Abstract
Through the lens of evolution, climate change is an agent of natural selection that forces populations to change and adapt, or face extinction. However, current assessments of the risk of biodiversity associated with climate change1 do not typically take into account how natural selection influences populations differently depending on their genetic makeup2. Here we make use of the extensive genome information that is available for Arabidopsis thaliana and measure how manipulation of the amount of rainfall affected the fitness of 517 natural Arabidopsis lines that were grown in Spain and Germany. This allowed us to directly infer selection along the genome3. Natural selection was particularly strong in the hot-dry location in Spain, where 63% of lines were killed and where natural selection substantially changed the frequency of approximately 5% of all genome-wide variants. A significant portion of this climate-driven natural selection of variants was predictable from signatures of local adaptation (R2 = 29–52%), as genetic variants that were found in geographical areas with climates more similar to the experimental sites were positively selected. Field-validated predictions across the species range indicated that Mediterranean and western Siberian populations—at the edges of the environmental limits of this species—currently experience the strongest climate-driven selection. With more frequent droughts and rising temperatures in Europe4, we forecast an increase in directional natural selection moving northwards from the southern end of Europe, putting many native A. thaliana populations at evolutionary risk.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data are available in the Supplementary Tables and Supplementary Data and are deposited in Figshare (https://doi.org/10.6084/m9.figshare.6756836 and https://doi.org/10.6084/m9.figshare.6480599). Genomes are available at http://1001genomes.org/data/GMI-MPI/releases/v3.1/. The seed collection can be obtained from the Arabidopsis Biological Resource Center (ABRC) under accession CS78942 (https://abrc.osu.edu/stocks/465820). The PLINK files for genome-wide association scans of fitness and climate are deposited in the AraGWAS Catalog (https://aragwas.1001genomes.org/; https://doi.org/10.21958/study:34).
Code availability
Field data cleaning and processing scripts are available from GitHub (https://github.com/MoisesExpositoAlonso/dryAR) and Zenodo (https://doi.org/10.5281/zenodo.2583224). Plant rosette area scripts are available from GitHub (http://github.com/MoisesExpositoAlonso/hippo) and Zenodo (https://doi.org/10.5281/zenodo.1039888). Inflorescence analysis scripts are available from GitHub (http://github.com/MoisesExpositoAlonso/hitfruit) and Zenodo (https://doi.org/10.5281/zenodo.2583262). Simulations of total selection coefficients inference and population allele frequency changes are available from GitHub (https://github.com/MoisesExpositoAlonso/selectioncorrelatedgenotypes) and Zenodo (https://doi.org/10.5281/zenodo.1408095).
Change history
30 September 2019
An Amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Urban, M. C. Accelerating extinction risk from climate change. Science 348, 571–573 (2015).
Hoffmann, A. A. & Sgrò, C. M. Climate change and evolutionary adaptation. Nature 470, 479–485 (2011).
Thurman, T. J. & Barrett, R. D. H. The genetic consequences of selection in natural populations. Mol. Ecol. 25, 1429–1448 (2016).
IPCC Climate Change 2013: The Physical Science Basis (eds Stocker, T. F. et al.) (Cambridge Univ. Press, 2014).
Jezkova, T. & Wiens, J. J. Rates of change in climatic niches in plant and animal populations are much slower than projected climate change. Proc. R. Soc. Lond. B 283, 20162104 (2016).
Nielsen, R. et al. Genomic scans for selective sweeps using SNP data. Genome Res. 15, 1566–1575 (2005).
Bonhomme, M. et al. Detecting selection in population trees: the Lewontin and Krakauer test extended. Genetics 186, 241–262 (2010).
Exposito-Alonso, M. et al. Genomic basis and evolutionary potential for extreme drought adaptation in Arabidopsis thaliana. Nat. Ecol. Evol. 2, 352–358 (2018).
Bay, R. A. et al. Genomic signals of selection predict climate-driven population declines in a migratory bird. Science 359, 83–86 (2018).
Coop, G., Witonsky, D., Di Rienzo, A. & Pritchard, J. K. Using environmental correlations to identify loci underlying local adaptation. Genetics 185, 1411–1423 (2010).
