. Bottom images show magnified views of amniotic ectoderm-like tissues. Nuclei were stained with DAPI. Scale bars, 40 µm (small panels) and 160 µm (large panels). Experiments were repeated five times with similar results. h, Thickness of amniotic ectoderm-like tissue as a function of time. n = 267, 167, 258 and 111 cysts for t = 24 h, 30 h, 36 h and 72 h, respectively. Data were pooled from n = 5 independent experiments. Red lines represent the median. i, In situ hybridization images of P-ELS at t = 24 h and t = 36 h for BMP4 (left) and AXIN2 (right). Scale bars, 40 µm. Experiments were repeated twice with similar results. j, In situ hybridization images of P-ELS at t = 36 h for ACTB (top) and B. subtilis dapB (bottom). Scale bars, 40 µm. Experiments were repeated twice with similar results. k, Left, bright-field and fluorescent micrographs showing P-ELS at t = 36 h blocking diffusion of fluorescein-labelled dextran (70 kDa) into the cell loading channel. Dextran was supplemented into the induction channel and diffused into the gel channel. Experiments were repeated twice with similar results. Scale bar, 40 µm. Right, plot showing relative fluorescence intensity within the cell-loading channel and induction channel. Data are normalized to the average intensity in the cell loading channel. Our characterization of dextran diffusion shows that the supporting posts and in-between expanding P-ELS effectively block morphogen diffusion across the tissues. n = 10 cysts. Data were pooled from n = 2 independent experiments. Red lines represent the median.
Extended Data Fig. 3 Progressive development of anteriorized embryonic-like sac.
a, Protocol for generating A-ELS. BMP4 (50 ng ml−1) stimulation from the induction channel leads to patterning of the epiblast-like cyst; cells directly exposed to BMP4 are specified to the amniotic ectoderm-like fate. Inhibition of BMP and Wnt signalling by adding noggin (50 ng ml−1) and IWP2 (5 µM) in the cell–loading channel prevents the epiblast-like compartment from losing pluripotency and initiating gastrulation-like events. Bright-field image shows an array of A-ELS at t = 36 h. Scale bar, 80 µm. Experiments were repeated three times with similar results. b, Percentage of A-ELS and uniformly squamous amniotic ectoderm-like cysts at t = 36 h. Representative phase-contrast micrographs are shown. Scale bar, 40 µm. n = 212 cysts from n = 4 independent experiments. Each dot represents data obtained from an independent experiment. Red lines represent the median. c, Cell number in each cyst as a function of time. n = 26 cysts for each time point. Data were pooled from n = 2 independent experiments. Red lines represent the median. d, Representative confocal micrographs showing A-ELS at t = 36 h stained for pSMAD1/5 or TFAP2A. Nuclei were stained with DAPI. Scale bars, 40 µm. Experiments were repeated three times with similar results. e, Representative confocal micrographs showing A-ELS at indicated time points stained for NANOG, E-cadherin and T. Nuclei were stained with DAPI. Scale bars, 40 µm. Experiments were repeated three times with similar results. f, Thickness of amniotic ectoderm-like tissue in A-ELS as a function of time. n = 71, 100 and 106 cysts for t = 24 h, 30 h and 36 h, respectively. Data were pooled from n = 3 independent experiments. Red lines represent the median.
Extended Data Fig. 4 Specification of human primordial germ cell-like cells in posteriorized embryonic-like sac.
