Abstract
PIEZO1 is a mechanosensitive channel that converts applied force into electrical signals. Partial molecular structures show that PIEZO1 is a bowl-shaped trimer with extended arms. Here we use cryo-electron microscopy to show that PIEZO1 adopts different degrees of curvature in lipid vesicles of different sizes. We also use high-speed atomic force microscopy to analyse the deformability of PIEZO1 under force in membranes on a mica surface, and show that PIEZO1 can be flattened reversibly into the membrane plane. By approximating the absolute force applied, we estimate a range of values for the mechanical spring constant of PIEZO1. Both methods of microscopy demonstrate that PIEZO1 can deform its shape towards a planar structure. This deformation could explain how lateral membrane tension can be converted into a conformation-dependent change in free energy to gate the PIEZO1 channel in response to mechanical perturbations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Any data relating to the findings presented in this Article are available from the corresponding authors upon reasonable request.
References
Coste, B. et al. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330, 55–60 (2010).
Coste, B. et al. Piezo proteins are pore-forming subunits of mechanically activated channels. Nature 483, 176–181 (2012).
Wu, J., Lewis, A. H. & Grandl, J. Touch, tension, and transduction – the function and regulation of Piezo ion channels. Trends Biochem. Sci. 42, 57–71 (2017).
Li, J. et al. Piezo1 integration of vascular architecture with physiological force. Nature 515, 279–282 (2014).
Ranade, S. S. et al. Piezo1, a mechanically activated ion channel, is required for vascular development in mice. Proc. Natl Acad. Sci. USA 111, 10347–10352 (2014).
Retailleau, K. et al. Piezo1 in smooth muscle cells is involved in hypertension-dependent arterial remodeling. Cell Reports 13, 1161–1171 (2015).
Cahalan, S. M. et al. Piezo1 links mechanical forces to red blood cell volume. eLife 4, (2015).
Wang, S. et al. Endothelial cation channel PIEZO1 controls blood pressure by mediating flow-induced ATP release. J. Clin. Invest. 126, 4527–4536 (2016).
Rode, B. et al. Piezo1 channels sense whole body physical activity to reset cardiovascular homeostasis and enhance performance. Nat. Commun. 8, 350 (2017).
Ranade, S. S. et al. Piezo2 is the major transducer of mechanical forces for touch sensation in mice. Nature 516, 121–125 (2014).
Woo, S. H. et al. Piezo2 is the principal mechanotransduction channel for proprioception. Nat. Neurosci. 18, 1756–1762 (2015).
Demolombe, S., Duprat, F., Honoré, E. & Patel, A. Slower Piezo1 inactivation in dehydrated hereditary stomatocytosis (xerocytosis). Biophys. J. 105, 833–834 (2013).
Andolfo, I. et al. Novel Gardos channel mutations linked to dehydrated hereditary stomatocytosis (xerocytosis). Am. J. Hematol. 90, 921–926 (2015).
Fotiou, E. et al. Novel mutations in PIEZO1 cause an autosomal recessive generalized lymphatic dysplasia with non-immune hydrops fetalis. Nat. Commun. 6, 8085 (2015).
Coste, B. et al. Gain-of-function mutations in the mechanically activated ion channel PIEZO2 cause a subtype of distal arthrogryposis. Proc. Natl Acad. Sci. USA 110, 4667–4672 (2013).
Guo, Y. R. & MacKinnon, R. Structure-based membrane dome mechanism for Piezo mechanosensitivity. eLife 6, e33660 (2017).
Saotome, K. et al. Structure of the mechanically activated ion channel Piezo1. Nature 554, 481–486 (2018).
Zhao, Q. et al. Structure and mechanogating mechanism of the Piezo1 channel. Nature 554, 487–492 (2018).
Wu, J., Goyal, R. & Grandl, J. Localized force application reveals mechanically sensitive domains of Piezo1. Nat. Commun. 7, 12939 (2016).
Wang, Y. et al. A lever-like transduction pathway for long-distance chemical- and mechano-gating of the mechanosensitive Piezo1 channel. Nat. Commun. 9, 1300 (2018).
Lewis, A. H. & Grandl, J. Mechanical sensitivity of Piezo1 ion channels can be tuned by cellular membrane tension. eLife 4, e12088 (2015
Cox, C. D. et al. Removal of the mechanoprotective influence of the cytoskeleton reveals PIEZO1 is gated by bilayer tension. Nat. Commun. 7, 10366 (2016).
