Abstract
Elucidating the cellular architecture of the human cerebral cortex is central to understanding our cognitive abilities and susceptibility to disease. Here we used single-nucleus RNA-sequencing analysis to perform a comprehensive study of cell types in the middle temporal gyrus of human cortex. We identified a highly diverse set of excitatory and inhibitory neuron types that are mostly sparse, with excitatory types being less layer-restricted than expected. Comparison to similar mouse cortex single-cell RNA-sequencing datasets revealed a surprisingly well-conserved cellular architecture that enables matching of homologous types and predictions of properties of human cell types. Despite this general conservation, we also found extensive differences between homologous human and mouse cell types, including marked alterations in proportions, laminar distributions, gene expression and morphology. These species-specific features emphasize the importance of directly studying human brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data can be accessed through the Allen Brain Atlas data portal at http://portal.brain-map.org/ and RNA-seq data from this study are publicly available and can be downloaded at http://celltypes.brain-map.org/. Data can be visualized and analysed using two complementary viewers: the RNA-seq Data Navigator (http://celltypes.brain-map.org/rnaseq/human) and the Cytosplore Viewer (https://viewer.cytosplore.org/), an extension of Cytosplore66 that presents a hierarchy of t-SNE maps of different subsets of MTG clusters67. An ontology of cell types can be navigated at http://bioportal.bioontology.org/ontologies/PCL. Controlled access raw RNA-seq data are registered with dbGAP (accession number phs001790) and have been deposited at the NeMO archive (https://nemoarchive.org/). Applications to access raw sequencing data should be submitted via dbGAP.
Code availability
The data and code used to produce figures are available from https://github.com/AllenInstitute/MTG_celltypes.
References
Glasser, M. F. et al. A multi-modal parcellation of human cerebral cortex. Nature 536, 171–178 (2016).
Azevedo, F. A. C. et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 513, 532–541 (2009).
Herculano-Houzel, S., Mota, B. & Lent, R. Cellular scaling rules for rodent brains. Proc. Natl Acad. Sci. USA 103, 12138–12143 (2006).
DeFelipe, J. The evolution of the brain, the human nature of cortical circuits, and intellectual creativity. Front. Neuroanat. 5, 29 (2011).
Poorthuis, R. B. et al. Rapid neuromodulation of layer 1 interneurons in human neocortex. Cell Rep. 23, 951–958 (2018).
Eyal, G. et al. Unique membrane properties and enhanced signal processing in human neocortical neurons. eLife 5, e16553 (2016).
Szegedi, V. et al. Plasticity in single axon glutamatergic connection to GABAergic interneurons regulates complex events in the human neocortex. PLoS Biol. 14, e2000237 (2016).
Benavides-Piccione, R., Ballesteros-Yáñez, I., DeFelipe, J. & Yuste, R. Cortical area and species differences in dendritic spine morphology. J. Neurocytol. 31, 337–346 (2002).
Gabbott, P. L. Subpial fan cell—a class of calretinin neuron in layer 1 of adult monkey prefrontal cortex. Front. Neuroanat. 10, 28 (2016).
Ramón y Cajal, S. La Textura del Sistema Nerviosa del Hombre y los Vertebrados (Nicolas Moya, 1904).
Lorente de Nó, R. La corteza cerebral del ratón. Trab. Lab. Invest. Bio. (Madrid) 20, 41–78 (1922).
Hill, R. S. & Walsh, C. A. Molecular insights into human brain evolution. Nature 437, 64–67 (2005).
Oberheim, N. A. et al. Uniquely hominid features of adult human astrocytes. J. Neurosci. 29, 3276–3287 (2009).
Boldog, E. et al. Transcriptomic and morphophysiological evidence for a specialized human cortical GABAergic cell type. Nat. Neurosci. 21, 1185–1195 (2018).
Zeng, H. et al. Large-scale cellular-resolution gene profiling in human neocortex reveals species-specific molecular signatures. Cell 149, 483–496 (2012).
Bakken, T. E. et al. A comprehensive transcriptional map of primate brain development. Nature 535, 367–375 (2016).
Hawrylycz, M. et al. Canonical genetic signatures of the adult human brain. Nat. Neurosci. 18, 1832–1844 (2015).
Ecker, J. R. et al. The BRAIN initiative cell census consortium: lessons learned toward generating a comprehensive Brain Cell Atlas. Neuron 96, 542–557 (2017).
