Abstract
Enzymes that catalyse CpG methylation in DNA, including the DNA methyltransferases 1 (DNMT1), 3A (DNMT3A) and 3B (DNMT3B), are indispensable for mammalian tissue development and homeostasis1,2,3,4. They are also implicated in human developmental disorders and cancers5,6,7,8, supporting the critical role of DNA methylation in the specification and maintenance of cell fate. Previous studies have suggested that post-translational modifications of histones are involved in specifying patterns of DNA methyltransferase localization and DNA methylation at promoters and actively transcribed gene bodies9,10,11. However, the mechanisms that control the establishment and maintenance of intergenic DNA methylation remain poorly understood. Tatton–Brown–Rahman syndrome (TBRS) is a childhood overgrowth disorder that is defined by germline mutations in DNMT3A. TBRS shares clinical features with Sotos syndrome (which is caused by haploinsufficiency of NSD1, a histone methyltransferase that catalyses the dimethylation of histone H3 at K36 (H3K36me2)8,12,13), which suggests that there is a mechanistic link between these two diseases. Here we report that NSD1-mediated H3K36me2 is required for the recruitment of DNMT3A and maintenance of DNA methylation at intergenic regions. Genome-wide analysis shows that the binding and activity of DNMT3A colocalize with H3K36me2 at non-coding regions of euchromatin. Genetic ablation of Nsd1 and its paralogue Nsd2 in mouse cells results in a redistribution of DNMT3A to H3K36me3-modified gene bodies and a reduction in the methylation of intergenic DNA. Blood samples from patients with Sotos syndrome and NSD1-mutant tumours also exhibit hypomethylation of intergenic DNA. The PWWP domain of DNMT3A shows dual recognition of H3K36me2 and H3K36me3 in vitro, with a higher binding affinity towards H3K36me2 that is abrogated by TBRS-derived missense mutations. Together, our study reveals a trans-chromatin regulatory pathway that connects aberrant intergenic CpG methylation to human neoplastic and developmental overgrowth.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The WGBS, ChIP–seq and RNA-sequencing data have been deposited in the GEO under accession number GSE118785.
Code availability
The source code for bioinformatic analysis is available on request.
References
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Wu, H. et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science 329, 444–448 (2010).
Challen, G. A. et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat. Genet. 44, 23–31 (2012).
Ley, T. J. et al. DNMT3A mutations in acute myeloid leukemia. N. Engl. J. Med. 363, 2424–2433 (2010).
Klein, C. J. et al. Mutations in DNMT1 cause hereditary sensory neuropathy with dementia and hearing loss. Nat. Genet. 43, 595–600 (2011).
Xu, G. L. et al. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature 402, 187–191 (1999).
Tatton-Brown, K. et al. Mutations in the DNA methyltransferase gene DNMT3A cause an overgrowth syndrome with intellectual disability. Nat. Genet. 46, 385–388 (2014).
Ooi, S. K. T. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448, 714–717 (2007).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Morselli, M. et al. In vivo targeting of de novo DNA methylation by histone modifications in yeast and mouse. eLife 4, e06205 (2015).
Kurotaki, N. et al. Haploinsufficiency of NSD1 causes Sotos syndrome. Nat. Genet. 30, 365–366 (2002).
Rayasam, G. V. et al. NSD1 is essential for early post-implantation development and has a catalytically active SET domain. EMBO J. 22, 3153–3163 (2003).
Rao, B., Shibata, Y., Strahl, B. D. & Lieb, J. D. Dimethylation of histone H3 at lysine 36 demarcates regulatory and nonregulatory chromatin genome-wide. Mol. Cell. Biol. 25, 9447–9459 (2005).
Xiao, T. et al. Phosphorylation of RNA polymerase II CTD regulates H3 methylation in yeast. Genes Dev. 17, 654–663 (2003).
Doynova, M. D., Markworth, J. F., Cameron-Smith, D., Vickers, M. H. & O’Sullivan, J. M. Linkages between changes in the 3D organization of the genome and transcription during myotube differentiation in vitro. Skelet. Muscle 7, 5 (2017).
Kuo, A. J. et al. NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol. Cell 44, 609–620 (2011).