Árnadóttir, J. & Chalfie, M. Eukaryotic mechanosensitive channels. Annu. Rev. Biophys. 39, 111–137 (2010).
Brohawn, S. G., Su, Z. & MacKinnon, R. Mechanosensitivity is mediated directly by the lipid membrane in TRAAK and TREK1 K+ channels. Proc. Natl Acad. Sci. USA 111, 3614–3619 (2014).
Brohawn, S. G., Campbell, E. B. & MacKinnon, R. Physical mechanism for gating and mechanosensitivity of the human TRAAK K+ channel. Nature 516, 126–130 (2014).
Moe, P. & Blount, P. Assessment of potential stimuli for mechano-dependent gating of MscL: effects of pressure, tension, and lipid headgroups. Biochemistry 44, 12239–12244 (2005).
Zhang, W. et al. Ankyrin repeats convey force to gate the NOMPC mechanotransduction channel. Cell 162, 1391–1403 (2015).
Jin, P. et al. Electron cryo-microscopy structure of the mechanotransduction channel NOMPC. Nature 547, 118–122 (2017).
Gaub, B. M. & Müller, D. J. Mechanical stimulation of Piezo1 receptors depends on extracellular matrix proteins and directionality of force. Nano Lett. 17, 2064–2072 (2017).
Hamill, O. P. & McBride, D. W., Jr. Induced membrane hypo/hyper-mechanosensitivity: a limitation of patch-clamp recording. Annu. Rev. Physiol. 59, 621–631 (1997).
Suchyna, T. M., Markin, V. S. & Sachs, F. Biophysics and structure of the patch and the gigaseal. Biophys. J. 97, 738–747 (2009).
Moroni, M., Servin-Vences, M. R., Fleischer, R., Sánchez-Carranza, O. & Lewin, G. R. Voltage gating of mechanosensitive PIEZO channels. Nat. Commun. 9, 1096 (2018).
Lacroix, J. J., Botello-Smith, W. M. & Luo, Y. Probing the gating mechanism of the mechanosensitive channel Piezo1 with the small molecule Yoda1. Nat. Commun. 9, 2029 (2018).
Ando, T., Uchihashi, T. & Scheuring, S. Filming biomolecular processes by high-speed atomic force microscopy. Chem. Rev. 114, 3120–3188 (2014).
Miyagi, A. & Scheuring, S. Automated force controller for amplitude modulation atomic force microscopy. Rev. Sci. Instrum. 87, 053705 (2016).
Helfrich, W. Elastic properties of lipid bilayers: theory and possible experiments. Z. Naturforsch. C 28, 693–703 (1973).
Legleiter, J., Park, M., Cusick, B. & Kowalewski, T. Scanning probe acceleration microscopy (SPAM) in fluids: mapping mechanical properties of surfaces at the nanoscale. Proc. Natl Acad. Sci. USA 103, 4813–4818 (2006).
Kiracofe, D. et al. VEDA: Virtual Environment for Dynamic AFM, https://nanohub.org/resources/veda (2012).
Guzman, H. V., Garcia, P. D. & Garcia, R. Dynamic force microscopy simulator (dForce): A tool for planning and understanding tapping and bimodal AFM experiments. Beilstein J. Nanotechnol. 6, 369–379 (2015).
García, R. & San Paulo, A. Attractive and repulsive tip-sample interaction regimes in tapping-mode atomic force microscopy. Phys. Rev. B 60, 4961–4967 (1999).
Weisstein, E. W. Spherical Cap, from MathWorld–a Wolfram web resource. http://mathworld.wolfram.com/SphericalCap.html.
Haselwandter, C. A. & MacKinnon, R. Piezo’s membrane footprint and its contribution to mechanosensitivity. eLife 7, e41968 (2018).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Acknowledgements
We thank M. Ebrahim and J. Sotiris at the Evelyn Gruss Lipper Cryo-EM Resource Center of Rockefeller University for assistance with cryo-EM data collection. Y.R.G. is a Howard Hughes Medical Institute Fellow of the Damon Runyon Cancer Research Foundation (DRG 2317-18). R.M. is an investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Y.-C.L., Y.R.G., R.M. and S.S. designed the study; Y.R.G. and J.L. purified and reconstituted the protein. Y.R.G. performed cryo-EM measurements. Y.-C.L. and A.M. performed HS-AFM experiments. Y.-C.L., Y.R.G., R.M. and S.S. analysed the data. Y.-C.L., Y.R.G., R.M. and S.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Peer review information Nature thanks Ardem Patapoutian, Victor Shahin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Fig. 1 Architecture and topology of the mechanosensitive channel PIEZO1.