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Krishnaswami, S. R. et al. Using single nuclei for RNA-seq to capture the transcriptome of postmortem neurons. Nat. Protoc. 11, 499–524 (2016).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Lake, B. B. et al. A comparative strategy for single-nucleus and single-cell transcriptomes confirms accuracy in predicted cell-type expression from nuclear RNA. Sci. Rep. 7, 6031 (2017).
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS ONE 13, e0209648 (2018).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Zhu, Y., Wang, L., Yin, Y. & Yang, E. Systematic analysis of gene expression patterns associated with postmortem interval in human tissues. Sci. Rep. 7, 5435 (2017).
Bakken, T. et al. Cell type discovery and representation in the era of high-content single cell phenotyping. BMC Bioinformatics 18, 559 (2017).
Werner, M. S. et al. Chromatin-enriched lncRNAs can act as cell-type specific activators of proximal gene transcription. Nat. Struct. Mol. Biol. 24, 596–603 (2017).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Liu, S. J. et al. Single-cell analysis of long non-coding RNAs in the developing human neocortex. Genome Biol. 17, 67 (2016).
von Economo, C. Cellular structure of the human cerebral cortex. (Karger, 2009).
Kalmbach, B. E. et al. h-Channels contribute to divergent intrinsic membrane properties of supragranular pyramidal neurons in human versus mouse cerebral cortex. Neuron 100, 1194–1208 (2018).
Hansen, D. V. et al. Non-epithelial stem cells and cortical interneuron production in the human ganglionic eminences. Nat. Neurosci. 16, 1576–1587 (2013).
Ma, T. et al. Subcortical origins of human and monkey neocortical interneurons. Nat. Neurosci. 16, 1588–1597 (2013).
Lee, S., Hjerling-Leffler, J., Zagha, E., Fishell, G. & Rudy, B. The largest group of superficial neocortical GABAergic interneurons expresses ionotropic serotonin receptors. J. Neurosci. 30, 16796–16808 (2010).
Raghanti, M. A. et al. Neuropeptide Y-immunoreactive neurons in the cerebral cortex of humans and other haplorrhine primates. Am. J. Primatol. 75, 415–424 (2013).
Xu, X., Roby, K. D. & Callaway, E. M. Immunochemical characterization of inhibitory mouse cortical neurons: three chemically distinct classes of inhibitory cells. J. Comp. Neurol. 518, 389–404 (2010).
Paul, A. et al. Transcriptional architecture of synaptic communication delineates GABAergic neuron identity. Cell 171, 522–539 (2017).
Miyoshi, G. et al. Genetic fate mapping reveals that the caudal ganglionic eminence produces a large and diverse population of superficial cortical interneurons. J. Neurosci. 30, 1582–1594 (2010).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Johansen, N. & Quon, G. scAlign: a tool for alignment, integration and rare cell identification from scRNA-seq data. Genome Biol. 20, 166 (2019).
Kilduff, T. S., Cauli, B. & Gerashchenko, D. Activation of cortical interneurons during sleep: an anatomical link to homeostatic sleep regulation? Trends Neurosci. 34, 10–19 (2011).
Belichenko, P. V., Vogt Weisenhorn, D. M., Myklóssy, J. & Celio, M. R. Calretinin-positive Cajal-Retzius cells persist in the adult human neocortex. Neuroreport 6, 1869–1874 (1995).
Sorensen, S. A. et al. Correlated gene expression and target specificity demonstrate excitatory projection neuron diversity. Cereb. Cortex 25, 433–449 (2015).
Lin, Y. et al. Evaluating stably expressed genes in single cells. Preprint at https://doi.org/10.1101/229815 (2018).
Colantuoni, C. et al. Temporal dynamics and genetic control of transcription in the human prefrontal cortex. Nature 478, 519–523 (2011).
Foley, N. M., Springer, M. S. & Teeling, E. C. Mammal madness: is the mammal tree of life not yet resolved? Phil. Trans. R. Soc. Lond. B 371, 20150140 (2016).
Markou, A., Chiamulera, C., Geyer, M. A., Tricklebank, M. & Steckler, T. Removing obstacles in neuroscience drug discovery: the future path for animal models. Neuropsychopharmacology 34, 74–89 (2009).
Nestler, E. J. & Hyman, S. E. Animal models of neuropsychiatric disorders. Nat. Neurosci. 13, 1161–1169 (2010).