Orlando, D. A. et al. Quantitative ChIP-seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Dhayalan, A. et al. The Dnmt3a PWWP domain reads histone 3 lysine 36 trimethylation and guides DNA methylation. J. Biol. Chem. 285, 26114–26120 (2010).
Papillon-Cavanagh, S. et al. Impaired H3K36 methylation defines a subset of head and neck squamous cell carcinomas. Nat. Genet. 49, 180–185 (2017).
Choufani, S. et al. NSD1 mutations generate a genome-wide DNA methylation signature. Nat. Commun. 6, 10207 (2015).
Tatton-Brown, K. et al. Mutations in epigenetic regulation genes are a major cause of overgrowth with intellectual disability. Am. J. Hum. Genet. 100, 725–736 (2017).
Heyn, P. et al. Gain-of-function DNMT3A mutations cause microcephalic dwarfism and hypermethylation of Polycomb-regulated regions. Nat. Genet. 51, 96–105 (2019).
Jeffries, A. R. et al. Growth disrupting mutations in epigenetic regulatory molecules are associated with abnormalities of epigenetic aging. Genome Res. 29, 1057–1066 (2019).
Williams, K., Christensen, J. & Helin, K. DNA methylation: TET proteins—guardians of CpG islands? EMBO Rep. 13, 28–35 (2011).
Boulard, M., Edwards, J. R. & Bestor, T. H. FBXL10 protects Polycomb-bound genes from hypermethylation. Nat. Genet. 47, 479–485 (2015).
Quintana, R. M. et al. A transposon-based analysis of gene mutations related to skin cancer development. J. Invest. Dermatol. 133, 239–248 (2013).
Rinaldi, L. et al. Loss of Dnmt3a and Dnmt3b does not affect epidermal homeostasis but promotes squamous transformation through PPAR-γ. eLife 6, e21697 (2017).
Jaju, R. J. et al. A novel gene, NSD1, is fused to NUP98 in the t(5;11)(q35;p15.5) in de novo childhood acute myeloid leukemia. Blood 98, 1264–1267 (2001).
Sendžikaitė, G., Hanna, C. W., Stewart-Morgan, K. R., Ivanova, E. & Kelsey, G. A DNMT3A PWWP mutation leads to methylation of bivalent chromatin and growth retardation in mice. Nat. Commun. 10, 1884 (2019).
Sidoli, S., Bhanu, N. V., Karch, K. R., Wang, X. & Garcia, B. A. Complete workflow for analysis of histone post-translational modifications using bottom-up mass spectrometry: from histone extraction to data analysis. J. Vis. Exp. 111, e54112 (2016).
Yuan, Z. F. et al. EpiProfile 2.0: a computational platform for processing epi-proteomics mass spectrometry data. J. Proteome Res. 17, 2533–2541 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Killick, R. et al. Optimal detection of changepoints with a linear computation cost. J. Am. Stat. Assoc. 107, 1590–1598 (2012).
Wambui, G. D. et al. The power of the pruned exact linear time (PELT) test in multiple changepoint detection. Am. J. Theor. Appl. Stat. 4, 581–586 (2015).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: Quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Hovestadt, V. et al. Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510, 537–541 (2014).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat. Genet. 44, 40–46 (2012).
Acknowledgements
We thank T. Bestor and members of the Lu, Majewski and Allis laboratories for critical reading of the manuscript. This research was supported by US National Institutes of Health (NIH) grants (P01CA196539 to N.J., B.A.G., C.D.A. and J.M.; R00CA212257 to C.L.; T32GM007739 and F30CA224971 to D.N.W.; T32GM008275 to D.M.M.; and R44GM116584 and R44GM117683 to M.-C.K.), the Rockefeller University (C.D.A.) and the Cedars Cancer Foundation (N.J.). The research was funded in part through the NIH/NCI Cancer Center Support Grant P30CA013696 and used the HICCC Flow Cytometry Shared Resource. This work was performed within the context of the I-CHANGE consortium and was supported by funding from Genome Canada, Genome Quebec, the Institute for Cancer Research of the CIHR, McGill University and the Montreal Children’s Hospital Foundation. B.A.G. is funded by a Leukemia and Lymphoma Society Dr. Robert Arceci Scholar Award; H.L. is funded by the National Natural Science Foundation of China (31725014 and 91753203); C.L. is the Giannandrea Family Dale F. Frey Breakthrough Scientist of the Damon Runyon Foundation (DFS-28-18), a Pew-Stewart Scholar for Cancer Research and supported by an AACR Gertrude B. Elion Cancer Research Grant; N.J. is a member of the Penny Cole Laboratory and the recipient of a Chercheur Boursier, Chaire de Recherche Award from the Fond de la Recherche du Québec en Santé; and S.P.-C. and A.S.H. are supported by a studentship and postdoctoral fellowship from the Fond de la Recherche du Québec en Santé, respectively.