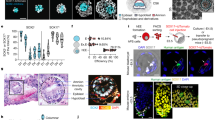
a, Specification of PGCs or PGCLCs in the M. fascicularis embryo (left; ref. 21) and P-ELS (right). b, Representative confocal micrographs showing P-ELS stained for TFAP2C, NANOG and SOX17 or BLIMP1 and SOX17 at indicated time points. Nuclei were stained with DAPI. TFAP2C+SOX17− and SOX17+TFAP2C− cells are marked by green arrows. TFAP2C+SOX17+ cells are identified as nascent, early-stage hPGCLCs. The amniotic ectoderm-like tissue is divided into four quadrants: two middle quadrants furthest away from the epiblast-like pole as central amniotic ectoderm-like compartment (CEN-AM), and two quadrants at the junction of epiblast-like and amniotic ectoderm-like compartments (EPI-AM). TFAP2C+SOX17+ hPGCLCs in the CEN-AM, EPI-AM and epiblast-like compartments are marked by blue, yellow and white arrowheads, respectively. Scale bars, 40 µm. Experiments were repeated three times with similar results. c, Left, dot plot of the numbers of TFAP2C+ and SOX17+ cells at indicated time points. Right, dot plot of the numbers of TFAP2C+SOX17+ and TFAP2C+ NANOG+SOX17+ hPGCLCs at indicated time points. n = 85, 92, 82 and 80 cysts for t = 24 h, 28 h, 32 h and 36 h, respectively. Data were pooled from n = 3 independent experiments. Red lines represent the median. d, Left, spatial distribution of TFAP2C+ and SOX17+ cells at indicated time points. Right, spatial distribution of TFAP2C+SOX17+ and TFAP2C+NANOG+SOX17+ hPGCLCs at indicated time points. n = 85, 92, 82 and 80 cysts for t = 24 h, 28 h, 32 h and 36 h, respectively. Data were pooled from n = 3 independent experiments. e, Representative confocal micrographs showing P-ELS at t = 36 h stained for TFAP2C, T and SOX17. Nuclei were stained with DAPI. T-expressing (TFAP2C+SOX17+T+) and T-non-expressing (TFAP2C+SOX17+T−) hPGCLCs are marked by blue and green arrowheads, respectively. Scale bars, 40 µm. Experiments were repeated three times with similar results. f, Representative confocal micrographs showing P-ELS treated with different doses of the GP130 inhibitor SC144 (top, 0.4 µM; bottom, 2 µM). P-ELS obtained at t = 36 h were stained for TFAP2C, NANOG and SOX17. Nuclei were stained with DAPI. TFAP2C+SOX17+ hPGCLCs in the EPI-AM and epiblast-like compartments are marked by yellow and white arrowheads, respectively. Higher doses of SC144 (8 µM) caused substantial cell death (data not shown). Scale bars, 40 µm. Experiments were repeated twice with similar results.
Extended Data Fig. 5 Microfluidic modelling of human epiblast and amnion development using H1 human ES cells and human induced pluripotent stem cells (hiPSCs) maintained in mTeSR medium as well as H9 human ES cells maintained in Essential 8 medium (E8-H9).
a, Microfluidic generation of epiblast-like cysts. The schematic shows culture conditions and cartoons of epiblast-like cyst development. Representative confocal micrographs show epiblast-like cysts generated from H1 human ES cell, hiPSC and E8–H9 human ES cell at t = 36 h, stained for OCT4, NANOG, SOX2, E-cadherin and laminin as indicated. Fluorescently labelled WGA was used for staining of plasma membrane. b, Microfluidic generation of P-ELS. Schematic shows culture conditions and cartoons of P-ELS development. Representative confocal micrographs show P-ELS generated from H1 human ES cell, hiPSC and E8–H9 human ES cell at t = 36 h stained for TFAP2A, OCT4 and T (top); CDX2, NANOG and T (middle); TFAP2C, NANOG and SOX17 (bottom). TFAP2C+NANOG+SOX17+ hPGCLCs are marked by white arrowheads. c, Microfluidic generation of A-ELS. Culture conditions and cartoons of A-ELS development are shown. Representative confocal micrographs show A-ELS generated from H1 human ES cell, hiPSC and E8–H9 human ES cell at t = 36 h stained for OCT4 and NANOG (top); TFAP2A, NANOG and T (bottom). Fluorescently labelled WGA was used for staining of plasma membrane. In a–c, nuclei were stained with DAPI. Scale bars, 40 µm. All experiments were repeated twice with similar results.
Extended Data Fig. 6 Exogenous Wnt or activin alone is insufficient to generate asymmetric embryonic-like sacs.
Schematics show different culture protocols in which WNT3A (50 ng ml−1), activin A (50 ng ml−1) and/or BMP4 (50 ng ml−1) were supplemented into basal medium in the induction channel as indicated. All cystic tissues were obtained at t = 30 h. a, Representative confocal micrographs showing cysts stained for CDX2, EOMES and T or OCT4 and NANOG. b, Representative confocal micrographs showing cysts stained for CDX2, EOMES and T or OCT4 and NANOG. c, Representative confocal micrographs showing cysts stained for CDX2, NANOG and T or TFAP2A and T. d, Representative confocal micrographs showing cysts stained for CDX2, EOMES and T or TFAP2A and T. In a–d, nuclei were stained with DAPI. Scale bars, 40 µm. All experiments were repeated three times with similar results.