DeFelipe, J., Alonso-Nanclares, L. & Arellano, J. I. Microstructure of the neocortex: comparative aspects. J. Neurocytol. 31, 299–316 (2002).
Aronesty, E. Comparison of sequencing utility programs. Open Bioinform. J. 7, 1–8 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Calvo, S. E., Clauser, K. R. & Mootha, V. K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, D1251–D1257 (2016).
Fortunato, S. & Barthélemy, M. Resolution limit in community detection. Proc. Natl Acad. Sci. USA 104, 36–41 (2007).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Aevermann, B. D. et al. Cell type discovery using single-cell transcriptomics: implications for ontological representation. Hum. Mol. Genet. 27, R40–R47 (2018).
Yanai, I. et al. Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification. Bioinformatics 21, 650–659 (2005).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Lyubimova, A. et al. Single-molecule mRNA detection and counting in mammalian tissue. Nat. Protocols 8, 1743–1758 (2013).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Höllt, T. et al. Cytosplore: Interactive immune cell phenotyping for large single-cell datasets. Comput. Graph. Forum 35, 171–180 (2016).
Höllt, T. et al. CyteGuide: Visual guidance for hierarchical single-cell analysis. IEEE Trans. Vis. Comput. Graph. 24, 739–748 (2018).
Acknowledgements
We thank the Tissue Procurement, Tissue Processing and Facilities teams at the Allen Institute for Brain Science for assistance with the transport and processing of postmortem and neurosurgical brain specimens; the Technology team at the Allen Institute for assistance with data management; our collaborators at the Swedish Medical Center and Harborview Medical Center in Seattle for coordinating human neurosurgical tissue collections; M. Vawter, J. Davis and the San Diego Medical Examiner’s Office for assistance with postmortem tissue donations; and the Molecular Biology, Histology and Imaging teams at the Allen Institute for Brain Science for performing chromogenic in situ hybridization experiments. A. M. Yanny provided technical assistance with RNAscope experiments. This work was funded by the Allen Institute for Brain Science and by US National Institutes of Health grant U01 MH114812-02 to E.S.L. Funding from NWO-AES projects 12721: Genes in Space and 12720: VANPIR (principal investigator A. Vilanova) for development of the Cytosplore Viewer is gratefully acknowledged. We thank B. van Lew for scripting and narration of Cytosplore instructional and use-case videos. Support for the development of NS-Forest v.2 and the provisional cell ontology was provided by the Chan–Zuckerberg Initiative DAF, an advised fund of the Silicon Valley Community Foundation (2018-182730). G.Q. is supported by NSF CAREER award 1846559. This publication is part of the Human Cell Atlas (https://www.humancellatlas.org/publications). The authors thank the Allen Institute founder, Paul G. Allen, for his vision, encouragement and support.
Author information
Authors and Affiliations
Contributions
E.S.L. conceptualized and supervised the study. E.S.L. and R.Y. conceptualized the Human Cell Types Program. R.D.H. and T.E.B. designed experiments. R.D.H., E.R.B., B. Long., J.L.C., B.P.L., S.I.S., K.B., J.G., D.H., S.-L.D., M.M., S.P., E.R.T., N.V.S., E.G., T.N.N. and Z.M. contributed to nuclei isolation and/or validation experiments. T.E.B. and J.A.M. analysed the data with contributions from N.J., O.P., Z.Y., O.F., J.G., S.S., G.Q. and M.H. K.A.S. and B.T. managed the snRNA-seq pipeline. L.T.G. developed data visualization tools. D.B., K.L., C.R. and M.T. performed snRNA-seq. A. Bernard and J.W.P. managed establishment of the snRNA-seq pipeline. A. Bernard and M.M. contributed to the development and management of histological methods and data generation. R.D., N.D., T.C., J.N. and A.O. processed postmortem brain tissues. A. Bernard and N.D. managed acquisition of postmortem and neurosurgical tissues. A. Beller, C.D.K., C.C., R.G.E., R.P.G., A.L.K. and J.G.O. contributed to neurosurgical tissue collections. B.A., M.K. and R.H.S. developed the semantic representation of clusters. J.E., T.H., A.M. and B. Lelieveldt developed the Cytosplore Viewer. L.T.G., J.A.M., D.F., L.N. and A. Bernard contributed to the development of the RNA-Seq Data Navigator. S.R., A.S. and S.M.S. provided programme management and/or regulatory compliance support. C.K. and A.R.J. provided institutional support and project oversight. E.S.L. and H.Z. directed the Allen Institute Cell Types Program. R.D.H., T.E.B. and E.S.L. wrote the paper with contributions from J.A.M. and J.L.C. and in consultation with all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Thomas Mrsic-Flogel, Rahul Satija and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Nuclei metadata summarized by cluster.