Author information
Authors and Affiliations
Contributions
D.N.W., S.P.-C., C.D.A., J.M. and C.L. conceived and designed the experiments. D.N.W., K.N.R., C.H., J.T.M., X.X., A.E.L., D.M.M., A.S.H., N.J., B.A.G. and C.L. performed cell-based experiments and analysed data. S.P.-C., H.C., X.C., H.N., E.B., A.D. and J.M. performed bioinformatic analysis on sequencing-based data. Y.Y., M.R.M., M.J.M., M.A.C., M.-C.K. and H.L. performed in vitro experiments with recombinant proteins and analysed data. H.C. and Y.Y. contributed equally as co-second authors to this manuscript. All authors contributed to the written manuscript.
Corresponding authors
Ethics declarations
Competing interests
EpiCypher (M.R.M., M.J.M., M.A.C. and M.-C.K.) is a commercial developer and supplier of platforms similar to those used in this study: recombinant semi-synthetic modified nucleosomes and the dCypher nucleosome-binding assay.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Taiping Chen, Jonathan Licht and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 H3K36me2 and H3K36me3 mark transcriptionally active euchromatin.
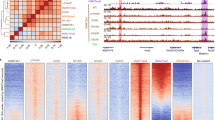
a, ChIP–seq normalized reads of H3K36me2 relative to the average percentage of CpG methylation in mouse MSCs for 100-kb non-overlapping bins (n = 25,624). Pearson’s correlation coefficient is indicated. b, Quantification of ChIP–seq normalized reads for averaged CpG methylation, DNMT3A1 and DNMT3B within H3K36me2 and H3K36me3 (n = 591), H3K27me3 (n = 283) or H3K9me3 (n = 545) domains. P values were <2.2 × 10−16 for all pair-wise comparisons, as determined by Wilcoxon’s rank-sum test (two-sided), except for DNMT3B (P = 3.73 × 10−14) between H3K27me3 and H3K9me3 domains. For box plots, the centre lines represent the median, the box limits are the 25th and 75th percentiles, the whiskers are the minimum to maximum values and discrete points represent outliers. c, Quantification of ChIP–seq normalized reads for gene expression, H3K4me1 and H3K27ac within H3K36me2 and H3K36me3 (n = 591), H3K27me3 (n = 283) or H3K9me3 (n = 545) domains. P values were <2.2 × 10−16 for all pair-wise comparisons, as determined by Wilcoxon’s rank-sum test (two-sided), except for gene expression (P = 1.72 × 10−6) between H3K27me3 and H3K9me3 domains. Box plots as in b. d, Three-dimensional genome browser representation of Hi-C chromatin conformation capture data from mouse myoblasts compared to ChIP–seq normalized reads for histone PTMs in mouse MSCs at Chr4: 52.2–99.1 Mb. The levels of CpG methylation (CpGme) are depicted as a heat map (blue, low; white, intermediate; red, high). For H3K36me3 and H3K36me2, data are representative of two independent ChIP–seq experiments; for H3K27me3 and H3K9me3, ChIP–seq was performed once and an independent ChIP was performed in which genomic regions of selective enrichment and depletion were confirmed by qPCR. WGBS was performed once.
