Abstract
Balanced fusion and fission are key for the proper function and physiology of mitochondria1,2. Remodelling of the mitochondrial inner membrane is mediated by the dynamin-like protein mitochondrial genome maintenance 1 (Mgm1) in fungi or the related protein optic atrophy 1 (OPA1) in animals3,4,5. Mgm1 is required for the preservation of mitochondrial DNA in yeast6, whereas mutations in the OPA1 gene in humans are a common cause of autosomal dominant optic atrophy—a genetic disorder that affects the optic nerve7,8. Mgm1 and OPA1 are present in mitochondria as a membrane-integral long form and a short form that is soluble in the intermembrane space. Yeast strains that express temperature-sensitive mutants of Mgm19,10 or mammalian cells that lack OPA1 display fragmented mitochondria11,12, which suggests that Mgm1 and OPA1 have an important role in inner-membrane fusion. Consistently, only the mitochondrial outer membrane—not the inner membrane—fuses in the absence of functional Mgm113. Mgm1 and OPA1 have also been shown to maintain proper cristae architecture10,14; for example, OPA1 prevents the release of pro-apoptotic factors by tightening crista junctions15. Finally, the short form of OPA1 localizes to mitochondrial constriction sites, where it presumably promotes mitochondrial fission16. How Mgm1 and OPA1 perform their diverse functions in membrane fusion, scission and cristae organization is at present unknown. Here we present crystal and electron cryo-tomography structures of Mgm1 from Chaetomium thermophilum. Mgm1 consists of a GTPase (G) domain, a bundle signalling element domain, a stalk, and a paddle domain that contains a membrane-binding site. Biochemical and cell-based experiments demonstrate that the Mgm1 stalk mediates the assembly of bent tetramers into helical filaments. Electron cryo-tomography studies of Mgm1-decorated lipid tubes and fluorescence microscopy experiments on reconstituted membrane tubes indicate how the tetramers assemble on positively or negatively curved membranes. Our findings convey how Mgm1 and OPA1 filaments dynamically remodel the mitochondrial inner membrane.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The atomic coordinates of Mgm1 have been deposited in the Protein Data Bank with accession number 6QL4. Maps obtained by subtomogram averaging were deposited in the Electron Microscopy Data Bank with accession numbers EMD-10062 (with PDB accession number 6RZT for the molecular model) and EMD-4584 for nucleotide-free Mgm1 on the outside of lipid tubes in a close-up view, and the overall tube structure, respectively. EMD-10063 (with PDB 6RZU) shows Mgm1 on the outside of a lipid tube in the GTPγS bound state. EMD-10064 (with PDB 6RZV) and EMD-10065 (with PDB 6RZW) show Mgm1 decorating the inside of a tube without and with GTPγS, respectively. All source data associated with the paper (beyond those deposited) are provided as Supplementary Information.
References
Nunnari, J. & Suomalainen, A. Mitochondria: in sickness and in health. Cell 148, 1145–1159 (2012).
Youle, R. J. & van der Bliek, A. M. Mitochondrial fission, fusion, and stress. Science 337, 1062–1065 (2012).
van der Laan, M., Horvath, S. E. & Pfanner, N. Mitochondrial contact site and cristae organizing system. Curr. Opin. Cell Biol. 41, 33–42 (2016).
Pernas, L. & Scorrano, L. Mito-morphosis: mitochondrial fusion, fission, and cristae remodeling as key mediators of cellular function. Annu. Rev. Physiol. 78, 505–531 (2016).
Wai, T. & Langer, T. Mitochondrial dynamics and metabolic regulation. Trends Endocrinol. Metab. 27, 105–117 (2016).
Jones, B. A. & Fangman, W. L. Mitochondrial DNA maintenance in yeast requires a protein containing a region related to the GTP-binding domain of dynamin. Genes Dev. 6, 380–389 (1992).
Alexander, C. et al. OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nat. Genet. 26, 211–215 (2000).
Delettre, C. et al. Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nat. Genet. 26, 207–210 (2000).
Wong, E. D. et al. The dynamin-related GTPase, Mgm1p, is an intermembrane space protein required for maintenance of fusion competent mitochondria. J. Cell Biol. 151, 341–352 (2000).