a, FACS gating scheme for nuclei sorts. b, FACS metadata for index-sorted single nuclei (n = 571) show significant variability in NeuN fluorescence intensity (NeuN–PE-A), size (forward-scatter area, FSC-A) and granularity (side-scatter area, SSC-A) across clusters. As expected, non-neuronal nuclei have almost no NeuN staining and are smaller (as inferred by lower FSC values). Error bars represent 95% bootstrapped confidence intervals on mean values (points). c, d, Scatter plots of single nuclei from all clusters (n = 15,928) plus median and interquartile interval of three quality control metrics grouped and coloured by cluster. c, Median total reads were approximately 2.6 million for all cell types, although slightly lower for non-neuronal nuclei. d, Median gene detection was highest among excitatory neuron types in L5 and L6 and a subset of types in L3, lower among inhibitory neuron types and significantly lower among non-neuronal types.
Extended Data Fig. 2 Small but consistent expression signature of donor tissue source.
a, mRNA quality was only slightly higher for nuclei isolated from neurosurgical (n = 722) versus postmortem (n = 15,206) donors (~3% more uniquely aligned reads and ~350 more genes detected). All nuclei were dissected from cortical L5 and sorted on the basis of NeuN-positive staining, and transcripts were sequenced to a median depth of approximately 2.5 million reads per nucleus. Median values (red points) and interquartile interval are indicated. b, Dot plot showing the proportion of nuclei isolated from neurosurgical and postmortem donors among human MTG clusters. Note that most nuclei from neurosurgical donors were isolated only from L5, so clusters enriched in other layers, such as L1 interneurons, have low representation of these donors. c, Highly correlated (Pearson’s) expression between nuclei from postmortem and neurosurgical donors among two subclasses of excitatory neurons and one subclass of inhibitory neurons. Nuclei were pooled and compared within these subclasses owing to the low sampling of individual clusters from neurosurgical donors. Average expression of n = 2,180, 1,636 and 815 postmortem nuclei and 127, 38 and 114 neurosurgical nuclei were included for the L5a excitatory, L4 excitatory and SST+ interneuron comparisons, respectively. d, Expression (log10(CPM + 1)) heat maps of the top-10 upregulated genes in nuclei from postmortem and neurosurgical donors including ribosomal genes and activity-dependent genes, respectively.
Extended Data Fig. 3 Cluster robustness.
a, Cluster separability (mean co-clustering within a cluster minus the maximum co-clustering between clusters) varied substantially among cell types (n = 15,928 nuclei), with a subset of neuronal types and all non-neuronal types being highly discrete. b, Scatter plots quantifying the separation of each cluster from its nearest neighbour. Left, cluster separability based on rounds of iterative clustering using all variable genes is correlated with the number of binary marker genes. Middle, all clusters express at least 30 genes with >twofold increased expression, but only a subset are binary markers. Right, A substantial fraction of markers of many clusters are unannotated. c, River plots of clusters that merge with more binary markers required for separation. Note that clusters that appear distinct on the basis of layer position (excitatory neurons in L2 and L3), morphology (interlaminar astroctyes in L1) or homology with mouse (SST+ interneuron subtypes) can have few binary markers. Marker genes for clusters defined by four markers are listed in Supplementary Table 2. d, Confusion plots comparing cluster membership of single nuclei (n = 15,928) in reference MTG clusters and clusters generated using a different iterative clustering pipeline. Above each plot are listed the parameter settings and total number of clusters detected. Point size is proportional to the number of nuclei and point colour corresponds to the Jaccard index, with darker colours corresponding to a higher Jaccard index and greater consistency between clustering. e, Box plots summarizing consistency of cluster membership of single nuclei (n = 15,928) across the four iterative clustering runs shown in c. Box plots show median, interquartile interval and full range of values. Top, the number of clusters that overlap each reference cluster. A cluster count of 1 indicates a one-to-one match, 0 indicates that a reference cluster was not detected and was merged with a related cluster, and >1 indicates that a reference cluster was split into sub-clusters. *The Exc L2−3 LINC00507 FREM3 reference cluster was consistently divided into subclusters. Bottom, reference clusters with higher Jaccard index values have more consistent membership of nuclei and therefore more distinct borders with related clusters. f, Violin plots of marker-gene expression for FREM3 subclusters (n = 2,284 nuclei) identified in one clustering run show relatively binary expression. In the violin plot, rows are genes and black dots correspond to median expression. Expression values are on a linear scale.
