Abstract
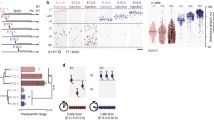
The diverse subtypes of excitatory neurons that populate the neocortex are born from apical progenitors located in the ventricular zone. During corticogenesis, apical progenitors sequentially generate deep-layer neurons followed by superficial-layer neurons directly or via the generation of intermediate progenitors. Whether neurogenic fate progression necessarily implies fate restriction in single progenitor types is unknown. Here we specifically isolated apical progenitors and intermediate progenitors, and fate-mapped their respective neuronal progeny following heterochronic transplantation into younger embryos. We find that apical progenitors are temporally plastic and can re-enter past molecular, electrophysiological and neurogenic states when exposed to an earlier-stage environment by sensing dynamic changes in extracellular Wnt. By contrast, intermediate progenitors are committed progenitors that lack such retrograde fate plasticity. These findings identify a diversity in the temporal plasticity of neocortical progenitors, revealing that some subtypes of cells can be untethered from their normal temporal progression to re-enter past developmental states.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets generated in this study are available in the GEO repository under accession number GSE122644.
Change history
15 April 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Jabaudon, D. Fate and freedom in developing neocortical circuits. Nat. Commun. 8, 16042 (2017).
Gaspard, N. et al. An intrinsic mechanism of corticogenesis from embryonic stem cells. Nature 455, 351–357 (2008).
Gao, P. et al. Deterministic progenitor behavior and unitary production of neurons in the neocortex. Cell 159, 775–788 (2014).
Okamoto, M. et al. Cell-cycle-independent transitions in temporal identity of mammalian neural progenitor cells. Nat. Commun. 7, 11349 (2016).
Yuzwa, S. A. et al. Developmental emergence of adult neural stem cells as revealed by single-cell transcriptional profiling. Cell Rep. 21, 3970–3986 (2017).
Mihalas, A. B. & Hevner, R. F. Clonal analysis reveals laminar fate multipotency and daughter cell apoptosis of mouse cortical intermediate progenitors. Development 145, dev164335 (2018).
Telley, L. et al. Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364, eaav2522 (2019).
Mihalas, A. B. et al. Intermediate progenitor cohorts differentially generate cortical layers and require Tbr2 for timely acquisition of neuronal subtype identity. Cell Rep. 16, 92–105 (2016).
Vasistha, N. A. et al. Cortical and clonal contribution of Tbr2 expressing progenitors in the developing mouse brain. Cereb. Cortex 25, 3290–3302 (2015).
Kowalczyk, T. et al. Intermediate neuronal progenitors (basal progenitors) produce pyramidal-projection neurons for all layers of cerebral cortex. Cerebral Cortex 19, 2439–2450 (2009).
Frantz, G. D. & McConnell, S. K. Restriction of late cerebral cortical progenitors to an upper-layer fate. Neuron 17, 55–61 (1996).
Desai, A. R. & McConnell, S. K. Progressive restriction in fate potential by neural progenitors during cerebral cortical development. Development 127, 2863–2872 (2000).
Telley, L. et al. Sequential transcriptional waves direct the differentiation of newborn neurons in the mouse neocortex. Science 351, 1443–1446 (2016).
Govindan, S., Oberst, P. & Jabaudon, D. In vivo pulse labeling of isochronic cohorts of cells in the central nervous system using FlashTag. Nat. Protoc. 13, 2297–2311 (2018).
Nagashima, F. et al. Novel and robust transplantation reveals the acquisition of polarized processes by cortical cells derived from mouse and human pluripotent stem cells. Stem Cells Dev. 23, 2129–2142 (2014).
Fishell, G. Striatal precursors adopt cortical identities in response to local cues. Development 121, 803–812 (1995).
Brüstle, O., Maskos, U. & McKay, R. D. Host-guided migration allows targeted introduction of neurons into the embryonic brain. Neuron 15, 1275–1285 (1995).
Cadwell, C. R. et al. Electrophysiological, transcriptomic and morphologic profiling of single neurons using Patch-seq. Nat. Biotechnol. 34, 199–203 (2016).
Fuzik, J. et al. Integration of electrophysiological recordings with single-cell RNA-seq data identifies neuronal subtypes. Nat. Biotechnol. 34, 175–183 (2016).
Kuwahara, A. et al. Tcf3 represses Wnt-β-catenin signaling and maintains neural stem cell population during neocortical development. PLoS ONE 9, e94408 (2014).
Pereira, J. D. et al. Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc. Natl Acad. Sci. USA 107, 15957–15962 (2010).