Meeusen, S. et al. Mitochondrial inner-membrane fusion and crista maintenance requires the dynamin-related GTPase Mgm1. Cell 127, 383–395 (2006).
Cipolat, S., Martins de Brito, O., Dal Zilio, B. & Scorrano, L. OPA1 requires mitofusin 1 to promote mitochondrial fusion. Proc. Natl Acad. Sci. USA 101, 15927–15932 (2004).
Ishihara, N., Fujita, Y., Oka, T. & Mihara, K. Regulation of mitochondrial morphology through proteolytic cleavage of OPA1. EMBO J. 25, 2966–2977 (2006).
Meeusen, S., McCaffery, J. M. & Nunnari, J. Mitochondrial fusion intermediates revealed in vitro. Science 305, 1747–1752 (2004).
Frezza, C. et al. OPA1 controls apoptotic cristae remodeling independently from mitochondrial fusion. Cell 126, 177–189 (2006).
Yamaguchi, R. et al. Opa1-mediated cristae opening is Bax/Bak and BH3 dependent, required for apoptosis, and independent of Bak oligomerization. Mol. Cell 31, 557–569 (2008).
Anand, R. et al. The i-AAA protease YME1L and OMA1 cleave OPA1 to balance mitochondrial fusion and fission. J. Cell Biol. 204, 919–929 (2014).
Faelber, K. et al. Crystal structure of nucleotide-free dynamin. Nature 477, 556–560 (2011).
Ford, M. G., Jenni, S. & Nunnari, J. The crystal structure of dynamin. Nature 477, 561–566 (2011).
Chappie, J. S., Acharya, S., Leonard, M., Schmid, S. L. & Dyda, F. G domain dimerization controls dynamin’s assembly-stimulated GTPase activity. Nature 465, 435–440 (2010).
Ingerman, E. et al. Dnm1 forms spirals that are structurally tailored to fit mitochondria. J. Cell Biol. 170, 1021–1027 (2005).
Ban, T., Heymann, J. A., Song, Z., Hinshaw, J. E. & Chan, D. C. OPA1 disease alleles causing dominant optic atrophy have defects in cardiolipin-stimulated GTP hydrolysis and membrane tubulation. Hum. Mol. Genet. 19, 2113–2122 (2010).
Kong, L. et al. Cryo-EM of the dynamin polymer assembled on lipid membrane. Nature 560, 258–262 (2018).
Reubold, T. F. et al. Crystal structure of the dynamin tetramer. Nature 525, 404–408 (2015).
Chiaruttini, N. et al. Relaxation of loaded ESCRT-III spiral springs drives membrane deformation. Cell 163, 866–879 (2015).
Gao, S. et al. Structure of myxovirus resistance protein a reveals intra- and intermolecular domain interactions required for the antiviral function. Immunity 35, 514–525 (2011).
Frohlich, C. et al. Structural insights into oligomerization and mitochondrial remodelling of dynamin 1-like protein. EMBO J 32, 1280–1292 (2013).
Kalia, R. et al. Structural basis of mitochondrial receptor binding and constriction by DRP1. Nature 558, 401–405 (2018).
Chappie, J. S. et al. A pseudoatomic model of the dynamin polymer identifies a hydrolysis-dependent powerstroke. Cell 147, 209–222 (2011).
Roux, A., Uyhazi, K., Frost, A. & De Camilli, P. GTP-dependent twisting of dynamin implicates constriction and tension in membrane fission. Nature 441, 528–531 (2006).
Antonny, B. et al. Membrane fission by dynamin: what we know and what we need to know. EMBO J. 35, 2270–2284 (2016).
Doublié, S. Preparation of selenomethionyl proteins for phase determination. Methods Enzymol. 276, 523–530 (1997).
Kabsch, W. XDS. Acta Cryst. D 66, 125–132 (2010).
Sparta, K. M., Krug, M., Heinemann, U., Mueller, U. & Weiss, M. S. Xdsapp2.0. J. Appl. Crystallogr. 49, 1085–1092 (2016).