Extended Data Fig. 4 Expression of cell-type-specific markers.
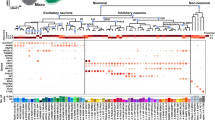
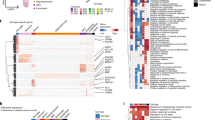
a, b, Heat maps of the top cell-type markers for inhibitory neurons (a) and excitatory neurons and non-neuronal cell types (b). Markers include many non-coding and unannotated genes (blue). Median expression values are shown on a logarithmic scale, with maximum expression values shown on the right side of each row. Up to five marker genes are shown for each cell type. Note that LOC genes were excluded from cluster names, and the best non-LOC marker genes were used instead. Dendrograms and cluster names are reproduced from Fig. 1. Marker genes for broad classes, as defined manually and using NS-Forest, are also shown in the top rows of each heat map.
Extended Data Fig. 5 Clusters in this study capture reported human cortical cell types and additional subtype diversity.
a–c, Dot plots showing the proportion of each MTG cluster that matches reported clusters based on a centroid expression classifier. a, Three of sixteen neuronal clusters reported by Lake et al.24 (n = 3,042 nuclei) match human MTG clusters one-to-one, and the remaining clusters map to multiple MTG clusters. *Ex3 was highly enriched in visual cortex and not detected in temporal cortex by Lake et al. b, Four of eighteen neuronal clusters and three of four non-neuronal clusters reported by Lake et al.27 (n = 10,319 nuclei) match human MTG clusters one-to-one, including two rare but distinct interneuron types (Inh L3–6 SST NPY and Inh L2–5 PVALB SCUBE3) and one rare but distinct excitatory type (Exc L4–5 FEZF2 SCN4B). c, Four neuronal clusters reported by Habib et al.28 (n = 5,433 nuclei) correspond to broad classes of inhibitory and excitatory neurons. Seven non-neuronal clusters include two astrocyte types that correspond to the types reported in this study, and one additional oligodendrocyte subtype. d, Sixteen clusters detected in L1 of human temporal cortex14 (n = 914 nuclei) are captured at finer subtype resolution in this study.
Extended Data Fig. 6 Excitatory neuron types express marker genes across multiple cortical layers.
a, Constellation diagram showing cluster relationships, relative frequencies and average layer position. b–e, Heat maps of log-transformed expression in individual nuclei ordered by cluster and then layer. Clusters are grouped on the basis of their dominant class-marker gene, which corresponds to position in superficial (LAMP5/LINC00507, c; RORB, b) and deep (THEMIS, d; FEZF2, e) layers.
Extended Data Fig. 7 RNAscope mFISH validation of ten excitatory neuron types.
a, Heat map summarizing combinatorial three-gene panels used for multiplex FISH assays to explore the spatial distribution of ten excitatory clusters. Gene combinations for each cluster are indicated by coloured boxes on the heat map. Maximum expression values for each gene are listed on the right of the heat map and gene-expression values are displayed on a log scale. Experiments were repeated on at least two donors for each probe combination with similar results. b, Gene combinations probed are listed above each image. Labelled cells are indicated by white arrows. Scale bar, 20 µm. c, Schematic representing the laminar distribution of clusters on the basis of the observed positions of labelled cells across at least three sections from at least two donors per cell type.
Extended Data Fig. 8 In situ validation of LOC, long non-coding RNA and antisense transcripts as cell-type-specific markers.