Extended Data Fig. 2 Distinct enrichment patterns between H3K36me2 and H3K36me3 at euchromatin.
a, Genome browser representation of ChIP–seq normalized reads for H3K36me3, H3K36me2, DNMT3A1 and DNMT3B in mouse MSCs at Chr8: 75.0–75.1 Mb. The levels of CpG methylation are depicted as a heat map (blue, low; white, intermediate; red, high) and RefSeq genes are annotated at the bottom. For H3K36me3, H3K36me2 and DNMT3A1, data are representative of two independent ChIP–seq experiments; for DNMT3B, ChIP–seq was performed once and an independent ChIP was performed in which genomic regions of selective enrichment and depletion were confirmed by qPCR. WGBS was performed once. b, Ratio of observed-to-expected ChIP–seq reads for H3K36me2 and H3K36me3 in annotated genomic regions. Numbers of expected reads were generated assuming equivalent genomic distribution to input. c, Heat maps representing ChIP–seq signal density for H3K36me2 and H3K36me3 in mouse MSCs across all gene bodies. Genes are ranked by the length of the first intron. Each gene is displayed as a row. d, Averaged ChIP–seq normalized signal across gene bodies stratified by expression quartile, represented as log2(ChIP signal/input signal) over input for DNMT3A1 (top) and DNMT3B (bottom) in parental MSCs. Sample sizes (which are the same for DNMT3A1 and DNMT3B) are n = 7,524, n = 7,435, n = 6,923 and n = 8,550 for the first, second, third and fourth quartiles, respectively; and n = 35,777 for the average. TES, transcription end site; TSS, transcription start site.
Extended Data Fig. 3 Genome-wide colocalization between H3K36me2 and DNMT3A2 in mouse ES cells.
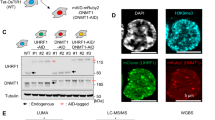
a, Immunoblots of lysates generated from parental and sgDnmt3a mouse ES cells (termed mESCs in the Extended Data figures) that ectopically express HA-tagged DNMT3A2. Vinculin was used as a loading control. Endogenous expression of the long isoform (DNMT3A1) and the short isoform (DNMT3A2) of DNMT3A is indicated. Data are representative of two independent experiments. b, ChIP–seq normalized reads of HA-tagged DNMT3A2 in sgDnmt3a relative to parental ES cells for 100-kb non-overlapping bins (n = 26,181). Pearson’s correlation coefficient is indicated. c, Genome browser representation of ChIP–seq normalized reads for H3K36me3, H3K36me2, DNMT3A2 and DNMT3B in ES cells at Chr12: 86.6–87.5 Mb. RefSeq genes are annotated at the bottom. The shaded areas indicate H3K36me2-enriched intergenic regions (orange) and H3K36me3-enriched genic regions (green) in parental cells. For H3K36me2 and DNMT3A2, ChIP–seq was performed once and an independent ChIP was performed in which genomic regions of selective enrichment and depletion were confirmed by qPCR. H3K36me3 and DNMT3B ChIP–seq were performed once. d, ChIP–seq normalized reads per 10-kb bin for DNMT3A2 (y axis) and DNMT3B (x axis) in ES cells (n = 246,285). Bins (dots) are colour-coded on the basis of differences between H3K36me3 and H3K36me2 ChIP–seq reads, to show selective enrichment for H3K36me2 (orange) or H3K36me3 (green). e, De novo methylation per bin by DNMT3A2 (brown) or DNMT3B (grey) after reintroduction into Dnmt1, Dnmt3a and Dnmt3b triple-knockout ES cells relative to H3K36me3. To generate bins, 1-kb genomic tiles (n = 2,462,755) were ranked by H3K36me3 enrichment in ES cells and grouped into 1,000 bins, ordered by rank (2,463 tiles per group). The dashed lines indicate H3K36me3 enrichment per bin. Goodness of fit was computed using a quadratic model (red lines). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Genetic ablation of Nsd1 and Nsd2 in mouse MSCs and Nsd1 in mouse ES cells.