Aldiri, I., Moore, K. B., Hutcheson, D. A., Zhang, J. & Vetter, M. L. Polycomb repressive complex PRC2 regulates Xenopus retina development downstream of Wnt/β-catenin signaling. Development 140, 2867–2878 (2013).
Mutch, C. A., Funatsu, N., Monuki, E. S. & Chenn, A. β-catenin signaling levels in progenitors influence the laminar cell fates of projection neurons. J. Neurosci. 29, 13710–13719 (2009).
Vitali, I. et al. Progenitor Hyperpolarization Regulates the Sequential Generation of Neuronal Subtypes in the Developing Neocortex. Cell 174, 1264–1276 (2018).
Haubensak, W., Attardo, A., Denk, W. & Huttner, W. B. Neurons arise in the basal neuroepithelium of the early mammalian telencephalon: a major site of neurogenesis. Proc. Natl Acad. Sci. USA 101, 3196–3201 (2004).
Llorca, A. et al. Heterogeneous progenitor cell behaviors underlie the assembly of neocortical cytoarchitecture. Preprint at https://www.biorxiv.org/content/10.1101/494088v1 (2018).
Soufi, A. & Dalton, S. Cycling through developmental decisions: how cell cycle dynamics control pluripotency, differentiation and reprogramming. Development 143, 4301–4311 (2016).
Sela, Y., Molotski, N., Golan, S., Itskovitz-Eldor, J. & Soen, Y. Human embryonic stem cells exhibit increased propensity to differentiate during the G1 phase prior to phosphorylation of retinoblastoma protein. Stem Cells 30, 1097–1108 (2012).
Takahashi, T., Nowakowski, R. S. & Caviness, V. S. Jr. The cell cycle of the pseudostratified ventricular epithelium of the embryonic murine cerebral wall. J. Neurosci. 15, 6046–6057 (1995).
Reillo, I. & Borrell, V. Germinal zones in the developing cerebral cortex of ferret: ontogeny, cell cycle kinetics, and diversity of progenitors. Cereb. Cortex 22, 2039–2054 (2012).
Naujok, O., Lentes, J., Diekmann, U., Davenport, C. & Lenzen, S. Cytotoxicity and activation of the Wnt/beta-catenin pathway in mouse embryonic stem cells treated with four GSK3 inhibitors. BMC Res. Notes 7, 273 (2014).
Kohwi, M., Lupton, J. R., Lai, S.-L., Miller, M. R. & Doe, C. Q. Developmentally regulated subnuclear genome reorganization restricts neural progenitor competence in Drosophila. Cell 152, 97–108 (2013).
Molyneaux, B. J., Arlotta, P., Hirata, T., Hibi, M. & Macklis, J. D. Fezl is required for the birth and specification of corticospinal motor neurons. Neuron 47, 817–831 (2005).
Hanashima, C., Li, S. C., Shen, L., Lai, E. & Fishell, G. Foxg1 suppresses early cortical cell fate. Science 303, 56–59 (2004).
Mizutani, K., Yoon, K., Dang, L., Tokunaga, A. & Gaiano, N. Differential Notch signalling distinguishes neural stem cells from intermediate progenitors. Nature 449, 351–355 (2007).
Knobloch, M. et al. A fatty acid oxidation-dependent metabolic shift regulates adult neural stem cell activity. Cell Rep. 20, 2144–2155 (2017).
Toma, K., Kumamoto, T. & Hanashima, C. The timing of upper-layer neurogenesis is conferred by sequential derepression and negative feedback from deep-layer neurons. J. Neurosci. 34, 13259–13276 (2014).
Seuntjens, E. et al. Sip1 regulates sequential fate decisions by feedback signaling from postmitotic neurons to progenitors. Nat. Neurosci. 12, 1373–1380 (2009).
Long, J. Z., Lackan, C. S. & Hadjantonakis, A.-K. Genetic and spectrally distinct in vivo imaging: embryonic stem cells and mice with widespread expression of a monomeric red fluorescent protein. BMC Biotechnol. 5, 20 (2005).
Chen, F. & LoTurco, J. A method for stable transgenesis of radial glia lineage in rat neocortex by piggyBac mediated transposition. J. Neurosci. Methods 207, 172–180 (2012).
Bocchi, R. et al. Perturbed Wnt signaling leads to neuronal migration delay, altered interhemispheric connections and impaired social behavior. Nat. Commun. 8, 1158 (2017).