a, LINC01164 specifically labels the Exc L3–5 RORB COL22A1 cluster (n = 160 nuclei). Left, violin plots showing expression of genes used for cluster validation by mFISH. Middle, read pile-ups shown for LINC01164 across all excitatory clusters (n = 24), viewed in UCSC genome browser. Red box indicates Exc L3–5 RORB COL22A1 cluster. Right, mFISH validation of cluster-specific marker genes. Laminar distribution of the Exc L3–5 RORB COL22A1 cluster marked by LINC01164 is consistent with the distribution shown using protein-coding marker genes (panel showing staining for RORB, MME and NTNG1 reproduced from Fig. 2c). Scale bars, 100 µm (low-magnification DAPI-stained columns); 5 µm (mFISH images). Experiments were repeated on two donors with similar results. b, The Exc L4–6 FEZF2 IL26 cluster (n = 344 nuclei) is specifically marked by INFG-AS1 and LOC105369818. Top, heat map showing expression of these genes along with protein-coding marker gene CARD11. Bottom, mFISH validation of cluster-specific marker genes. Experiments were repeated on three donors with similar results. Scale bars, 5 µm. Right, read pile-ups shown for INFG-AS1 across all excitatory clusters, viewed in UCSC genome browser. Red box indicates Exc L4–6 FEZF2 IL26 cluster. c, Violin plots showing expression of LOC105376081 in the Exc L3–5 RORB ESR1 cluster (n = 1,428 nuclei). Right, ISH for LOC105376081 shows expression in L4 (red bar), consistent with the anatomical location of Exc L3–5 RORB ESR1 (left, laminar distribution from Fig. 2). Scale bars, 100 µm. d, Violin plots showing expression of LOC401134 and the protein-coding gene CRYM in three L3–5 RORB-expressing clusters (n = 1,674 nuclei). Right, mFISH showing three possible combinations for the genes assayed as indicated by labelled arrows. Scale bars, 10 µm. Experiments were repeated on two donors with similar results. e, LOC102723415 labels a subset of PVALB clusters (n = 618 nuclei) as shown in the violin plots on the left and mFISH images on the right (clusters indicated by labelled arrows). Scale bars, 5 µm. Experiments were repeated on two donors with similar results. For all violin plots, rows are genes, black dots correspond to median expression and maximum expression (CPM) is listed on the far right. Expression values are on a linear scale. Asterisks indicate lipofuscin in mFISH images.
Extended Data Fig. 9 Laminar distribution of superficial excitatory neuron types validated by smFISH.
a, smFISH (image, 100×) was performed with probes against SLC17A7, CUX2, CBLN2, RFXP1, GAD2, COL5A2, LAMP5, PENK and CARTPT mRNA. Spots for each gene are pseudocoloured as indicated in the bottom right legend. Layer demarcations are indicated in magenta. Scale bar, 300 µm. b, Spot indications for each gene, pseudocoloured as indicated in the bottom right legend, as in a. a,a′, Superficial L2 cells express SLC17A7(lavender), CUX2 (magenta) and LAMP5 (mint). b,b′, At deeper locations in L2, an example of an SLC17A7-expressing cell with CUX2, LAMP5 and COL5A2 expression. Note that LAMP5 expression (mint) decreases in CUX2/SLC17A7-expressing cells, whereas COL5A2/CUX2-expressing cells increase with depth along L2 and L3 (see, c,c′; d,d′; e,e′). c, Probe density (spots per 100 µm2) for nine genes assayed across L1–L4 (and partially L5) of human MTG. The cortical slice was approximately 0.5 mm wide and 2 mm deep. Points correspond to cellular locations in situ where the y axis is the cortical depth from the pial surface and the x axis is the lateral position. Point size and colour correspond to probe density. Cells that lack probe expression are shown as small grey points. Experiments were repeated on three donors with similar results. d, In situ location of cells mapped to indicated cell types and classes (different panels) on the basis of expression levels of nine genes shown in a. Numbers indicate qualitative calls of the layer to which each cell belongs based on cytoarchitecture, and 0 indicates that the cell was not annotated.
Extended Data Fig. 10 Layer distributions and frequencies of inhibitory neuron types.