Terwilliger, T. C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. D 65, 582–601 (2009).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Echols, N. et al. Graphical tools for macromolecular crystallography in PHENIX. J. Appl. Crystallogr. 45, 581–586 (2012).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Sievers, F. & Higgins, D. G. Clustal omega. Curr. Protoc. Bioinform. 48, 1.25.1–1.25.33 (2014).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Yofe, I. & Schuldiner, M. Primers-4-Yeast: a comprehensive web tool for planning primers for Saccharomyces cerevisiae. Yeast 31, 77–80 (2014).
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Ieva, R. et al. Mgr2 functions as lateral gatekeeper for preprotein sorting in the mitochondrial inner membrane. Mol. Cell 56, 641–652 (2014).
Morgenstern, M. et al. Definition of a high-confidence mitochondrial proteome at quantitative scale. Cell Rep. 19, 2836–2852 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Wilson-Kubalek, E. M., Brown, R. E., Celia, H. & Milligan, R. A. Lipid nanotubes as substrates for helical crystallization of macromolecules. Proc. Natl Acad. Sci. USA 95, 8040–8045 (1998).
Hagen, W. J. H., Wan, W. & Briggs, J. A. G. Implementation of a cryo-electron tomography tilt-scheme optimized for high resolution subtomogram averaging. J. Struct. Biol. 197, 191–198 (2017).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Castaño-Díez, D., Kudryashev, M., Arheit, M. & Stahlberg, H. Dynamo: a flexible, user-friendly development tool for subtomogram averaging of cryo-EM data in high-performance computing environments. J. Struct. Biol. 178, 139–151 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Whitford, P. C. et al. Excited states of ribosome translocation revealed through integrative molecular modeling. Proc. Natl Acad. Sci. USA 108, 18943–18948 (2011).
Noel, J. K. et al. SMOG 2: a versatile software package for generating structure-based models. PLOS Comput. Biol. 12, e1004794 (2016).
Harvey, M. J. & De Fabritiis, G. AceCloud: molecular dynamics simulations in the cloud. J. Chem. Inf. Model. 55, 909–914 (2015).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Theile, C. S. et al. Site-specific N-terminal labeling of proteins using sortase-mediated reactions. Nat. Protoc. 8, 1800–1807 (2013).
Meglei, G. & McQuibban, G. A. The dynamin-related protein Mgm1p assembles into oligomers and hydrolyzes GTP to function in mitochondrial membrane fusion. Biochemistry 48, 1774–1784 (2009).
Roux, A. et al. Membrane curvature controls dynamin polymerization. Proc. Natl. Acad. Sci. USA 107, 4141–4146 (2010).
Rujiviphat, J. et al. Mitochondrial genome maintenance 1 (Mgm1) protein alters membrane topology and promotes local membrane bending. J. Mol. Biol. 427, 2599–2609 (2015).
Mühleip, A. W. et al. Helical arrays of U-shaped ATP synthase dimers form tubular cristae in ciliate mitochondria. Proc. Natl Acad. Sci. USA 113, 8442–8447 (2016).
Tarasenko, D. et al. The MICOS component Mic60 displays a conserved membrane-bending activity that is necessary for normal cristae morphology. J. Cell Biol. 216, 889–899 (2017).
Barbot, M. et al. Mic10 oligomerizes to bend mitochondrial inner membranes at cristae junctions. Cell Metab. 21, 756–763 (2015).
Bohnert, M. et al. Central role of Mic10 in the mitochondrial contact site and cristae organizing system. Cell Metab. 21, 747–755 (2015).
Hessenberger, M. et al. Regulated membrane remodeling by Mic60 controls formation of mitochondrial crista junctions. Nat. Commun. 8, 15258 (2017).
Lee, H., Smith, S. B. & Yoon, Y. The short variant of the mitochondrial dynamin OPA1 maintains mitochondrial energetics and cristae structure. J. Biol. Chem. 292, 7115–7130 (2017).