Hancock, A. M. et al. Adaptation to climate across the Arabidopsis thaliana genome. Science 334, 83–86 (2011).
Fitzpatrick, M. C. & Keller, S. R. Ecological genomics meets community-level modelling of biodiversity: mapping the genomic landscape of current and future environmental adaptation. Ecol. Lett. 18, 1–16 (2015).
Kingsolver, J. G. et al. The strength of phenotypic selection in natural populations. Am. Nat. 157, 245–261 (2001).
Savolainen, O., Lascoux, M. & Merilä, J. Ecological genomics of local adaptation. Nat. Rev. Genet. 14, 807–820 (2013).
Gompert, Z. et al. Experimental evidence for ecological selection on genome variation in the wild. Ecol. Lett. 17, 369–379 (2014).
1001 Genomes Consortium. 1,135 genomes reveal the global pattern of polymorphism in Arabidopsis thaliana. Cell 166, 481–491 (2016).
Gompert, Z., Egan, S. P., Barrett, R. D. H., Feder, J. L. & Nosil, P. Multilocus approaches for the measurement of selection on correlated genetic loci. Mol. Ecol. 26, 365–382 (2017).
Zhou, X., Carbonetto, P. & Stephens, M. Polygenic modeling with Bayesian sparse linear mixed models. PLoS Genet. 9, e1003264 (2013).
Charlesworth, B. The effects of deleterious mutations on evolution at linked sites. Genetics 190, 5–22 (2012).
Kojima, K. & Lewontin, R. C. in Mathematical Topics in Population Genetics (ed. Kojima, K.) 367–388 (Springer, 1970)
Neher, R. A. Genetic draft, selective interference, and population genetics of rapid adaptation. Annu. Rev. Ecol. Evol. Syst. 44, 195–215 (2013).
Anderson, J. T., Lee, C.-R. & Mitchell-Olds, T. Strong selection genome-wide enhances fitness trade-offs across environments and episodes of selection. Evolution 68, 16–31 (2014).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005).
Nosil, P. et al. Natural selection and the predictability of evolution in Timema stick insects. Science 359, 765–770 (2018).
Fournier-Level, A. et al. A map of local adaptation in Arabidopsis thaliana. Science 334, 86–89 (2011).
Manzano-Piedras, E., Marcer, A., Alonso-Blanco, C. & Picó, F. X. Deciphering the adjustment between environment and life history in annuals: lessons from a geographically-explicit approach in Arabidopsis thaliana. PLoS ONE 9, e87836 (2014).
Abatzoglou, J. T., Dobrowski, S. Z., Parks, S. A. & Hegewisch, K. C. TerraClimate, a high-resolution global dataset of monthly climate and climatic water balance from 1958–2015. Sci. Data 5, 170191 (2018).
Samaniego, L. et al. Anthropogenic warming exacerbates European soil moisture droughts. Nat. Clim. Change 8, 421–426 (2018).
Asner, G. P., Nepstad, D., Cardinot, G. & Ray, D. Drought stress and carbon uptake in an Amazon forest measured with spaceborne imaging spectroscopy. Proc. Natl Acad. Sci. USA 101, 6039–6044 (2004).
Aitken, S. N. & Bemmels, J. B. Time to get moving: assisted gene flow of forest trees. Evol. Appl. 9, 271–290 (2016).