a, b, Constellation diagram showing cluster relationships, relative frequencies and average layer position for LAMP5/PAX6 (n = 2,320 nuclei) (a) and SST/PVALB (n = 1,844 nuclei) (b) classes of inhibitory neurons. c, Chromogenic ISH for TH, a marker of Inh L5–6 SST TH and NPY, a marker of Inh L3−6 SST NPY, from the Allen Human Brain Atlas. Left columns show greyscale images of the Nissl section nearest the ISH section shown on the right for each gene. Red dots show cells positive for the gene assayed by ISH. Experiments were repeated 9 (NPY) and 40 (TH) times with similar results. Chromogenic ISH for Th and Npy in mouse TEa from the Allen Mouse Brain Atlas are to the right of the human images. Experiments were repeated six (Npy) and two (Th) times with similar results. Scale bars, human, 250 µm; mouse, 100 µm. d, RNAscope mFISH for markers of Inh L2−5 PVALB SCUBE3. Left, inverted DAPI-stained cortical column with red dots marking cells positive for the genes GAD1, PVALB and NOG (scale bar, 250 µm). Middle, cells positive for GAD1, PVALB and the specific marker genes NOG (top: scale bar, 10 µm) and COL15A1 (bottom: scale bar, 10 µm). White arrows mark triple-positive cells. Experiments were repeated on three donors with similar results. Right, counts of GAD1+PVALB+NOG+ cells across layers (expressed as percentage of total triple-positive cells). Data are mean ± s.d. and dots show the data points for individual specimens (n = 3 subjects). Violin plots show gene-expression distributions across clusters in the PVALB subclass (n = 802 nuclei) for the chandelier cell marker UNC5B and the Inh L2−5 PVALB SCUBE3 cluster markers NOG and COL15A1. Rows are genes, black dots correspond to median expression and maximum expression (CPM) is listed on the far right. Expression values are on a linear scale. e, Inverted DAPI-stained cortical column illustrating laminar positions of cells labelled with interneuron class markers. Green dots mark GAD1+/Gad1+, ADARB2+/Adarb2+ and LHX6−/Lhx6− cells (that is, ADARB2 branch interneurons); blue dots mark GAD1+/Gad1+, ADARB2−/Adarb2− and LHX6+/Lhx6+ cells (that is, LHX6 branch interneurons); and pink dots mark GAD1+/Gad1+, ADARB2+/Adarb2+ and LHX6+/Lhx6+ cells (that is, Inh L2–6 LAMP5 CA1 cells in human or Lamp5 Lhx6 cells in mouse). f, Representative images of cells labelled with the GAD1, ADARB2 and LHX6 gene panel for human (top) and mouse (bottom). Left to right: cells double positive for GAD1 and ADARB2; cells double positive for GAD1 and LHX6; and GAD1, ADARB2 and LHX6 triple-positive cells. Scale bars, 15 µm (human), 10 µm (mouse). Experiments were repeated on three donors and three mice with similar results.
Extended Data Fig. 11 Aligning snRNA-seq and scRNA-seq data from human and mouse cortex.
a, Heat map of Pearson’s correlations between average MetaNeighbour AUROC scores (n = 384 gene sets) for three broad classes of human and mouse cortical cell types. Rows and columns are ordered by average-linkage hierarchical clustering. b, Human (blue; n = 3,594 nuclei) and mouse (orange; n = 6,595 cells) inhibitory neurons projected on the first two principal components of a PCA combining expression data from both species. Almost 20% of expression differences are explained by species, whereas 6% are explained by major classes of interneurons. c, Number of highly differentially expressed (>tenfold change) genes (out of 14,551 orthologous genes) between homologous cell types matched between species (n = 37 types), mouse cortical area22 (n = 103 types) and sample type26 (n = 11 types). Box plots show median, interquartile interval, range and outlier values. d, Schematic of scAlign analysis to align RNA-seq data from human nuclei and mouse cells. e, t-SNE plots of human (blue; n = 3,503 nuclei) and mouse (orange; n = 4,127 cells) excitatory neurons after alignment with scAlign and coloured by species and cluster. Arrow highlights two human nuclei that cluster with the mouse-specific (M) L5 PT Chrna6 cluster. f, t-SNE plots of human (blue; n = 670 nuclei) and mouse (orange; n = 671 cells) non-neuronal cells coloured by species and cluster. g, t-SNE plots of human (blue; n = 3,594 nuclei) and mouse (orange; n = 6,595 cells) inhibitory neurons after alignment with scAlign (as in Fig. 5c) and Seurat and coloured by species. h, Consistently higher accuracy and alignment of inhibitory neurons using scAlign versus Seurat with several neural network architectures and parameter settings. Box plots show median and interquartile interval of values.