a, Top, bottom and side views of PIEZO1 (PDB 6B3R) in cartoon representation (top) and embedded in the micelle density map (EMD-7042), contoured at 6σ. CTD, C-terminal domain. b, Top, topology of PIEZO1 shown rainbow-coloured (with N terminus in blue and C terminus in red), except for the structurally unsolved TM1–TM12 regions (shown in grey). Helices are represented as cylinders, loops as solid lines and unresolved regions as dotted lines. Bottom, top view of transmembrane helices, labelled as in the topology. Red squares outline four-transmembrane units that constitute the arm. TM21–TM24 are at the ‘elbow’ of the arm. The hypothetical position of the unresolved units TM1–TM4, TM5–TM8 and TM9–TM12 are indicated (dashed outline).
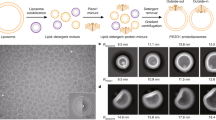
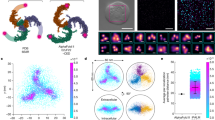
Extended Data Fig. 2 Image processing procedure to determine the Rc of side-view PIEZO1 channels, and the intrinsic Rc of the vesicles in which they are embedded.
a, The input 2D-averaged image of one side of the vesicle (here the PIEZO1 occupied side). b, Edge detection output using the Canny method. c, Edge detection output overlaid with the selection polygon. d, Edges in the selected polygon region annotated with numbers in different colours for easier identification. e, f, Measured (blue circle) and fitted (red dotted line) circles of the edges corresponding to the outer (e) and inner (f) boundaries of the vesicle membrane. Centre coordinates, the radius and its 95% confidence interval are shown. The unit of all values is in pixels. g, Fitted circles (red dashed lines) overlaid onto the edge detection output. Radii with the confidence interval of outer and inner boundaries are shown in units of nm. h, Fitted circles (red dashed lines) overlaid onto the input 2D-averaged image.
Extended Data Fig. 3 Estimation of Fts in HS-AFM at various Aratio, on the basis of experimental tip motion analysis and numerical simulation.
a, b, Average tip motions observed at different Aratio in buffer solution during HS-AFM imaging on mica (a) and a supported lipid bilayer (1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC):DOPS, 4:1) (b). c, d, Forces caused by the elastic response of the cantilever (top), the hydrodynamic damping with the medium (middle) and the total force that governs the tip motion (bottom). e, f, Sum of the drive force and tip-sample interaction force. g, h, Reconstructed Fts trajectories during a single oscillation cycle, based on the point-mass model (equation (7), equation (8), Methods). i, Comparison between peak forces obtained from reconstructed Fts trajectories on mica (blue dashed line) and membrane (red dashed line), and peak forces simulated using different surface stiffness using VEDA38 (dashed lines and grey shadowed area). The VEDA simulation is performed by using the amplitude-modulation-approach curve tool with the following settings: discrete approach steps within a defined z-range, acoustic excitation, Afree = 2 nm, Hertz contact model (Etip = 130 GPa, νtip = 0.3, νsample = 0.5, ν is the Poisson’s ratio) with a tip radius of 1 nm, and other HS-AFM experimental parameters (for example, k, Q and ω0). j, Comparison between average forces obtained from reconstructed Fts trajectories on mica (blue dashed line) and membrane (red dashed line) and values calculated through equation (1) (thick black dashed line). k, Comparison between peak forces obtained from experimental Fts trajectories on membrane at different Afree values. Using second-order polynomial fitting, the peak force reconstructed in the condition of Afree = 1.5 nm can be well-described by y = −688.7x2 + 633.4x + 55 (black dashed line) with R2 = 0.99. This fitting allows us to estimate the upper bound of force (peak force) applied to PIEZO1 channels at any given Aratio. Tip trajectories are representative of ≥5 independent experiments using ≥3 different HS-AFM cantilevers.