Teo, C. H., Vishwanathan, S. V. N. & Smola, A. Bundle methods for regularized risk minimization. J. Mach. Learn. Res. 11, 311–365 (2010).
Acknowledgements
We thank members of the Jabaudon laboratory and E. Azim for comments on the manuscript. We thank O. Raineteau for providing plasmids. We thank A. S. Lopes, A. Benoit, the FACS facility, the Genomics Platform and the Bioimaging Facility of the University of Geneva for technical assistance. Work in the Jabaudon laboratory is supported by the Swiss National Science Foundation, the Fondation des HUG, and the Carigest Foundation. P.O. is supported by a fellowship from iGE3.
Author information
Authors and Affiliations
Contributions
P.O. and D.J. designed the experiments. P.O. performed the experiments with the help of C.C. and G.B.; S.F. performed the electrophysiology experiments and the collection of cells for Patch-seq with the help of P.O.; N.B. performed the bioinformatic analysis; P.O. and D.J. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
: The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks André Goffinet and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Donor APs rapidly integrate the VZ and behave normally.
a, FACS of FlashTag+ cells 1 h after labelling. b, Donor APs integrate the host VZ at discrete sites. Only a few APs (median = 2) are present at each site, such that daughter neurons found at single integration sites at P7 are probably born from a small number of initial APs (oligoclonal analysis, Figs. 1g, 4d, Extended Data Fig. 8b). c, Left, photomicrograph of a transplanted AP showing a radial glia morphology (maximum projection). Right, juxtaventricular mitosis in a transplanted AP. Experiments in a–c were repeated at least three times with similar results. d, The progeny of transplanted APs progressively migrate towards the cortical plate. The time course of this migration is similar to that of the host cells, as assessed by comparison with the migration of FlashTag-labelled endogenous cells. Box plots show median and interquartile range. n = 46 cells (RFP 6 h), 21 cells (RFP 12 h), 30 cells (RFP 24 h), 30 cells (RFP 48 h), 31 cells (RFP 72 h), 36 cells (FlashTag 6 h), 25 cells (FlashTag 12 h), 33 cells (FlashTag 24 h), 39 cells (FlashTag 48 h), 31 cells (FlashTag 72 h).
Extended Data Fig. 2 Analysis of single integration sites in AP15→15 and AP12→12—the transplantation procedure does not affect the neurogenic competence of APs.
a, Isochronically transplanted E15.5 APs (AP15→15) essentially generate SL neurons. Photomicrographs: within each donor litter, illustrations are clustered by integration site. Where applicable, a vertical black line delineates distinct host pups within a given litter. Experiments were repeated eight times with similar results. b, Isochronically transplanted E12.5 APs (AP12→12) generate DL and SL neurons. Photomicrographs: within each donor litter, illustrations are clustered by integration site. Where applicable, a vertical black line delineates distinct host pups within a given litter. Experiments were repeated six times with similar results. c, The laminar distribution of daughter neurons in the AP15→15 condition is replicated by in utero electroporation of a piggyBac-transposon construct at E15.5, in the absence of transplantation. d, The laminar distribution of daughter neurons in the AP12→12 condition is replicated by in utero electroporation of a piggyBac-transposon construct at E12.5. c, d, Box plots show median and interquartile range. In bar graphs, values are shown as mean ± s.e.m.; n = 5 pups (EporPB15), 7 experimental litters (AP15→15), 3 pups (EporPB12), 6 experimental litters (AP12→12). Two-way ANOVA with post hoc Tukey test. Epor, in utero electroporation. AP15→15 and AP12→12 distribution plots are reproduced from Fig. 1.
Extended Data Fig. 3 Analysis of single integration sites in AP15→12 and lack of temporal respecification in gliogenic AP17→12.
a, E15.5 APs transplanted into an E12.5 host (AP15→12) generate DL and SL neurons. Photomicrographs: within each donor litter, illustrations are clustered by integration site. Where applicable, a vertical black line delineates distinct host pups within a given litter. Experiments were repeated six times with similar results. b, Laminar distribution of daughter neurons across conditions. AP15→15 and AP12→12 distribution plots are reproduced from Extended Data Fig. 2 to enable direct comparison across conditions. Values are shown as mean ± s.e.m.; n = 7 experimental litters (AP15→15), 6 experimental litters (AP12→12), 8 experimental litters (AP15→12). Two-way ANOVA with post hoc Tukey test. c, E17.5 APs transplanted into an E12.5 host (AP17→12) still almost exclusively generate glial cells. Arrowheads show examples of GFAP+ cells. Values are shown as mean ± s.e.m.; n = 3 experimental litters per group. Two-tailed t-test. ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 4 Molecular markers of AP15→12 daughter neurons and approach used to identify transplanted nascent neurons.