Acknowledgements
This project was supported by ERC grants MitoShape (ERC-2013-CoG-616024 to O.D.) and ScaleCell (ERC- CoG-772230 to F.N.), grants from the Deutsche Forschungsgemeinschaft (SFB958/A12 and SFB740/C07 to O.D., SFB958/A04 and SFB740/D07 to F.N., SFB894/A20 to M.v.d.L., IRTG1830 to M.v.d.L. and F.W., SFB807 to R.S.), the Max Planck Society, a Humboldt fellowship to J.K.N., a pre-doctoral fellowship of the Boehringer Ingelheim Fonds to F.W., a Sofja Kovalevskaja Award from the Alexander von Humboldt Foundation to M.K., and a DOC Fellowship of the Austrian Academy of Sciences to M.H. We thank Y. Roske for help with crystallographic data collection, structure solution and Isothermal titration calorimetry measurements, T. Brandt for help and assistance in preparing cryo-EM samples, D. Mills for cryo-EM maintenance, B. Purfürst for support in the negative-stain EM analyses, T. Bock-Bierbaum for helpful comments on the manuscript, E. Werner from Research Network Services for his careful work on the videos, A. Xavier for help with Mgm1 fluorescence labelling, and the entire BESSY team for generous support during data collection at beamlines BL14.1, BL14.2 or BL14.3.
Peer review information
Nature thanks Harry Low, Tom Shemesh and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
K.F. designed the construct, grew the crystals and solved the structure. L.D. determined the cryo-ET reconstructions with support from A.M., R.S. and M.K.; J.K.N. and F.N. conducted and analysed molecular modelling and molecular dynamics simulations; F.W. and A.v.d.M. performed yeast-growth assays; and A.-K. P. together with N.C. carried out the tube-pulling assay. J.K.N., F.W. and A.-K.P. contributed equally to this study. J.S. purified the protein and J.S. and K.F. carried out the liposome co-sedimentation and GTPase assays; H.L. performed the analytical ultracentrifugation assays; E.R. and M.H. grew initial crystals of related Mgm1 constructs; C.M. and S.K. analysed yeast mitochondria using electron microscopy; K.F., L.D., J.K.N., C.M., A.R., M.v.d.L., W.K. and O.D. designed the research and interpreted structural data. K.F., L.D., J.K.N., M.v.d.L., W.K. and O.D. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Structure determination and analysis.
a, SDS–PAGE of recombinantly expressed and purified Mgm1. M, marker proteins; NI, whole-cell lysate, non-induced; I, whole-cell lysate, induced; R, whole-cell lysate, resuspended, collected cells; D, whole-cell lysate, disrupted cells; CL, cleared lysate; FT, flow-through; W, buffer wash; PC, after cleavage by PreScission Protease; L, as loaded onto gel filtration column (n = 5 independent experiments). b, Selenium sites and experimental density at 1.4σ before model building and refinement of the G domain (top left), stalk (top right), BSE (bottom left) and paddle domain (bottom right). c, Ribbon diagram of Mgm1 dimer, indicating the positions of confirmed methionines in ball-and-stick representation. Anomalous difference density is contoured at 2.5σ in magenta. An anomalous difference map was calculated from refined phases, resulting in discrete difference peaks indicating the positions of selenium atoms. Four selenium sites in the G domain, three in the BSE, two in the paddle domain and three in the stalk were used to determine the structure and verify the sequence assignment in the model. d, Mutations resulting in impaired lipid binding60 or in temperature-sensitive inner mitochondrial membrane fusion deficits10 were mapped onto the crystal structure. Mutations localize to the G interface, the G domain/BSE interface, stalk interface-1 or the paddle domain. e, Sequence conservation of nine Mgm1 sequences (see Supplementary Fig. 1 for alignments) was plotted on the surface of an Mgm1 monomer. Magenta, high conservation; cyan, low conservation. Residues investigated in this study are labelled and interfaces and contact sites are circled.
