Abstract
Cation–chloride cotransporters (CCCs) mediate the electroneutral transport of chloride, potassium and/or sodium across the membrane. They have critical roles in regulating cell volume, controlling ion absorption and secretion across epithelia, and maintaining intracellular chloride homeostasis. These transporters are primary targets for some of the most commonly prescribed drugs. Here we determined the cryo-electron microscopy structure of the Na–K–Cl cotransporter NKCC1, an extensively studied member of the CCC family, from Danio rerio. The structure defines the architecture of this protein family and reveals how cytosolic and transmembrane domains are strategically positioned for communication. Structural analyses, functional characterizations and computational studies reveal the ion-translocation pathway, ion-binding sites and key residues for transport activity. These results provide insights into ion selectivity, coupling and translocation, and establish a framework for understanding the physiological functions of CCCs and interpreting disease-related mutations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The three-dimensional cryo-EM density maps have been deposited into the Electron Microscopy Data Bank under accession numbers EMD-0470, EMD-0471, EMD-0472, EMD-0473, EMD-0474 and EMD-0475. The coordinates are deposited into the Protein Data Bank with accession numbers 6NPH, 6NPJ, 6NPK and 6NPL. All simulation starting models and trajectories are available upon request.
References
Gamba, G. Molecular physiology and pathophysiology of electroneutral cation–chloride cotransporters. Physiol. Rev. 85, 423–493 (2005).
Haas, M. & Forbush, B., III. The Na–K–Cl cotransporter of secretory epithelia. Annu. Rev. Physiol. 62, 515–534 (2000).
Arroyo, J. P., Kahle, K. T. & Gamba, G. The SLC12 family of electroneutral cation-coupled chloride cotransporters. Mol. Aspects Med. 34, 288–298 (2013).
Russell, J. M. Sodium–potassium–chloride cotransport. Physiol. Rev. 80, 211–276 (2000).
Kaila, K., Price, T. J., Payne, J. A., Puskarjov, M. & Voipio, J. Cation–chloride cotransporters in neuronal development, plasticity and disease. Nat. Rev. Neurosci. 15, 637–654 (2014).
Gagnon, K. B. & Delpire, E. Physiology of SLC12 transporters: lessons from inherited human genetic mutations and genetically engineered mouse knockouts. Am. J. Physiol. Cell Physiol. 304, C693–C714 (2013).
Duarte, J. D. & Cooper-DeHoff, R. M. Mechanisms for blood pressure lowering and metabolic effects of thiazide and thiazide-like diuretics. Expert Rev. Cardiovasc. Ther. 8, 793–802 (2010).
ALLHAT Officers and Coordinators for the ALLHAT Collaborative Research Group. Major outcomes in high-risk hypertensive patients randomized to angiotensin-converting enzyme inhibitor or calcium channel blocker vs diuretic: The Antihypertensive and Lipid-Lowering Treatment to Prevent Heart Attack Trial (ALLHAT). J. Am. Med. Assoc. 288, 2981–2997 (2002).
Schrier, R. W. Use of diuretics in heart failure and cirrhosis. Semin. Nephrol. 31, 503–512 (2011).
Markadieu, N. & Delpire, E. Physiology and pathophysiology of SLC12A1/2 transporters. Pflug. Arch. 466, 91–105 (2014).
Flemmer, A. W., Gimenez, I., Dowd, B. F., Darman, R. B. & Forbush, B. Activation of the Na–K–Cl cotransporter NKCC1 detected with a phospho-specific antibody. J. Biol. Chem. 277, 37551–37558 (2002).
Hartmann, A. M. & Nothwang, H. G. Molecular and evolutionary insights into the structural organization of cation chloride cotransporters. Front. Cell. Neurosci. 8, 470 (2015).
Payne, J. A. Molecular operation of the cation chloride cotransporters: ion binding and inhibitor interaction. Curr. Top. Membr. 70, 215–237 (2012).
Gamba, G. et al. Molecular cloning, primary structure, and characterization of two members of the mammalian electroneutral sodium–(potassium)–chloride cotransporter family expressed in kidney. J. Biol. Chem. 269, 17713–17722 (1994).
Yamada, J. et al. Cl− uptake promoting depolarizing GABA actions in immature rat neocortical neurones is mediated by NKCC1. J. Physiol. 557, 829–841 (2004).
Abbas, L. & Whitfield, T. T. Nkcc1 (Slc12a2) is required for the regulation of endolymph volume in the otic vesicle and swim bladder volume in the zebrafish larva. Development 136, 2837–2848 (2009).