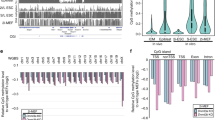
a, Immunoblots of lysates from parental and H3K36-methyltransferase-knockout mouse MSC clonal lines for NSD1, NSD2 and SETD2. Vinculin was used as a loading control. b, Immunoblots of lysates from parental and sgNsd1 mouse ES cells that express HA-tagged DNMT3A. sgNsd1 cells were rescued with ectopic expression of wild-type NSD1 or the C2023A catalytic mutant. Vinculin was used as a loading control. c, Immunoblots of lysates generated from parental, sgSetd2 and sgNsd1/2 MSCs for H3K36me3 and H3K36me2. Total H3 was used as a loading control. d, Ratios of ChIP–seq reads for H3K36me2 and H3K36me3 in mouse MSCs between target chromatin (mouse) and reference spike-in chromatin (Drosophila). Data are representative of two independent experiments. e, Quantitative mass spectrometry measurement of the abundance of histone PTMs in acid-extracted histones derived from the indicated MSC lines. Data are from one experiment. f, Ratios of ChIP–seq reads for H3K36me2 and H3K36me3 in ES cells between target chromatin (mouse) and reference spike-in chromatin (Drosophila). Data are from one experiment. g, Density plots of H3K36me2 levels at intergenic (n = 1,165), exonic (n = 13,601) and intronic (n = 12,364) regions for parental (grey) and sgNsd1/2 (orange) MSCs. P values were determined by Wilcoxon’s rank-sum test (two-sided). h, Density plots of H3K36me2 levels at intergenic (n = 1,165), exonic (n = 13,601) and intronic (n = 12,364) regions for parental (grey) and sgNsd1 (orange) ES cells. P values were determined by Wilcoxon’s rank-sum test (two-sided). The immunoblot experiments in a–c were independently repeated twice with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 5 H3K36me2 depletion impairs genomic targeting of DNMT3A and reduces intergenic CpG methylation.
a, Immunoblots of lysates from parental, sgSetd2 and sgNsd1/2 mouse MSCs that ectopically express HA-tagged DNMT3A1 or DNMT3B. β-tubulin was used as a loading control. b, Immunoblots of lysates from sgDnmt3a, parental and sgNsd1 mouse ES cells that ectopically express HA-tagged DNMT3A2 or DNMT3B. Vinculin was used as a loading control. c, Genome browser representation of ChIP–seq normalized reads for H3K36me3, H3K36me2 and DNMT3A1 in parental and sgNsd1/2 MSCs at Chr17: 87.9–88.7 Mb. The levels of CpG methylation (CpGme) are depicted as a heat map (blue, low; white, intermediate; red, high) and RefSeq genes are annotated at the bottom. The shaded area indicates H3K36me2-enriched intergenic region in parental cells. For H3K36me2, H3K36me3 and DNMT3A1 in parental cells, data are representative of two independent ChIP–seq experiments. DNMT3A1 ChIP–seq in sgNsd1/2 cells and WGBS in both lines were performed once. d, Genome browser representation of ChIP–seq normalized reads for H3K36me3, H3K36me2 and DNMT3A2 in parental and sgNsd1 ES cells at Chr17: 87.9-88.7 Mb. The levels of CpG methylation are depicted as a heat map (blue, low; white, intermediate; red, high) and RefSeq genes are annotated at the bottom. The shaded area indicates the H3K36me2-enriched intergenic region in parental cells. For H3K36me2 and DNMT3A2, ChIP–seq was performed once and an independent ChIP was performed in which genomic regions of selective enrichment and depletion were confirmed by qPCR. WGBS and H3K36me3 ChIP–seq were performed once. e, Per cent change in averaged CpG methylation between parental and sgNsd1/2 MSCs relative to changes in ChIP–seq normalized reads of H3K36me2 for 100-kb non-overlapping bins (n = 25,611). Pearson’s correlation coefficient is indicated. f, Per cent change in averaged CpG methylation between parental and sgNsd1 ES cells relative to changes in ChIP–seq normalized reads of H3K36me2 for 100-kb non-overlapping bins (n = 26,044). Pearson’s correlation coefficient is indicated. g, The enrichment (per cent input) of H3K36me2 at various intergenic regions in parental (black) and sgNsd1 (orange) ES cells rescued with ectopic expression of wild-type NSD1 or the C2023A catalytic mutant was measured by ChIP–qPCR. Each data point represents a genomic locus (n = 10 for H3K36me2-enriched regions, n = 4 for H3K36me2-depleted regions). Data are mean ± s.d. P values were determined by one-way ANOVA. The immunoblot experiments in a, b were independently repeated twice with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 6 The PWWP domain of DNMT3A preferentially binds to H3K36me2.
a, The dCypher approach for interrogating chromatin readers (Methods). Biotinylated nucleosomes are immobilized on streptavidin donor beads and GST-tagged DNMT3APWWP on glutathione AlphaLISA acceptor beads. Laser excitation of the donor generates singlet oxygen, which diffuses to activate emission from the acceptor. Fluorescence counts are directly proportional to the amount of donor–acceptor complex bridged by the nucleosome-reader interaction. b, Quantification of AlphaLISA counts for isolated GST-tagged DNMT3APWWP interaction, titrated against H3K36-modified nucleosomes. Data are the mean of two replicates. c, ITC titration and fitting curves of human DNMT3APWWP with H3.3(1–42) K36-modified peptides. d, ITC titration and fitting curves of human DNMT3APWWP with H3.1(1–42) K36-modified peptides.