Extended Data Fig. 2 Comparison of Mgm1 and dynamin.
a, Monomers of Mgm1 (left) and dynamin (right) coloured by domain. b, The G domain and BSE domain of nucleotide-free Mgm1 and dynamin (grey, PDB: 5A3F) were superimposed on the BSE domains with a Cα root-mean-square deviation (r.m.s.d.) of 2.6 Å and 40% sequence identity. Both structures are in the closed state. The nucleotide-binding site is indicated. c, Superposition of the upper part of the stalk between Mgm1 and dynamin. In contrast to dynamin, the stalk in Mgm1 is kinked. d, Comparison between the stalk dimers of Mgm1 (left) and dynamin (right). In both proteins, the dimer buries a total surface area of 1,200 Å2. However, in Mgm1, interface-2 is shifted towards the paddle, resulting in a V-shaped dimer, whereas the dynamin dimer is X-shaped. e, Association of two dimers in the respective tetrameric crystal structures. In dynamin, the assembly of dimers occurs via two interfaces (interface-1 and interface-3), whereas only interface-1 is present in Mgm1.
Extended Data Fig. 3 Biochemical and negative-stain electron microscopy analysis.
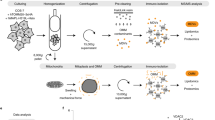
a, Liposome-binding assays (see also Figs. 1c, 2d) and quantification for Mgm1 mutants. Error bars indicate s.d. of 4 independent measurements. b, Isothermal titration calorimetry experiments showing binding of GTPγS to Mgm1 with a Kd of 9 ± 3 µM, binding number n = 1.01, deviation represents root-mean-square (r.m.s.) error of the fit (n = 1). c, Sedimentation equilibrium of wild-type Mgm1 (black) and Mgm1(F840D) (red) was performed at a protein concentration of 1 mg ml−1 at 8,000 r.p.m. and 20 °C. The protein distribution in the cell was monitored by absorbance at 280 nm. Solid lines represent fits to a molecular mass of Mr = 146 ± 6 kDa for wild-type Mgm1 and 78 ± 5 kDa for the Mgm1(F840D) (deviation represents r.m.s. error of the fit), indicating dimeric and monomeric association states at given conditions. The upper panel shows the original data and fits, the lower panels show the residuals from fit to data. d, GTPase assays using HPLC analysis. Error bars show s.d. of the mean of 4 independent experiments (each with 4 or 5 data points). e, Control experiments for negative-stain electron microscopy analysis of Mgm1-mediated membrane remodelling. Scale bars, 200 nm. f, Mgm1 binds to liposomes and forms tubes of different diameters with or without nucleotides present. Scale bars, 100 nm. g, Representative electron micrographs for Mgm1 mutants. Proteins with mutations in dimer interface-2 (F840D), in the membrane-binding site (R748A/K749A), the disulfide bond in the paddle domain (C812S and C821S) or in the putative paddle–paddle contact (F779D/S780D) show severe defects in tube formation or in the assembly of a regular liposome decoration compared to Mgm1. Scale bars, 100 nm. n = 2 independent experiments for e–g.
Extended Data Fig. 4 Yeast assays.
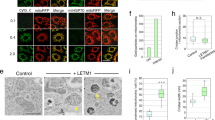
a, Schematic overview of yeast complementation experiments. In the presence of 2% glucose, expression of chromosomally encoded Mgm1 from the GAL1 promoter is suppressed. Yeast cells irreversibly lose the mitochondrial genome in the absence of Mgm1 (that is, become ρ0) and cannot switch to respiratory growth upon glucose depletion (as shown by the shift from low glucose conditions to the oxidation of ethanol produced during the fermentation of glucose). By co-expressing wild-type yeast Mgm1 or the corresponding Mgm1 mutants, functionality of the Mgm1 variants is assessed through various rescue parameters. b, Representative growth curve for the unmodified yeast