Flagella, M. et al. Mice lacking the basolateral Na–K–2Cl cotransporter have impaired epithelial chloride secretion and are profoundly deaf. J. Biol. Chem. 274, 26946–26955 (1999).
Somasekharan, S., Tanis, J. & Forbush, B. Loop diuretic and ion-binding residues revealed by scanning mutagenesis of transmembrane helix 3 (TM3) of Na–K–Cl cotransporter (NKCC1). J. Biol. Chem. 287, 17308–17317 (2012).
Moore-Hoon, M. L. & Turner, R. J. The structural unit of the secretory Na+–K+–2Cl− cotransporter (NKCC1) is a homodimer. Biochemistry 39, 3718–3724 (2000).
Pedersen, M., Carmosino, M. & Forbush, B. Intramolecular and intermolecular fluorescence resonance energy transfer in fluorescent protein-tagged Na–K–Cl cotransporter (NKCC1): sensitivity to regulatory conformational change and cell volume. J. Biol. Chem. 283, 2663–2674 (2008).
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl−-dependent neurotransmitter transporters. Nature 437, 215–223 (2005).
Ye, Z. Y., Li, D. P., Byun, H. S., Li, L. & Pan, H. L. NKCC1 upregulation disrupts chloride homeostasis in the hypothalamus and increases neuronal activity-sympathetic drive in hypertension. J. Neurosci. 32, 8560–8568 (2012).
Gupta, K. et al. The role of interfacial lipids in stabilizing membrane protein oligomers. Nature 541, 421–424 (2017).
Isenring, P. & Forbush, B. III. Ion and bumetanide binding by the Na–K–Cl cotransporter. Importance of transmembrane domains. J. Biol. Chem. 272, 24556–24562 (1997).
Gagnon, K. B., England, R. & Delpire, E. A single binding motif is required for SPAK activation of the Na–K–2Cl cotransporter. Cell. Physiol. Biochem. 20, 131–142 (2007).
Parvin, M. N., Gerelsaikhan, T. & Turner, R. J. Regions in the cytosolic C-terminus of the secretory Na+–K+–2Cl− cotransporter NKCC1 are required for its homodimerization. Biochemistry 46, 9630–9637 (2007).
Nezu, A., Parvin, M. N. & Turner, R. J. A conserved hydrophobic tetrad near the C terminus of the secretory Na+–K+–2Cl− cotransporter (NKCC1) is required for its correct intracellular processing. J. Biol. Chem. 284, 6869–6876 (2009).
Monette, M. Y. & Forbush, B. Regulatory activation is accompanied by movement in the C terminus of the Na–K–Cl cotransporter (NKCC1). J. Biol. Chem. 287, 2210–2220 (2012).
Rinehart, J. et al. Sites of regulated phosphorylation that control K–Cl cotransporter activity. Cell 138, 525–536 (2009).
Warmuth, S., Zimmermann, I. & Dutzler, R. X-ray structure of the C-terminal domain of a prokaryotic cation-chloride cotransporter. Structure 17, 538–546 (2009).
Parvin, M. N. & Turner, R. J. Identification of key residues involved in the dimerization of the secretory Na+–K+–2Cl− cotransporter NKCC1. Biochemistry 50, 9857–9864 (2011).
Harding, M. M. Metal-ligand geometry relevant to proteins and in proteins: sodium and potassium. Acta Crystallogr. D 58, 872–874 (2002).
Krishnamurthy, H., Piscitelli, C. L. & Gouaux, E. Unlocking the molecular secrets of sodium-coupled transporters. Nature 459, 347–355 (2009).
Faham, S. et al. The crystal structure of a sodium galactose transporter reveals mechanistic insights into Na+/sugar symport. Science 321, 810–814 (2008).
Weyand, S. et al. Structure and molecular mechanism of a nucleobase-cation-symport-1 family transporter. Science 322, 709–713 (2008).
Wahlgren, W. Y. et al. Substrate-bound outward-open structure of a Na+-coupled sialic acid symporter reveals a new Na+ site. Nat. Commun. 9, 1753 (2018).
Perez, C., Koshy, C., Yildiz, O. & Ziegler, C. Alternating-access mechanism in conformationally asymmetric trimers of the betaine transporter BetP. Nature 490, 126–130 (2012).
Dutzler, R., Campbell, E. B. & MacKinnon, R. Gating the selectivity filter in ClC chloride channels. Science 300, 108–112 (2003).
Knoers, N. V. Gitelman syndrome. Adv. Chronic Kidney Dis. 13, 148–154 (2006).
Wang, L., Dong, C., Xi, Y. G. & Su, X. Thiazide-sensitive Na+–Cl− cotransporter: genetic polymorphisms and human diseases. Acta Biochim. Biophys. Sin. 47, 325–334 (2015).