Extended Data Fig. 4 Classification of PIEZO1 channels using cross-correlation analysis on characteristic dimensions measured upon force application.
a, b, Cross-correlation density maps of smallest halo radius (Rmin) at low applied force versus largest halo radius (Rmax) at highest applied force (a) and of smallest halo radius (Rmin) versus maximum central height (Hmax) at lowest applied force (b). Along the diagonal direction of the cross-correlation density maps, the molecules separate into two peaks. This suggests the existence of two subtypes of PIEZO1 with different sizes in HS-AFM force-sweep movies. Therefore, we assigned the molecules in the peaks as type-1 (about 70%, white dashed circle) and type-2 (about 30%, yellow dashed circle) PIEZO1, respectively. The same molecules populate in the same subtype according to all analysed criteria. The total number of analysed PIEZO1 particles is 143, from 11 HS-AFM movies acquired on ≥5 different samples, days and HS-AFM tips.
Extended Data Fig. 5 Mechanical response of type-2 PIEZO1 to applied force.
a, HS-AFM image at \(\left\langle {F}_{{\rm{HS-AFM}}}\right\rangle \) of about 52 pN, of type-2 PIEZO1 viewed from the extracellular face. b, Height section profile (top) and radial height profile (bottom) of the topography displayed in a. c, Dimensional analysis of single type-2 PIEZO1 particle in a force-sweep HS-AFM experiment. Similar to the type-1 PIEZO1 (as reported in the main text), each single molecule (left) is 360-fold-symmetry averaged (centre) and a kymograph (right) across the centre profile (dashed line in 360-fold image) is calculated. The kymograph highlights the halo expansion (dashed line) as a function of force. Bottom, force as function of frame acquisition or time during the force-sweep HS-AFM movie acquisition. The yellow coloured area corresponds to the image acquisition of the particle shown (left). d, Normalized probability density maps of halo radius (R) as a function of force. Type-2 PIEZO1 also shows structural reversibility. The total number of analysed type-2 PIEZO1 particles is 43, from 11 HS-AFM movies acquired on ≥5 different samples, days and HS-AFM tips.
Extended Data Fig. 6 PIEZO1 has an exceptional low 2D density of transmembrane helices.
Transmembrane-helix 2D-density analysis for channels and transporters of known 3D structure. The transmembrane-helix density is estimated by the total number of transmembrane helices divided by the occupied membrane area of each known structure. A total of 201 channels and transporters structures have been analysed. A general trend seems to be that the channels are somewhat less densely packed than transporters, which is possibly a signature of the existence of a protein-free pore region. The PIEZO1 structure is an outlier: it is of exceptional size and has an exceptionally low density of transmembrane helices, (less than 0.2 of a helix per square nanometre. This unique feature might be a signature, and prerequisite for PIEZO1 mechanosensing.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion.
Video 1
HS-AFM force-sweep video of the extracellular face of Piezo1 channels undergoing reversible conformational changes under force application. The top right molecule is the molecule shown in the middle panel in Fig. 4c. The bottom graph displays the calculated <FHS-AFM> during video acquisition. The HS-AFM video was recorded at 187 nm x 187 nm image size (280 x 280 pixels) and 1.0 s per frame.
Video 2
HS-AFM force-sweep video of Piezo1 channels exposed to a reverse force cycle. The bottom graph displays the calculated <FHS-AFM> (red), Aset/Afree ratio (blue) and the Z-piezo displacement (green) as a function of frame acquisition or time during the force-sweep HS-AFM video acquisition shown in Fig. 5a. The HS-AFM video was recorded at 350 nm x 350 nm image size (350 x 350 pixels) and 1.0 s per frame.
Video 3
Example of lateral expansion analysis of a single Piezo1 particle shown in Fig. 5b. The single molecule (left) is 360-fold symmetry averaged (middle) and a kymograph (right) across the center of the Piezo1 calculated (dashed line in 360-fold image). The kymograph highlights the outer radius (halo) expansion as a function of force application.
Rights and permissions
About this article
Cite this article
Lin, YC., Guo, Y.R., Miyagi, A. et al. Force-induced conformational changes in PIEZO1. Nature 573, 230–234 (2019). https://doi.org/10.1038/s41586-019-1499-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1499-2
This article is cited by
-
Matrix stiffness affects tumor-associated macrophage functional polarization and its potential in tumor therapy
Journal of Translational Medicine (2024)
-
Scaling up Functional Analyses of the G Protein-Coupled Receptor Rhodopsin
Journal of Molecular Evolution (2024)
-
Piezo1, the new actor in cell volume regulation
Pflügers Archiv - European Journal of Physiology (2024)
-
Aberrant mechanical loading induces annulus fibrosus cells apoptosis in intervertebral disc degeneration via mechanosensitive ion channel Piezo1
Arthritis Research & Therapy (2023)
-
A simple method to make, trap and deform a vesicle in a gel
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.