a, Reduced expression of the SL marker CUX1 in DL AP15→12 daughter neurons. b, Increased expression of the DL marker CTIP2 in DL AP15→12 daughter neurons. c, Increased expression of the DL marker TBR1 in DL AP15→12 daughter neurons. Photomicrographs showing pattern of expression of CUX1, CTIP2 and TBR1 are reproduced from Fig. 2. AP15→12 distribution plots in a–c have been reproduced from Fig. 2 for direct comparison. d, Left, schematic of the chronic EdU-labelling approach used to distinguish between nascent donor neurons and neurons born in the host. Centre, photomicrograph showing examples of an EdU+ and an EdU− donor neuron. Right, quantification of the fraction of EdU− labelled neurons at P7 (that is, transplanted cells that never underwent division in the host). Experiments were repeated three times with similar results. e, Heterochronically transplanted E15.5 nascent neurons (N15→12) migrate to the superficial layers, as they would have done in their original host. Values are shown as mean ± s.e.m.; n = 3 experimental litters (N15→12), 8 experimental litters (AP15→12). AP15→12 data in e have been reproduced from Fig. 1f for direct comparison. M, mitosis; S, S phase.
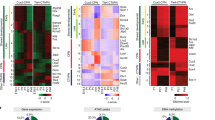
Extended Data Fig. 5 Repression of AP15-type transcriptional programs and re-induction of AP12-type transcriptional programs in AP15→12.
a, SVM classification of AP15 and AP15→12. Box plots indicate median and interquartile range. n = 26 cells (AP15), 19 cells (AP15→12). b, Expression of the AP15 transcripts used in the model. c, Expression of the AP15→12 transcripts used in the model. In b, c, n = 20 cells (AP12), 26 cells (AP15), 19 cells (AP15→12). d, Examples of hybridizations from the Allen Developing Mouse Brain Atlas (© 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas. Available from: http://developingmouse.brain-map.org/) corresponding to genes shown in Fig. 3d. e, The expression of potassium channels is repressed in AP15→12. n = 20 cells (AP12), 26 cells (AP15), 19 cells (AP15→12).
Extended Data Fig. 6 Lack of temporal respecification of AP12→15.
a, Parameters used to define AP12→15 identity. Letters refer to panels in this figure. b, The resting membrane potential of AP12→15 remains at AP12 values. Box plots indicate median and interquartile range; n = 27 cells (AP15), 21 cells (AP12), 12 cells (AP12→15). Kruskal–Wallis test with post hoc Dunn’s test. c, Neurogenic divisions in AP12→15 remain at AP12 values. n = 3 experimental litters per group. One-way ANOVA with post hoc Tukey test. d, Number of cells per integration site at P7. Each point represents one oligoclone. The oligoclone size of AP12→15 remains at AP12→12 values. n = 37 oligoclones (AP15→15), 54 oligoclones (AP12→12), 78 oligoclones (AP12→15). Kruskal–Wallis test with post hoc Dunn’s test. AP15, AP12, AP12→12 and AP15→15 data in b–d have been reproduced from Fig. 3 for direct comparison. e, AP12→15 still generate CTIP2+ daughter neurons. n = 3 experimental litters per group. f, AP12→15 still generate TBR1+ daughter neurons. n = 3 experimental litters (AP15→15 and AP12→12), 2 experimental litters (AP12→15). g, Mismigration of CUX1-expressing daughter neurons into DL in AP12→15. n = 3 experimental litters (AP12→12), 4 experimental litters (AP12→15). Photomicrographs showing pattern of expression of CUX1, CTIP2 and TBR2 are reproduced from Fig. 2. AP12→12 and AP15→15 data in e–g have been reproduced from Fig. 2 for direct comparison. Data in c–g are shown as mean ± s.e.m. One-way ANOVA with post hoc Tukey test (e–g). h, EdU pulse labelling at E17 and E18 shows progressive mismigration of increasingly later-born neurons in AP12→15. Box plots show mean and interquartile range. n = 19 cells (E17), 12 cells (E18). i, Summary of the mismigration phenotype in AP12→15. **P < 0.01, ***P < 0.001.
Extended Data Fig. 7 Analysis of single integration sites in AP12→15.
E12.5 APs transplanted into an E15.5 host (AP12→15) generate DL and SL neurons. Photomicrographs: within each donor litter, illustrations are clustered by integration site. Where applicable, a vertical black line delineates distinct host pups within a given litter. Experiments were repeated six times with similar results.