Alguel, Y., Cameron, A. D., Diallinas, G. & Byrne, B. Transporter oligomerization: form and function. Biochem. Soc. Trans. 44, 1737–1744 (2016).
Kowarz, E., Loscher, D. & Marschalek, R. Optimized Sleeping Beauty transposons rapidly generate stable transgenic cell lines. Biotechnol. J. 10, 647–653 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Ru, H. et al. Molecular mechanism of V(D)J recombination from synaptic RAG1–RAG2 complex structures. Cell 163, 1138–1152 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Swint-Kruse, L. & Brown, C. S. Resmap: automated representation of macromolecular interfaces as two-dimensional networks. Bioinformatics 21, 3327–3328 (2005).
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013).
Källberg, M. et al. Template-based protein structure modeling using the RaptorX web server. Nat. Protoc. 7, 1511–1522 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Betz, R.M. Dabble v.2.6.3. https://doi.org/10.5281/zenodo.836914 (2017).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Salomon-Ferrer, R., Götz, A. W., Poole, D., Le Grand, S. & Walker, R. C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald. J. Chem. Theory Comput. 9, 3878–3888 (2013).
Case, D. A. et al. AMBER (University of California, San Francisco, 2017).
Hopkins, C. W., Le Grand, S., Walker, R. C. & Roitberg, A. E. Long-time-step molecular dynamics through hydrogen mass repartitioning. J. Chem. Theory Comput. 11, 1864–1874 (2015).
Ryckaert, J., Ciccotti, G. & Berendsen, H. J. Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph.14, 33–38, 27–28 (1996).
Roe, D. R. & Cheatham, T. E. III. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Acknowledgements
We thank D. Bushnell and L. Montabana at Stanford-SLAC Cryo-EM facilities, Z. Li at HMS, C. Xu and K. Song at the UMass cryo-EM facility, and C. López and C. Yoshioka at the OHSU Multiscale Microscopy Core for help with electron microscopy data collection. We thank the staff at the Beckman Center Cell Sciences and Imaging Facility for their help with fluorescence imaging. This work was made possible by support from Stanford University and the Harold and Leila Y. Mathers Charitable Foundation to L.F., an NIH Kirschstein-NRSA fellowship and Medical Scientist Training Program funding to T.A.C., a Dean’s fellowship to J.Z., National Science Foundation Graduate Research Fellowships to N.R.L. and A.W., and a Stanford Graduate Fellowship to A.W. We thank A. Brunger and M. Maduke for helpful comments.
Author information
Authors and Affiliations
Contributions
T.A.C. performed molecular biology, biochemistry, cryo-EM sample preparation and functional studies and assisted in preparation of the figures and manuscript. B.J.O. collected and processed negative-stain and cryo-EM data, analysed the results of multi-body refinement, and assisted in preparation of the figures and manuscript. J.Z. performed biochemistry, negative-stain, cryo-EM grid preparation and optimization, cryo-EM data collection and model building and assisted in preparation of the figures and manuscript. N.R.L., A.W. and S.A.H. performed molecular dynamics simulations, analysed data and contributed to preparation of the figures and manuscript. D.-H.C. helped with the initial electron microscopy studies. R.O.D. oversaw the molecular dynamics simulations and contributed to the manuscript preparation. M.L. oversaw the cryo-EM, assisted cryo-EM data processing and contributed to the manuscript preparation. L.F. directed the project, oversaw biochemistry, functional studies, cryo-EM sample preparation and data collection and wrote the manuscript with input and support from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biochemical characterization of NKCC1 and cryo-EM data processing workflow for N-terminal truncated NKCC1.
a, NKCC1 in size-exclusion chromatography. Experiments were repeated six times independently with similar results. b, Representative negative-stain EM micrograph of purified NKCC1 in digitonin. Experiments were repeated six times independently with similar results. c, Two-dimensional class averages from negative-stained particles reveal the dimeric structure of NKCC1, with two soluble domains visible beneath the TMD and detergent micelle. Experiments were repeated six times independently with similar results. d, Representative cryo-EM micrograph of N-terminal truncated NKCC1 particles purified in digitonin. e, Two-dimensional class averages of N-terminal truncated NKCC1. f, Classification and refinement workflow used to obtain final cryo-EM maps of N-terminal truncated NKCC1. Two processing strategies were followed to obtain a high-resolution map of the TMD (left), or a composite map of the TMD and soluble domain following Relion3 multibody refinement (right).