strain transformed with an empty vector (e.v.), the engineered yeast strain (PGAL1-MGM1) complemented with yeast Mgm1 or an empty vector control (n = 3 independent experiments). c, Time-dependent expression of Mgm1, mitochondrially encoded cytochrome c oxidase subunit 1 (Cox1) and the nuclear-encoded mitochondrial heat shock protein Ssc1 (loading control) was assessed by western-blot analysis of isolated mitochondria upon transfer of yeast cells from a glucose-rich to a glucose-depleted medium containing 2% ethanol as the carbon source. = marks the long and short isoforms of Mgm1, ∼ is an unspecific band and * marks an Mgm1 degradation product (n = 2 independent experiments). Uncropped blots are shown in Supplementary Fig. 2. d, Western-blot analysis of isolated mitochondria from PGAL1-MGM1 yeast grown in glucose-containing medium containing plasmids that encode the respective mutant (n = 3 independent experiments). e, f, Yeast growth complementation assays with Mgm1 mutants containing mutations in the dimer interface and the paddle–paddle contacts. F805D in yeast corresponds to F840D in C. thermophilum and N675A corresponds to I700D. F779D/S780D in C. thermophilum corresponds to M745D/S746D in yeast. Representative growth curves are shown from n = 3 independent experiments. Data in Fig. 3b and Extended Data Fig. 4e are derived from the same experiment; the controls are shown in all graphs as a reference. g, Mitochondrial morphology of the indicated yeast strains was assessed by fluorescence microscopy. DNA and mitochondria were stained with DAPI and DiOC6, respectively. Three representative images from n = 2 independent cultures are shown. Dimensions of the images are 7.5 µm × 7.5 µm. h, Overexpression of Mgm1(Y520A) (with a mutation in the tetramer interface) leads to a strong dominant-negative effect on respiratory yeast growth (in media containing 3% glycerol as the carbon source). Representative growth curves are shown from n = 3 independent experiments. i, Overexpression of Mgm1(Y520A) leads to only a partial loss of mitochondrial DNA, as assayed by Cox1 expression. n = 3 independent experiments. j, k, Overexpression of dominant-negative Mgm1(Y520A) leads to a fragmentation of the mitochondrial network. Representative images and quantification of mitochondrial morphology in cells from n = 3 independent cultures, data displayed as mean ± s.d. l, Representative electron micrographs of yeast ultrathin sections assaying mitochondrial ultrastructure. Compared to mitochondria in wild-type yeast transformed with empty vector or pMgm1, mitochondria from cells expressing Mgm1(Y520A) showed a substantial loss of cristae and altered crista shape, as indicated by an increased diameter of the crista junctions and lumen and shorter crista length. Scale bars, 70 nm. m, Quantification of cristae morphology. WT+pMgm1: nmito = 208, ncristae = 132; WT+e.v.: nmito = 201, ncristae = 135; WT+pMgm1(Y520A): nmito = 202, ncristae = 81; 2 independent experiments. ***P < 0.0001 (Gaussian approximation); Mann–Whitney U-test (two-sided, 95% confidence interval); cristae number graph shows mean ± s.e.m.: WT+pMgm1: (4.8 ± 0.2) nm; WT+e.v.: (3.8 ± 0.2) nm; WT+pMgm1(Y520A): (1.4 ± 0.2) nm; crista length graph shows mean ± s.e.m.: WT+pMgm1: (153 ± 5) nm; WT+e.v.: (147 ± 5) nm; WT+pMgm1(Y520A): (115 ± 5) nm; crista diameter graph shows mean ± s.e.m.: WT+pMgm1 junction: (19.9 ± 0.5) nm; WT+e.v. junction: (21.0 ± 0.5) nm; WT+pMgm1(Y520A) junction: (26 ± 1) nm; WT+pMgm1 lumen: (24.7 ± 0.6) nm; WT+e.v. lumen: (25.8 ± 0.7) nm; WT+pMgm1(Y520A) lumen: (35 ± 2) nm.
Extended Data Fig. 5 Cryo-ET analysis.