Extended Data Fig. 8 Heterochronically transplanted IPs (IP15→12) generate SL neurons.
a, Schematic showing EdU-based labelling and transplantation procedure. b, E15.5 EdU+ cells transplanted into an E12.5 host (EdU15→12) still give rise to SL neurons (compare with Fig. 1d). Box plot shows median and interquartile range. Top right, modal distribution of daughter neurons in SL vs DL. Values are shown as mean ± s.e.m.; n = 3 experimental litters. Bottom right, laminar distribution of daughter neurons at single integration sites. Values refer to number of integration sites in each category. c, Heterochronically transplanted EdU-labelled progenitors (EdU15→12) essentially generate SL neurons. Photomicrographs: within each donor litter, illustrations are clustered by integration site. Experiments were repeated three times with similar results. d, Ten hours after FlashTag labelling, most cells have differentiated into IPs (that is, KI67+TBR2+ cells). Values are shown as mean ± s.e.m.; n = 3 pups. e, IP15→12 essentially give rise to SL neurons. Photomicrographs: within each donor litter, illustrations are clustered by integration site. Only EdU+ neurons (filled arrowheads) were included in this analysis. Experiments were repeated three times with similar results. f, Number of cells per integration site at P7. Each point represents one oligoclone. AP15→12 data has been reproduced from Fig. 3g for direct comparison. Values are shown as mean ± s.e.m.; n = 39 oligoclones (AP15→12), 17 oligoclones (IP15→12). g, IP15→12 daughter neurons express CUX1 but not CTIP2. Photomicrographs showing pattern of expression of CUX1 and CTIP2 are reproduced from Fig. 2. Values are shown as mean ± s.e.m.; N = 2 experimental litters (CUX1), 3 experimental litters (CTIP2). In b, vertically aligned cells belong to single integration sites. FT, FlashTag; IZ, intermediate zone.
Extended Data Fig. 9 Temporal dynamics of Wnt signalling in the developing cortex.
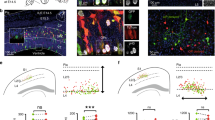
a, Top, temporally dynamic expression of Wnt transcripts in the developing cortex (data from www.unige.ch/genebrowser.unige.ch/telagirdon)8. Values are shown as mean ± s.e.m.; n = 189 APs, 268 neurons (E12), 207 APs, 223 neurons (E13), 134 APs, 219 neurons (E14), 301 APs, 213 neurons (E15). Right, example of corresponding in situ hybridizations from the Allen Developing Mouse Brain Atlas (© 2008 Allen Institute for Brain Science. Allen Developing Mouse Brain Atlas. Available from: http://developingmouse.brain-map.org/). b, Dynamic expression of Wnt transcripts in the developing cortex (expression landscapes from www.unige.ch/genebrowser.unige.ch/telagirdon)8. The overall pattern is early–high to late–low (a). c, AP15 express Wnt receptor-related transcriptional programs (data from ref. 8). Box plots indicate median and interquartile range. Right, expression of individual receptor-related transcripts in AP12 and AP15. n = 189 APs (E12), 301 APs (E15).
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-3, Discussion and References.
Supplementary Table
Supplementary Table 1| Source data of radial position, laminar location and oligoclone identity of all neurons in this study. This file contains the values for the normalized radial position, laminar location and oligoclone identity of all the neurons used for analysis in the following conditions: AP15→15, AP12→12, AP15→12, EdU15→12, IP15→12 and AP12→15.
Rights and permissions
About this article
Cite this article
Oberst, P., Fièvre, S., Baumann, N. et al. Temporal plasticity of apical progenitors in the developing mouse neocortex. Nature 573, 370–374 (2019). https://doi.org/10.1038/s41586-019-1515-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1515-6
This article is cited by
-
Integrating single-cell transcriptomics with cellular phenotypes: cell morphology, Ca2+ imaging and electrophysiology
Biophysical Reviews (2024)
-
Patch-seq: Advances and Biological Applications
Cellular and Molecular Neurobiology (2024)
-
A Spacetime Odyssey of Neural Progenitors to Generate Neuronal Diversity
Neuroscience Bulletin (2023)
-
Application of Lineage Tracing in Central Nervous System Development and Regeneration
Molecular Biotechnology (2023)
-
Loss of neurodevelopmental-associated miR-592 impairs neurogenesis and causes social interaction deficits
Cell Death & Disease (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.