Extended Data Fig. 2 Cryo-EM data processing workflow for full-length NKCC1.
a, Representative cryo-EM micrograph of full-length NKCC1 particles purified in digitonin. b, Two-dimensional class averages of full-length NKCC1. Several averages show high resolution features in the TMD; however, the soluble domain is fuzzy and disordered, indicative of conformational flexibility between these two domains. c, Classification and refinement workflow used to obtain final cryo-EM maps of full-length NKCC1. Two processing strategies were followed to obtain a high-resolution map of the TMD (left), or a composite map of the TMD and soluble domain following Relion3 multibody refinement (right).
Extended Data Fig. 3 Cryo-EM 3D reconstruction analysis, statistics and representative density.
a, Angle distributions for particles contributing to the high-resolution map of the TMD for full-length NKCC1. b, FSC curves: gold-standard FSC curve between the two half maps with indicated resolution at FSC = 0.143 (red); FSC curve between the atomic model of the TMD and the final map with indicated resolution at FSC = 0.5 (blue); FSC curve between half map 1 (orange) or half map 2 (green) and the atomic model refined against half map 1. c, Local resolution of the high-resolution TMD map as determined with ResMap. d, Slice view of the angle distributions for particles contributing to the map of the soluble domain for full-length NKCC1 obtained from Relion3 multibody refinement. e, FSC curves: gold-standard FSC curve between the two half maps with indicated resolution at FSC = 0.143 (red); FSC curve between the atomic model of the soluble domain and the final map with indicated resolution at FSC = 0.5 (blue); FSC curve between half map 1 (orange) or half map 2 (green) and the atomic model refined against half map 1. f, Local resolution of the soluble domain map as determined with ResMap. g, The cryo-EM density maps are of high quality throughout the TMD and show clear side chain features in CTD. Densities for all transmembrane helices, as well as helices and strands in the CTD are shown as wire mesh (6σ). The transporter is shown as sticks.
Extended Data Fig. 4 Principal component analysis of domain movements from multibody refinement.
a, Contribution of individual eigenvectors to the total variance in rotation and translation between the TMD and soluble domain. Eigenvectors 1–3 contribute more than 50% of the total variance in the rotations and translations between domains. b–d, Histograms of amplitude along eigenvector 1 (b), eigenvector 2 (c) and eigenvector 3 (c). All eigenvector amplitude histograms are monomodal, suggesting that the rotations and/or translations are continuous in nature. e–g, Representation of the extremes of rotation and translation between TMD and soluble domain along eigenvector 1 (e), eigenvector 2 (f) and eigenvector 3 (g). For simplicity of visualization, the maps at either extreme of an individual eigenvector were aligned on the TMDs. Blue arrows indicate the direction of movement of the soluble domain relative to the TMD.
Extended Data Fig. 5 Structure of soluble domain.
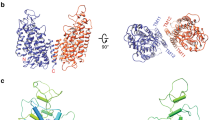
a, A single subunit of the NKCC soluble domain in ribbon representation. The N- and C-terminal halves of the CTD that show a similar structure are coloured in green and orange, respectively. b, The dimer of the soluble domain. One subunit is shown as a surface and the other is shown as a ribbon. c, Close-up view of the TMD–soluble domain interface. Two subunits are coloured in dark green and gold, respectively. The TMD is shown as a surface representation and the soluble domain as a ribbon representation. The NTD and the C-terminal end of the protein are located near the interface. d, The intracellular surface of the TMD. The surface is coloured according to electrostatic potential (red, −10 kT e−1; blue, +10 kT e−1).
Extended Data Fig. 6 Uptake activities of interface mutants and characterizations of all NKCC1 mutants in this study.
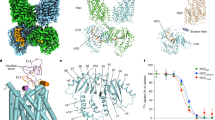
a, Uptake activities of NKCC1 mutants at the TMD–cytosolic domain interface. 86Rb+ uptake of NKCC1 mutants was normalized to that of the wild type (mean ± s.e.m., n = 4 independent experiments except for wild type, n = 7 independent experiments and for wild type with bumetanide, n = 3 independent experiments). b, Wild-type NKCC1 and mutants (also including those in Figs. 1e, 3e, 4f) in size-exclusion chromatography. The GFP fusion protein is monitored by fluorescence. Experiments were repeated three times independently with similar results. c, The expression level of wild-type NKCC1 and mutants as shown by western blot. Experiments were repeated three times independently with similar results. For gel source data, see Supplementary Fig. 2. d, The membrane localization of wild-type NKCC1 and mutants. The fluorescence images are shown for HEK293 cells expressing NKCC1–GFP. Experiments were repeated three times independently with similar results.