a, b, f, g, Electron micrographs on the left show one tomographic slice of each sample. The density maps below obtained by subtomogram averaging are bandpass-filtered to the Fourier pixel value at 0.143 of the FSC curve. The masked FSC curves of each subtomogram average are indicated with resolutions obtained at 0.5 and 0.143 FSC. a, Mgm1 on the outside of a galactocerebroside-containing lipid tube in the apo form. On the right, a larger box size was used for processing in order to visualize the complete protein coat decorating the lipid tube. b, Mgm1 in the GTPγS-bound form on the outside of galactocerebroside-containing lipid tubes are very similar to the apo form, whereas nucleotide-free dynamin assembles differently compared to the guanosine-5′-[(β,γ)-methyleno]triphosphate-bound form22. c, GTPase assays of Mgm1 in the presence of lipid tubes containing galactocerebroside, n = 4, errors represent s.d. from the mean. d, Low-resolution cryo-ET reconstructions of GTPγS-bound Mgm1 assembled on the outside of Folch membrane tubes of different diameters, as measured between bilayer centres. On the basis of the pitch angle θ and the tube diameter d, the number of helical repeats (n-start) was estimated as n = 2πrtanθ/h, where the filament radius r = d/2+4 nm and the width from paddle tip to tip h is 13 nm. Although the basic filament architecture appears very similar, the filaments adapt their orientation to the curvature of the membrane tube. e, Representative electron micrographs showing Mgm1 coating the inner surface of a membrane tube (top) or both sides of the membrane tube (below). f, g, Cryo-ET reconstruction of Mgm1 in the apo and GTPγS-bound form on the inside of tubulated Folch liposomes, as in a and b. Grey scale bars, 10 nm; black scale bars, 100 nm.
Extended Data Fig. 6 Mgm1 tetramers in crystal and membrane lattices.
a–d, Mgm1 assemblies in the presence of GTPγS on the outer (a, c) and inner surface (b, d) of a membrane tube. a, b, Surface representations of flexibly fitted Mgm1 molecules, showing their arrangement in the protein lattice. c, d, Fit into the corresponding cryo-ET volume. Note that the membrane density and, consequently, the paddle–membrane contact, is more prominent in the GTPγS-bound form compared with the nucleotide-free form (Figs. 4b, 5a). e, Comparison of Mgm1 tetramers in the crystal lattice (blue) with tetramers fitted to the subtomogram average volumes obtained for the external (orange) and internal surface lattice (pink). Fitting the paddle and the BSE and G domains required only minor rearrangements. f, Tetramers in the crystal lattice pack into a linear assembly. Crystal contacts between two tetramers are mediated by the BSE domain of one tetramer (blue) and the stalk domain of the neighbouring tetramer (grey), resulting in an open interface-1. When comparing intra- and inter-tetramer interactions, BSE domain residues E533, E534 and Y537 in α2B bind to different sites of the adjacent stalks.
Extended Data Fig. 7 Molecular dynamics simulations.
a, Schematic of a 4-start helix. b, Mgm1 filaments in a 4-start helix, as in the cryo-ET volume on the outside of lipid tubes. The filament is defined as a continuous string of stalk domains connected by alternating interface-1 and interface-2. With this arrangement, filaments have a radius of 22 nm (axis to the centre of the stalk) and pitch of 54 nm. c, A string of dimers in contact through identical interfaces-1, as in the crystal structure, results in a left-handed helical arrangement with a large pitch, similar to the cryo-ET filament of the outside decoration. d, Snapshot of the stalk tetramer structure in the molecular dynamics simulation box. Analysis of the stalk tetramer conformation in molecular dynamics simulations gives information about the structural preferences of the filament in the absence of other domains. Geometrical parameters are drawn on the structure. d is the distance between the centres of mass of neighbouring dimers (marked as filled black circles). 95% of the variation in d is between 6.8 and 7.7 nm. v1–v4 are vectors pointing along each stalk monomer, defining angles θ1, θ2, and \({\theta }_{2}^{^{\prime} }\) as shown. α is the net in-plane rotation defined by v2 × v3, and is related to the local radius of curvature of a filament containing the tetramer. α can be simply written as a difference of the two interface angles, α = θ2 − θ1, where positive/negative α implies positive/negative curvature; θ2 > θ1 results in positive curvature and θ2 < θ1 results in negative curvature. β is the relative rotation angle of one dimer relative to the next, which controls the pitch and, therefore, the handedness of the helix. β is defined by the angle between the vectors v1 × v2 and