Extended Data Fig. 7 Redistribution of DNMT3A to H3K36me3-marked gene bodies after loss of H3K36me2.
a, Averaged ChIP–seq normalized signal across all gene bodies, represented as log2(ChIP signal/input signal) for DNMT3A1 (grey, n = 14,959) and DNMT3B (green, n = 14,959) in parental mouse MSCs, and for DNMT3A1 (orange, n = 14,311) in sgNsd1/2 MSCs. b, Quantification of ChIP–seq normalized reads of DNMT3A1 in parental, sgNsd1/2 and triple-knockout (TKO) MSCs at 10-kb non-overlapping bins enriched for H3K36me3 in parental cells (top 20% of bins, n = 54,624). Reads were normalized by read depth. P values were <2.2 × 10−16 for all pair-wise comparisons, as determined by Wilcoxon’s rank-sum test (two-sided). For box plots, the centre lines represent the median, the box limits are the 25th and 75th percentiles, the whiskers are the minimum to maximum values and discrete points represent outliers. c, Genome browser representation of ChIP–seq normalized reads for H3K36me2, H3K36me3 and DNMT3A1 in parental, sgNsd1/2 and triple-knockout MSCs at Chr10: 74.8–77.8 Mb. RefSeq genes are annotated at the bottom. The shaded areas indicate H3K36me3-enriched genic regions in parental cells. For H3K36me2 and H3K36me3, data in parental and sgNsd1/2 cells are representative of two independent ChIP–seq experiments. ChIP–seq in triple-knockout cells for H3K36me2 and H3K36me3 was performed once. For DNMT3A1, data in parental cells are representative of two independent ChIP–seq experiments. ChIP–seq in sgNsd1/2 and triple-knockout cells for DNMT3A1 was performed once. d, Immunoblots of lysates from parental, sgNsd1/2 and triple-knockout MSCs that ectopically express HA-tagged wild-type or D333A-mutant DNMT3A1. β-actin was used as a loading control. Data are representative of two independent experiments. e, Fold enrichment of wild-type or PWWP-mutant (D333A) DNMT3A1 at gene-body versus intergenic regions in parental (black) and sgNsd1/2 (orange) MSCs was measured by ChIP–qPCR. Each data point represents a genomic locus (n = 10). Data are mean ± s.d. P values were determined by one-way ANOVA. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 Assessment of binding affinity between DNMT3B and H3K36me2 in vitro and in cells.
a, Immunoblots of His–MBP-tagged recombinant DNMT3A and DNMT3B PWWP domains. b, Immunoblots of recombinant His–MBP-tagged wild-type DNMT3A and DNMT3B PWWP domains bound to H3K36-modified recombinant nucleosomes following the in vitro pull-down assay. c, ChIP–seq normalized reads per bin for DNMT3A1 (grey) and DNMT3B (blue) in parental mouse MSCs and DNMT3B (red) in sgSetd2 MSCs relative to H3K36me2. To generate bins, 1-kb genomic tiles were ranked by H3K36me2 enrichment in parental MSCs and grouped into 1,000 bins, ordered by rank. The dashed line indicates H3K36me2 enrichment per bin. d, Genome browser representation of ChIP–seq normalized reads for H3K36me2, H3K36me3 and DNMT3B in parental and sgSetd2 MSCs at Chr10: 83.4–84.3 Mb. RefSeq genes are annotated at the bottom. The shaded areas indicate H3K36me2-enriched intergenic regions (orange) and H3K36me3-enriched genic regions (green) in parental cells. For H3K36me2 and H3K36me3, data are representative of two independent ChIP–seq experiments; for DNMT3B, ChIP–seq was performed once and an independent ChIP was performed in which genomic regions of selective enrichment and depletion were confirmed by qPCR. The immunoblot experiments in a, b were independently repeated twice with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 TBRS-associated mutations in DNMT3A are loss-of-function and result in DNA hypomethylation similar to that observed in Sotos syndrome.