Extended Data Fig. 7 Molecular characterization of posterior and anterior primitive streak-like cell development.
a, Schematic showing posterior primitive streak-like cell development in P-ELS at t = 48 h with BMP4 (50 ng ml−1) supplemented into basal medium in the cell-loading channel. Right, bright-field image shows an array of P-ELS at t = 48 h. Scale bar, 80 µm. b, Representative confocal micrographs show P-ELS stained for E-cadherin and N-cadherin at indicated time points. Thickening of the epiblast-like tissue before the onset of cell dissemination from the PrePS-EPI-like compartment was evident. Outlined regions are magnified in the panel below. c, Dot plot of the thickness of the epiblast-like tissue at indicated time points. For t = 24 h, n = 107 and 116 for epiblast-like cysts and P-ELS, respectively. For t = 30 h, n = 79 and 135 for epiblast-like cysts and P-ELS, respectively. Data were pooled from n = 4 independent experiments. Red lines represent the median. d, Representative confocal micrographs showing P-ELS stained for CDX2, EOMES and T at indicated time points. Outlined regions are magnified in the panel below. Intensity maps show relative intensities of corresponding markers as indicated. e, Representative confocal micrographs show P-ELS stained for TFAP2C, NANOG and SOX17 (left) or BLIMP1 and SOX17 (right) at t = 48 h. Outlined regions are magnified in the panel below. f, Schematic showing anterior primitive streak-like cell development in P-ELS at t = 48 h with BMP4 (50 ng ml−1) and activin A (50 ng ml−1) supplemented into the cell-loading and induction channels, respectively. Right, bright-field image shows an array of P-ELS at t = 48 h. Scale bar, 80 µm. g, Representative confocal micrographs showing staining for E-cadherin, N-cadherin, CDX2, EOMES and T at indicated time points. Outlined regions are magnified in the panel below. h, Representative confocal micrographs show staining for CDX2, EOMES and T at t = 48 h. Outlined regions are magnified in the panel below. Intensity maps show relative intensities of corresponding markers as indicated. i, Representative confocal micrographs show staining for TFAP2C, NANOG and SOX17 at t = 48 h. Outlined regions are magnified in the panel below. In all immunostaining micrographs, nuclei were stained with DAPI; scale bars, 40 µm. All experiments were repeated twice with similar results.
Extended Data Fig. 8 Cell-type identification and characterization using scRNA-seq.
a, Workflow. P-ELS were collected from six microfluidic devices at t = 48 h and were dissociated into single cells. AMLCs obtained using the Transwell method (Transwell-AMLC) and human ES cells cultured on tissue culture plates were dissociated into single cells before the cells were mixed at a 1:2 ratio. scRNA-seq was conducted using 10x Genomics and Illumina HiSeq 4000. Single-cell transcriptome data of P-ELS, Transwell-AMLC and human ES cell were merged and analysed. b, t-SNE plot generated from scRNA-seq data of a total of 9,966 cells, revealing six distinct, colour-coded cell populations (human ES cell, Transwell-AMLC, AMLC, hPGCLC, MeLC1 and MeLC2). Cell numbers of each population are indicated. c, Violin plots of log-transformed, normalized expression levels of genes associated with pluripotency (POU5F1 (also known as OCT4), SOX2, NANOG, PODXL and DPPA4), hPGC (SOX17, TFAP2C, NANOS3, BLIMP1 and PDPN), amniotic ectoderm (TFAP2A, GATA3, HAND1, TCIM (also known as C8orf4) and IGFBP3), mesoderm (T, EOMES, MIXL1, LHX1, MESP2, MESP1, GATA6, LEF1, CDX2 and SNAI2) and HOX proteins (HOXB6, HOXB7, HOXB8, HOXB9 and HOXA10) in the six cell populations as indicated. All cells in b are used for violin plots. d, Heat map of relative expression (Z-score) of top-20 gene signatures distinguishing each cell population. e, DEGs between different cell clusters (MeLC1 against human ES cell; MeLC2 against MeLC1; hPGCLC against human ES cell; AMLC against human ES cell; AMLC against Transwell-AMLC). Top, red and blue bars indicate the numbers of up- and downregulated DEGs, respectively, in indicated pairwise comparisons. Bottom, enrichment of GO terms and representative genes in DEGs from indicated pairwise comparisons. f, Heat map of log-transformed expression levels of selected genes among indicated cell types including those reported by others21,23. Genes are grouped on the basis of their associations with embryonic cell fates, cellular functions, developmental signalling or top-20 DEG status. Left, comparisons among M. fascicularis epiblast (CyEPI) lineage, human ES cell