Acknowledgements
We thank P. Lang, A. Hancock and T. Karasov for comments on the manuscript, and members of the Weigel and Burbano laboratories for discussions; X. Picó for advice on experimental design, I. Bezrukov for advice on image-processing replicability; and B. Mendez-Vigo, C. Alonso-Blanco, A. López Quirós, M. López Herránz and M. Ángel Mora Plaza for assistance during sowing in Madrid. This work was funded by an EMBO Short Term Fellowship (M.E.-A.), ERC Advanced Grant IMMUNEMESIS and the Max Planck Society (D.W.). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Consortia
Contributions
M.E.-A., H.A.B. and D.W. conceived the project outline. M.E.-A. designed, implemented and coordinated the project. M.E.-A. carried out statistical analyses with advice from R.N. H.A.B., O.B., R.N. and D.W. supervised the project and discussed the interpretation of analyses. M.E.-A. prepared the first draft and the final manuscript was written by M.E.-A., H.A.B., O.B., R.N. and D.W. M.E.-A. carried out the experiment in Tübingen and in Madrid with technical support of the 500 Genomes Field Experiment Team. Specifically, M.E.-A. designed the field experiment with advice of D.W., H.A.B., O.B., R.S., F.G.-A., G.W. and F.V. M.E.-A. and R.S. coordinated logistics. M.E.-A., D.W., O.B., R.S. and F.G.-A. provided materials. M.E.-A. bulked seeds. M.E.-A., R.G.R., J.R. and R. Wedegärtner aliquoted seeds. M.E.-A., R.G.R. and R. Wedegärtner set up field sties. M.E.-A., R.G.R., R. Wedegärtner, F.W., P.L. and E.S. imaged plants. M.E.-A., R.G.R. and H.A.B. sowed seeds in Madrid. M.E.-A., F.V., D.L., R. Wedegärtner, D.K.S., B.R., P.L., J.K., R. Wu, W.X., K.V. and S.K. sowed seeds in Tübingen. M.E.-A., R.G.R., P.L., G.C., E.S., V.M., A.-L.V.d.W., J.D., D.T.N.T. and W.Z. thinned seedlings. M.E.-A., R.G.R., R. Wedegärtner and L.R. maintained field experiment. M.E.-A. conducted image processing. M.E.-A., B.R., P.L., R. Wu, K.V., G.C., E.S., S.K., W.Z. and M.Z. repaired the foil tunnel in Tübingen. M.E.-A., R.G.R., F.V., P.L., K.V., C.F., T.K. and C.B. harvested fresh material in Madrid and M.E.-A., D.W., R.S., D.L., R. Wedegärtner, B.R., P.L., R. Wu, K.V., G.C., E.S., V.M., S.P., E.D., C.F., L.R., C.G. and E.C. harvested fresh material in Tübingen. M.E.-A., R. Wedegärtner, V.M., C.G. and L.R. monitored flowering and photographed harvested material in Tübingen, and R.G.R. in Madrid. M.E.-A. carried out field data analysis. All authors commented on and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Tom Mitchell-Olds and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
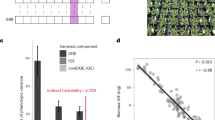
Extended Data Fig. 1 Field experiment setup and phenotyping.
a, View inside the foil tunnel in Germany. b, Spatial distribution of blocks and replicates in a split block design. Two independent blocks per watering condition were set up, with four spatially separated blocks within these two blocks. The 517 genotypes were then randomized within spatial blocks. The surface area of these blocks were the size of 18 or 26 quickpot trays for replicates with a high density of plants (a small population was grown per pot) or low density of plants (individual plants were grown per pot), respectively. In total, we placed 346 quickpot trays in Spain and 346 in Germany. Each quickpot tray had 40 cells of which 36 were used to sow plants (the corners were excluded). In total, 23,154 pots were successfully planted, which included about 14,500 pots with single plants (after thinning), and about 9,500 pots in which 30 seeds were planted and left to grow into small population without intervention. c, d, Soil water content (c) and soil surface temperature (d) retrieved from the 34 sensors monitoring each experimental block and the conditions outside the tunnel. Violin plots represent the distribution of recorded temperature and moisture values that were recorded every 10 min during the day and the night from three days after the day of sowing (16 November 2015 in Spain and 22 October 2015 in Germany) until the day at which all plants had completed their cycle and were dry (25 April 2016 in Spain and 15 April 2016 in Germany; n = 10,000 subsampled values of the time series). e, Set-up and examples of image-based high-throughput in situ phenotyping. (1) Customized dark box (Fotomatón) for image acquisition and (2) example tray with the corresponding green (3) and red (4) segmentation image products. Image-based monitoring was done for 23,154 pots, of which 375 pots were identified as failed replicates using the red flags placed in the field by experimenters (2 and 4). (5) Example image of a cut inflorescence of an adult plant and (6) segmentation of the inflorescence from background, (7) inflorescence path or skeletonization and (8) branch and end-point detection. In total, inflorescence images of 13,849 pots were taken and analysed. The variables extracted from inflorescence image processing (6–8) were used as predictors in a linear model that accurately estimated the number of fruits on an inflorescence, a relationship that was calibrated by manually counting fruits of small, medium and large representative plants (R2 = 0.97, n = 11, P = 10−4). a, e, All photographs were taken by M.E.-A.