Extended Data Fig. 12 Quantifying human and mouse cell-type homology and comparing cell-type frequencies between species.
a–c, Heat maps with inferred cell-type homologies highlighted in blue boxes. For each pair of clusters, the shade of grey indicates the minimum proportion of samples that co-cluster. Homologies for human and mouse inhibitory neurons (a), excitatory neurons (b) and non-neuronal cells (c) were predicted on the basis of shared cluster membership using mouse cells from two cortical areas (V1 and ALM) and two unsupervised alignment algorithms (scAlign and Seurat). d, Mouse V1 and mouse ALM excitatory neurons were aligned with scAlign. Blue boxes indicate V1 and ALM clusters that align to the same human clusters in b and are members of homologous cell types. Note that cell types can be matched at higher resolution within species than between species, as expected. e, Left to right: violin plots (n = 10,525 nuclei) showing expression of specific markers of the putative extratelencephalic EXC L4–5 FEZF2 SCN4B cluster (black box) and NPTX1, a gene expressed by all non-PT excitatory neurons. Each row represents a gene, the black dots in each violin represent median gene expression within clusters, and the maximum expression value for each gene is shown on the right-hand side of each row. Expression values are shown on a linear scale. Representative inverted DAPI-stained cortical column (scale bar, 200 µm) with red dots marking the position of cells positive for the genes SLC17A7 and FAM84B and negative for NPTX1 illustrates the relative abundance of the EXC L4–5 FEZF2 SCN4B type in human MTG. Representative examples (arrows) of FAM84B (scale bar, 25 µm) and POU3F1-expressing cells (scale bar, 25 µm). Expression of Fam84b in mouse TEa (scale bar, 75 µm) is shown in the adjacent panel. f, mFISH for NPTX1, a marker of non-PT excitatory types and SLC17A7, shows that NPTX1 labels most SLC17A7+ cells across all cortical layers. Boxed region shown at higher the magnification to the right. One SLC17A7+ cell (white arrow) is NPTX1−, but all other all other SLC17A7+ cells are NPTX1+. Scale bars, 200 µm (left); 50 µm (right). Right, representative inverted DAPI-stained cortical column with red dots that represent SLC17A7+, NPTX1− and POU3F1+ cells. Scale bar, 200 µm. e, f, Experiments were repeated on three donors (human) and two mice with similar results. g, ISH validation of layer distributions in human MTG and mouse primary visual cortex (data from ref. 22). Cells are labelled by cluster marker genes in human (RORB+/CNR1−/PRSS12+) and mouse (Scnn1a+/Hsd11b1+). ISH was performed on three human donors with similar results. For mouse, one experiment was performed.
Extended Data Fig. 13 Marker genes with relatively conserved expression in homologous cell types between human and mouse.
Expression heat maps of homologous cell-type markers in human cortical nuclei and mouse cortical cells. Rows, median expression based on intronic and exonic reads and log-transformed (log10(CPM + 1)). Values listed on the right side of each heat map indicate the maximum expression level (CPM) for each gene. Columns: single nuclei (human) or cells (mouse) grouped by homologous types identified in this study. For each homologous type, up to ten marker genes were identified based on relatively specific expression (median CPM >1 in six or fewer clusters and ordered by τ score) in both species. Note that many more genes support individual homologies but may not be cell-type-specific markers.
Supplementary information
Supplementary Information
A guide for Supplementary Tables 1-6.
Supplementary Table 1
Gene expression signature of donor tissue type - see Supplementary Information document for full description.
Supplementary Table 2
Cell type meta-data and marker genes - see Supplementary Information document for full description.
Supplementary Table 3
Provisional cell type ontology - see Supplementary Information document for full description.
Supplementary Table 4
Human and mouse homologous cortical cell types - see Supplementary Information document for full description.
Supplementary Table 5
Gene expression summary - see Supplementary Information document for full description.
Supplementary Table 6
Divergence of expression across gene families - see Supplementary Information document for full description.
Video 1
: Cytosplore viewer for exploration of gene expression data in human cortical neurons. Exploration of gene expression variation within and between layer 2 and 3 excitatory neuron types in human middle temporal gyrus.
Rights and permissions
About this article
Cite this article
Hodge, R.D., Bakken, T.E., Miller, J.A. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019). https://doi.org/10.1038/s41586-019-1506-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1506-7
This article is cited by
-
txci-ATAC-seq: a massive-scale single-cell technique to profile chromatin accessibility
Genome Biology (2024)
-
Morphological diversification and functional maturation of human astrocytes in glia-enriched cortical organoid transplanted in mouse brain
Nature Biotechnology (2024)
-
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Nature Communications (2024)
-
TISSUE: uncertainty-calibrated prediction of single-cell spatial transcriptomics improves downstream analyses
Nature Methods (2024)
-
Cellular development and evolution of the mammalian cerebellum
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.