and MeLC1. Right, comparisons among human ES cell, AMLC and Transwell-AMLC. Top-20 DEGs are identified from those upregulated in AMLC against human ES cell (AMLC > human ES cell). Bottom, comparisons between CyEPI, CyPGC, human ES cell and hPGCLC. g, Heat map of correlation coefficients among indicated cell types including those reported by others21,23,25,26. Comparisons between human ES cell, MeLC1, MeLC2 and CyEPI lineages are based on ontogenic genes identified for CyEPI (651 in common out of 776)23. Comparisons between hPGCLC, CyPGC and hPGC are based on ontogenic CyPGC genes (477 in common out of 544)21. Comparisons between human ES cell, Transwell-AMLC and AMLC are based on top-500 DEGs identified in AMLC against human ES cell. Correlation coefficient is calculated using averages of log-transformed expression of common genes. In f, g, CyEPI lineages: ICM, inner cell mass, E6; Pre, pre-implantation epiblast, E7–E9; PostE and PostL, post-implantation early (E13–E14) and late (E16–E17) epiblast, respectively; Gas1/2a/2b, gastrulating cells, E13–E17. For CyPGC and hPGC: ePGC, early CyPGC, E13–E20; lPGC, late gonadal CyPGC, E36–E55, or late gonadal hPGC, E45–E48. Embryonic days are indicated below the heat maps, and colour bars above and besides the heat maps indicate cell types. All genes are listed in Supplementary Table 1.
Extended Data Fig. 9 Inductive effect of amniotic ectoderm-like cells on the onset of gastrulation-like events.
a, Schematic showing P-ELS; the PrePS-EPI-like compartment is divided into four quadrants (R1, R2, R3 and R4) for quantification. Basal medium (BM) comprises E6 and FGF2 (20 ng ml−1). BMP4 (50 ng ml−1) was supplemented into the cell-loading channel. b, Fluorescent and composite images showing dynamic T expression in the PrePS-EPI-like compartment at indicated time points. Experiments were repeated three times with similar results. c, Top, dot plots of relative T intensity in different quadrants of the PrePS-EPI-like compartment at indicated time points. Red lines represent the median. Bottom, spatial maps of average relative T intensity at indicated time points. n = 22 (24 h), 20 (30 h) and 20 (36 h) cysts. Data were pooled from n = 2 independent experiments. d, Live imaging with T-mNeonGreen human ES cell reporter line to track dynamic T expression in the PrePS-EPI-like compartment during the development of P-ELS. e, Characterization of T-mNeonGreen human ES cell reporter line showing co-localization of neon green signal and immunostaining of T. f, Representative confocal micrographs showing development of AMLCs by culturing human ES cells on Transwell membranes in basal medium supplemented with BMP4 (50 ng ml−1) for 48 h. Cells were stained for CDX2, NANOG and T (top) or TFAP2A and OCT4 (bottom). g, Representative confocal micrographs showing AMLCs stained for CDX2, NANOG and T (top) or TFAP2A and OCT4 (bottom). AMLCs were generated from human ES cells by first culturing in basal medium supplemented with BMP4 for 48 h before switching to fresh basal medium for another 48 h. h, Transwell co-culture assays of AMLCs and human ES cells. Human ES cells were first differentiated into AMLCs by culturing on Transwell membranes in basal medium supplemented with BMP4 for 48 h. Culture medium was then switched to fresh basal medium before undifferentiated human ES cells were seeded onto Transwell membranes and co-cultured with AMLCs for another 48 h. Representative confocal micrographs show staining for CDX2, NANOG and T (top); TFAP2A, OCT4 and T (middle); E-cadherin, N-cadherin and T (bottom). i, Transwell co-culture assays of AMLCs and human ES cells. Human ES cells were first differentiated into AMLCs by culturing on Transwell membranes in basal medium supplemented with BMP4 (50 ng ml−1) for 48 h. Culture medium was then switched to fresh basal medium before undifferentiated human ES cells were seeded onto the lower dish and co-cultured with AMLCs for another 48 h. Representative confocal micrographs show staining of cells on the lower dish for OCT4 and T (top); NANOG and T (middle); TFAP2C, NANOG and SOX17 (bottom), or cells on the Transwell membrane for TFAP2C, NANOG and SOX17 as indicated. In all immunostaining micrographs, nuclei were stained with DAPI. Boxed images show magnified views of selected areas. In h, insets show human ES cell colonies seeded after AMLC differentiation as marked by white arrowheads. Scale bars in b, d, e, 40 µm. Scale bars in f–i, 160 µm (main panels) and 10 µm (insets). Experiments were repeated three times with similar results.