Extended Data Fig. 7 Molecular dynamics simulations.
a, Ion probability densities of ions within translocation pathway (top) and individual traces (bottom) for simulations performed with K+, Na+, and Cl− initially present in the translocation pathway (model 1, see Methods). b, Ion probability densities of ions within translocation pathway (top) and individual traces (bottom) for simulations performed with only K+ and Na+ initially present in the translocation pathway (model 1, see Methods). In the latter set of simulations, Cl– spontaneously explores regions that form stable chloride-binding sites. Probability density maps were calculated as described in Methods. The top four simulation traces represent the minimum distance of an ion of a particular type from the pre-determined centre of each binding site (see Methods). The bottom simulation trace shows the number of Cl− ions bound within any of the three chloride-binding sites at once.
Extended Data Fig. 8 Ion stability in molecular dynamics simulations.
a–d, Comparison of relative stabilities of ions across multiple simulation conditions. Here, stability refers to the amount of time an ion resides within 3 Å of the pre-determined centre of each binding site (see Methods). Each dot corresponds to a result from a single simulation. Closed circles indicate simulations with each cation placed in its predicted site; open circles indicate simulations with either cation swapped into the other cation’s predicted binding site. Blue circles correspond to measurements for Na+ and pink circles for K+. For a and b, we examined the relative stabilities of cations to determine whether the predicted Na+ and K+ binding sites exhibit a preference for their respective cations. a, For simulations started with all ions bound, Na+ resided longer in the Na+ site compared to K+ when K+ was initially placed within the Na+ site (‘Na+/K+/2Cl−’ vs ‘Na+/K+ swapped, 2 Cl−’). The same trend appeared for simulations with just cations initially bound within the cavity (‘Na+/K+’ vs ‘Na+/K+ swapped’). b, For simulations started with all ions bound, K+ resided for longer in the K+ site compared to Na+ when Na+ was initially placed within the K+ site (‘Na+/K+/2Cl−’ vs ‘Na+/K+ swapped, 2 Cl−’). For simulations with just cations initially bound, K+ and Na+ appeared to leave after similar amounts of time—perhaps because, particularly in the absence of Cl−, the K+ site is quite accessible to the intracellular solvent, and ions of either type could dissociate rapidly from this area. c, K+ remained within its initial position much longer compared to Na+ in the presence of two Cl− ions. In simulations with Na+ and 2 Cl− ions bound, Na+ often immediately dissociated from its site, perhaps indicating that the Na+ site on its own has a weaker affinity for binding cations in this state compared to the K+ site, the increased proximity of which to Cl− site 1 probably helps to stabilize K+ in the site. d, Chloride ions spontaneously visited and remained within 3 Å of the primary intracellular chloride site (site 2) for longer in the presence of both cations compared to in the presence of either cation alone. e, In one simulation, in which Na+ and K+ were initially placed in a swapped configuration with no Cl− ions bound, we observed escape of K+ (pink) followed by destabilization of the Na+ ion (blue) within the K+ site (123 ns) accompanied by Cl− binding. Na+ then proceeded to move down towards the intracellular side (220 ns) before rebinding for a 25-ns period within the proposed Na+-binding site.
Supplementary information
Supplementary Information
This file contains: Supplementary Figure 1 - sequence alignment of CCC transporters; Supplementary Figure 2 - source image for gel electrophoresis in Extended Data Figure 6c; Supplementary Table 1 -structural mapping of NCC mutations in Gitelman Syndrome and Supplementary References.
Video 1
Visualization of movement between NKCC1 TM and CTD along eigenvector 1, revealing a swiveling type motion between individual domains.
Video 2
Visualization of movement between NKCC1 TM and CTD along eigenvector 2, revealing a rocking type motion between individual domains.
Video 3
Visualization of movement between NKCC1 TM and CTD along eigenvector 3, revealing a rocking type motion between individual domains.
Rights and permissions
About this article
Cite this article
Chew, T.A., Orlando, B.J., Zhang, J. et al. Structure and mechanism of the cation–chloride cotransporter NKCC1. Nature 572, 488–492 (2019). https://doi.org/10.1038/s41586-019-1438-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1438-2
This article is cited by
-
Structure and thiazide inhibition mechanism of the human Na–Cl cotransporter
Nature (2023)
-
Structural basis for inhibition of the Cation-chloride cotransporter NKCC1 by the diuretic drug bumetanide
Nature Communications (2022)
-
Structural insights into the mechanism of the sodium/iodide symporter
Nature (2022)
-
Structure and function of an Arabidopsis thaliana sulfate transporter
Nature Communications (2021)
-
Cryo-EM structures of the TTYH family reveal a novel architecture for lipid interactions
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.