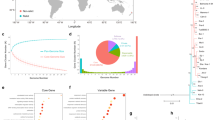
Extended Data Fig. 2 Geographical distributions of accessions and fitness values.
a, Locations of A. thaliana accessions used in this experiment (orange), 1001 Genomes accessions (blue) and all sightings of the species in GBIF (black, https://doi.org/10.15468/dl.c3twww). b, c, Geographical origin of the 502 Eurasian native A. thaliana lines used in this study and their raw fitness data (number of offspring produced in each pot) in the experiments in Spain with low precipitation (b) and in Germany with high precipitation (c). Note the most successful genotypes in the Spanish experiment (b) come not only from central Spain, but also from other areas of the distribution with extreme climates, including north Sweden, eastern Europe, the Caucasus and Siberia, which supports our previous observations8. Note also that for the German experiment (c), there could be multiple explanations for the visual trend that lower latitude genotypes had an apparent high fitness, namely that those Mediterranean genotypes are more diverse (some with higher and some with lower fitness values than average), or that the climate in Germany during 2015–2016 favoured genotypes from warmer areas. d, e, Idealized representation of distributions of alleles associated with fitness in Spain and Germany as inferred from genome-wide environmental niche models (see Supplementary Methods I.VI). The most-significant fitness-associated SNPs (if any) in each 0.5-Mb window of the genome were modelled (n = 414 in Spain, n = 279 in Germany). Colour scale indicates the percentage of locally present alleles with respect to the maximum number of positive fitness-related alleles identified in each experiment (maps were created using R v.3.4).
Extended Data Fig. 3 Simulation study of selection and genome-wide association model comparison.
Out of the A. thaliana genome matrix of 1,353,386 SNPs and 515 plants, we simulated that 1,000 randomly selected alleles were under natural selection (that is, the true selection coefficient drawn from a normal distribution around zero). Summing the selection coefficients that each of the 515 genotypes had based on their row in the genome matrix, we calculated their relative lifetime fitness. a, We then inferred the total selection coefficients s using a linear model as in Fig. 1 and compared them to the inferred estimates from a GBLUP-based genome-wide association (GWA) model (that is, a population-structure-corrected GWA), which we call direct selection coefficients in the main text (Supplementary Methods I.IV). Neither model infers the true simulated selection coefficients effectively. b, We were interested in studying the change in frequency of these 1,000 alleles after one generation of selection using individual-based simulations that sampled the genotypes for reproduction given their relative fitness. We compared the change in allele frequency in one generation (Δp = p1 − p0) with the inferred total selection coefficients and direct selection coefficients. The former, total selection coefficients, by summarizing direct and indirect effects of selection, perfectly coincide with the directionality of allele frequency changes. c, The total magnitude of allele frequency change also depends on the starting allele frequency of the allele under selection, which is described by the classic population genetics equation Δp = p(1 − p)s. Using our total selection coefficients and direct selection coefficients with this equation, we found an almost perfect relationship between predicted and realized allele frequency changes using total selection coefficients. d, To test whether the relationship in c can be extrapolated—that is, the relationship is not overly sensitive to differences in allele frequency and linkage disequilibrium in other A. thaliana subpopulations, we ran the individual-based simulations with a subset of 50 Spanish genotypes (out of the 515 genotypes) and repeated the comparison in c. The results indicate that total selection coefficients calculated from linear models are appropriate to understand the changes in allele frequencies in response to selection even when extrapolating to subpopulations with slightly different allele frequencies and linkage patterns. The regression statistics reported are: the adjusted R2, slope and regression P value, which were calculated using linear models of true compared with predicted values with n = 515. The 95% confidence interval of the regression is shaded in grey. (The code used to simulate and analyse these data can be found at https://doi.org/10.5281/zenodo.1408095.).
Extended Data Fig. 4 Genome maps of survival and seed set components of fitness.