a, Immunoblots of the recombinant His–MBP-tagged DNMT3A PWWP domain containing TBRS-associated mutations. MBP alone was used as a control. b, Immunoblots of soluble or chromatin-associated lysates generated from cells that ectopically express HA-tagged wild-type or mutant (W297del or I310N) DNMT3A. β-tubulin and lamin B1 were used as loading controls for the soluble and chromatin-associated fractions, respectively. c, Immunoblots of lysates from parental mouse MSCs that ectopically express HA-tagged wild-type or TBRS-associated mutant (W297del, I310N or Y365C) DNMT3A. Data are from one experiment. β-tubulin was used as a loading control. d, Immunoblots of nucleosomes bound to HA-tagged wild-type or mutant (W297del, I310N or Y365C) DNMT3A after ChIP with anti-HA-tag antibodies. Total H3 was used as a loading control and to normalize for differences in protein expression and nucleosome pull-down efficiency between samples. e, Genome browser representation of ChIP–seq normalized reads for H3K36me2, wild-type DNMT3A1 and I310N-mutant DNMT3A1 in mouse MSCs at Chr13: 62.2–66.8 Mb. The levels of CpG methylation (CpGme) are depicted as a heat map (blue, low; white, intermediate; red, high) and RefSeq genes are annotated at the bottom. f, ChIP–seq normalized reads of wild-type or PWWP-mutant (I310N) DNMT3A1 relative to that of H3K36me2 for 100-kb non-overlapping bins (wild type: n = 25,694; I310N: n = 25,757). Pearson’s correlation coefficient is indicated. For H3K36me2 and wild-type DNMT3A1, data are representative of two independent ChIP–seq experiments; for I310N-mutant DNMT3A1, ChIP–seq was performed once and an independent ChIP was performed in which genomic regions of selective enrichment and depletion were confirmed by qPCR. WGBS was performed once. g, Unsupervised hierarchical clustering of publicly available Infinium Human Methylation 450K array profiles of blood samples from patients with TBRS, Sotos syndrome and Weaver syndrome and healthy control individuals, based on the top 1,000 most-variable probes. h, Model depicting the changes in chromatin landscape in human developmental overgrowth syndromes (Sotos syndrome and TBRS). In normal development, DNMT3A and DNMT3B act in parallel to methylate CpG dinucleotides at H3K36me2-enriched intergenic and H3K36me3-enriched genic regions, respectively. Haploinsufficiency of NSD1 and depletion of intergenic H3K36me2 levels in Sotos syndrome abrogate PWWP-mediated intergenic recruitment of DNMT3A, which leads to hypomethylation of intergenic DNA and the redistribution of DNMT3A to H3K36me3-enriched gene bodies. Mutations within the PWWP domain of DNMT3A in TBRS impair chromatin occupancy and reduce cellular levels of the protein—thereby also causing hypomethylation of intergenic DNA. The immunoblot experiments in a, c, d were independently repeated twice with similar results. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Figure 1
Source Gel images (15 display items). One for each figure panel that contains immunoblot analysis, with red box indicating cropped region.
Supplementary Table 1
Sanger sequencing verification results for CRISPR-Cas9 gene editing of Setd2, Nsd1, Nsd2, and Dnmt3a in mMSCs and mESCs.
Supplementary Table 2
Primers used for ChIP-qPCR analysis in this study.
Rights and permissions
About this article
Cite this article
Weinberg, D.N., Papillon-Cavanagh, S., Chen, H. et al. The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape. Nature 573, 281–286 (2019). https://doi.org/10.1038/s41586-019-1534-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1534-3
This article is cited by
-
Epigenetics in diabetic cardiomyopathy
Clinical Epigenetics (2024)
-
Polygenic risk score model for renal cell carcinoma in the Korean population and relationship with lifestyle-associated factors
BMC Genomics (2024)
-
Interrogating epigenetic mechanisms with chemically customized chromatin
Nature Reviews Genetics (2024)
-
Targeted DNA integration in human cells without double-strand breaks using CRISPR-associated transposases
Nature Biotechnology (2024)
-
Structure-guided functional suppression of AML-associated DNMT3A hotspot mutations
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.