Extended Data Fig. 10 Mesoderm induction in posteriorized embryonic-like sac is inhibited by IWP2 but not by IWR1.
a, Schematic shows Transwell co-culture protocol. Human ES cells were first differentiated into AMLCs by culturing in basal medium supplemented with BMP4 (50 ng ml−1) for 48 h. Culture medium was then switched to fresh basal medium supplemented with IWP2 (5 µM). At this point, undifferentiated human ES cells were seeded onto Transwell membranes and co-cultured with AMLCs for another 48 h. Representative confocal micrographs showing staining for CDX2, NANOG and T (top) or TFAP2A and OCT4 (bottom). Insets show human ES cell colonies seeded after AMLC differentiation, as marked by white arrowheads. Scale bars, 160 µm (main panels) and 10 µm (insets). b, Live imaging with the TCF/Lef:H2B-GFP H9 human ES cell reporter line to track Wnt–β-catenin signalling dynamics during embryonic-like sac development with or without the Wnt inhibitors IWR1 (10 µM) or IWP2 (5 µM) supplemented into the induction channel as indicated. Scale bars, 40 µm. c, Representative confocal micrographs show P-ELS obtained at t = 48 h with IWR1 (10 µM) supplemented into the induction channel as indicated. P-ELS were stained for CDX2, EOMES and T (top) or TFAP2C, NANOG and SOX17 (bottom). Outlined regions are magnified in the panel below. Scale bars, 40 µm. In a, c, nuclei were stained with DAPI. Experiments were repeated twice with similar results.
Supplementary information
Supplementary Table 1
List of genes used for data analysis.
Supplementary Table 2
List of primary antibodies.
Video 1
Representative time-lapse video showing synchronized, progressive development of pluripotent epiblast-like cysts from human ES cells through epithelization and lumenogenesis in the microfluidic device. Legends indicate medium conditions in the cell loading and induction channels. Time stamps indicate culture time (t). Scale bar, 80 µm. Experiments were repeated twice with similar results.
Video 2
Representative time-lapse video showing synchronized, progressive development of posteriorized embryonic-like sacs from human ES cells in the microfluidic device. Legends indicate medium conditions in the cell loading and induction channels. Time stamps indicate culture time (t). Scale bar, 80 µm. Experiments were repeated three times with similar results.
Video 3
Representative time-lapse video showing dynamics of amniogenesis and gastrulating cell formation in posteriorized embryonic-like sacs in the microfluidic device. Legends indicate medium conditions in the cell loading and induction channels. Time stamps indicate culture time (t). Scale bar, 80 µm. Experiments were repeated twice with similar results.
Video 4
Representative time-lapse video showing dynamics of brachyury (T) expression in the incipient gastrulating epiblast-like cells during the development of a posteriorized embryonic-like sac. Legends indicate medium conditions in the cell loading and induction channels. Time stamps indicate culture time (t). Scale bar, 80 µm. Experiments were repeated twice with similar results.
Rights and permissions
About this article
Cite this article
Zheng, Y., Xue, X., Shao, Y. et al. Controlled modelling of human epiblast and amnion development using stem cells. Nature 573, 421–425 (2019). https://doi.org/10.1038/s41586-019-1535-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1535-2
This article is cited by
-
Modelling post-implantation human development to yolk sac blood emergence
Nature (2024)
-
Microfluidic high-throughput 3D cell culture
Nature Reviews Bioengineering (2024)
-
Derivation of human primordial germ cell-like cells in an embryonic-like culture
Nature Communications (2024)
-
Bioelectric stimulation controls tissue shape and size
Nature Communications (2024)
-
Hypoblast from human pluripotent stem cells regulates epiblast development
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.