Manhattan plots of SNPs significantly associated with fitness, using LM-GEMMA with the relative fitness averages of the 515 genotypes and 1,353,386 SNPs in 8 different environments. SNPs that were significant after multiple comparison correction using FDR (black and grey for alternating chromosomes) or Bonferroni (red) approaches are shown. a, Analyses using only the survival fitness component. b, Analyses using only the fecundity component.
Extended Data Fig. 5 Genome-wide environmental selection model testing and alternative projections.
a, Conceptual workflow of field validation procedure with data from published experiments25,26. This workflow was used for Fig. 3. b, Null expectation of predictability following the workflow shown in a with datasets randomizing fitness with genotypes. We could not find any model combination that had non-zero predictability (that is, all 95% bootstrap confidence intervals overlap with zero). c, Projections of selection intensity with real datasets as in Fig. 3 using different climate change scenarios available from WorldClim (http://worldclim.org/). The higher the predicted CO2 emissions, the stronger the predicted increase in selection intensity. rcp, representative concentration pathway. d, Map of predictions of selection change in 2050 as in Fig. 3 but representing the net number of local alleles increasing or decreasing in selection Total n = 10,752 SNPs. Only changes in s of more than 5% between present and future projections were considered.
Extended Data Fig. 6 Distribution of synonymous, nonsynonymous and neutral polymorphisms across space.
a, b, Fraction of all genome-wide nonsynonymous a and synonymous b mutations present in the local genotype (n = 502 locations). c, Ratio of nonsynonymous to synonymous fractions (Kn/Ks). d–f, Spearman’s ρ correlation of Kn/Ks with degrees longitude (d), degrees latitude (e) and precipitation in July (f) (n = 502 for all comparisons). High selection intensity (Fig. 3c) coincided with locations where natural lines have a lower-than-average ratio of nonsynonymous to synonymous polymorphisms (Spearman’s ρ = −0.276, P = 3 × 10−10; a, b), high local genetic diversity π (Spearman’s ρ = 0.187, P = 2.63×10−5; a, b) and elevated Tajima’s D (Spearman’s ρ = 0.161, P = 3 × 10−4; d–f and Fig. 3e) (see Supplementary Table 11). Various demographic scenarios could partially explain some of these patterns in isolation: bottlenecks can reduce the nonsynonymous polymorphisms because they are typically at low frequency, or high diversity might be found in old, large populations. However, all patterns are in agreement with stronger natural selection having acted more efficiently on nonsynonymous mutations in southern latitudes. In addition, high diversity could be driven by strong natural selection fluctuating over time, with alternative polymorphisms having been selected depending on interannual precipitation cycles (Fig. 3f). All in all, we did not find evidence that the warm edge of the geographical distribution of A. thaliana is formed by an increase in drift, which would cause small populations to accumulate nonsynonymous deleterious mutations and become genetically depauperate. Rather, our observations and predictions indicate that the species’ warm geographical limit is primarily defined by the environmental tolerance limits, where climate-driven natural selection limits the survival of individuals and populations outside the species range and only a few, highly specialized genotypes can survive near the range edges.
Supplementary information
Supplementary Information
This file contains Supplemental Methods: Appendix I and II.
Supplementary Tables
This file contains Supplementary Tables 1-11.
Supplementary Data
This file contains Supplementary Datasets 1-4.
Rights and permissions
About this article
Cite this article
Exposito-Alonso, M., 500 Genomes Field Experiment Team., Burbano, H.A. et al. Natural selection on the Arabidopsis thaliana genome in present and future climates. Nature 573, 126–129 (2019). https://doi.org/10.1038/s41586-019-1520-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1520-9
This article is cited by
-
Genomics for monitoring and understanding species responses to global climate change
Nature Reviews Genetics (2024)
-
Bottom-up perspective – The role of roots and rhizosphere in climate change adaptation and mitigation in agroecosystems
Plant and Soil (2024)
-
Re-focusing sampling, design and experimental methods to assess rapid evolution by non-native plant species
Biological Invasions (2024)
-
Pioneer Arabidopsis thaliana spans the succession gradient revealing a diverse root-associated microbiome
Environmental Microbiome (2023)
-
Genomic signals of local adaptation in Picea crassifolia
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.