v3 × v4 viewed along vf. vf is a unit vector in the direction of the filament defined by connecting the centres of mass of the two dimers. The elastic coordinates of a helical filament are the curvature κ and the twist τ. Positive/negative κ yields helices that bind to positive/negative membrane curvature. κ and τ can be approximately related to α and β, and the relations are indicated in the figure. e, Schematic of the curvature κ and the twist τ. For helices with a low pitch, κ is approximately the inverse radius of curvature (1/r). f, The angles θ1, θ2 and \({\theta }_{2}^{^{\prime} }\) are plotted over a portion (2.8 μs out of a total of 12 μs) of the simulation period. g, Distributions of θ1, θ2 and \({\theta }_{2}^{^{\prime} }\) over the whole simulation period. θ2 and \({\theta }_{2}^{^{\prime} }\) are, in principle, identical and the similarity of the distributions indicates sufficient sampling. In the crystal structure, θ1 = 123° and θ2/\({\theta }_{2}^{^{\prime} }\) = 142°/144°. The flexibilities of interface-1 and interface-2 are similar, as seen from the similar distribution widths. The peak of the θ1 distribution is centred on the parameters obtained for the crystal packing, whereas θ2/\({\theta }_{2}^{^{\prime} }\) is different, which may indicate that additional domain contacts present in the crystal stabilize a different configuration of interface-2. h, Using the relations shown in d, θ1 and θ2 at each snapshot are used to estimate the distribution of the curvature. The curvature distribution is centred near 0, which indicates that the stalk filament (at zero twist) prefers weakly curved or flat membranes. i, The angle β is plotted over a portion (2.8 μs out of a total of 12 μs) of the simulation period. j, k, The distributions of β (j) and τ (k) over the whole simulation period. A negative β or τ indicates that the stalk filament prefers a left-handed twist, but right-handed twists are thermally accessible. Note that no substantial correlation is seen between θ1, θ2/\({\theta }_{2}^{^{\prime} }\) and β.
Extended Data Fig. 8 Mgm1 attachment to membranes of different curvature.
Tube-pulling experiments, as described in Figs. 4d, 5c. Mgm1 was labelled with a fluorescein tag (green) and GUVs with Rhod-PE (red). Positive force is defined as pointing from the bead to the GUV. a, Tubes were pulled outward of single GUVs held by a micropipette (n = 8 independent experiments in the absence of GTP, n = 10 independent experiments in the presence of GTP). b, Representative time-lapse images of nucleation and growth of Mgm1 polymers on tubes pulled away from a GUV (right), and corresponding force measurements (left). c, Representative examples for tubes pulled into single GUVs adhering to the glass surface (n = 7 independent experiments in the absence of GTP, n = 7 independent experiments in the presence of GTP). d, Same as in b, but for tubes pulled into GUVs. ΔF is shown, as absolute forces were difficult to measure. Although Mgm1 covered the GUV surface in the experiments shown in c and d, it apparently did not oligomerize along the entire inward-pulled tube, as judged from the fluorescence signal. This probably reflects decreased diffusion of Mgm1 along the tube lumen. However, when the tube is not fully covered, a GTP-dependent shape change of the Mgm1 coat in the tube would not induce a force, as previously demonstrated for dynamin61. Therefore, the force increase probably results from the GTP-dependent remodelling of the Mgm1 coat on the GUV. In the case of outer decoration, Mgm1 oligomerizes on the tube and the GUV. In this case, the force increase can be caused by GTP-dependent alterations of the Mgm1 coat on the GUV and/or tube expansion. We note that these experiments gave no hint of GTP-driven constriction of membrane tubes.
Extended Data Fig. 9 Model of Mgm1 action.
a, On the basis of the close similarity of the G domains and BSE domains of Mgm1 and dynamin (Extended Data Fig. 2b), we propose that Mgm1 and dynamin perform similar power strokes. Dimerization of the G domain would link neighbouring Mgm1 filaments. The power stroke would then result in negative torque in the direction of the membrane normal. In b, a circle with a dot indicates a vector towards the viewer and a circle with an x indicates vector in the opposite direction. The arrow represents the direction of the torque. Note that power-stroke torque is independent of membrane curvature and helix handedness. During the power stroke, the helix pitch remains constant because of the G domain contacts. Unwinding or winding of filaments then translates into a change in helix diameter. Inter-paddle contacts must be weak or absent as the filaments slide past each other. b, The power-stroke torque applies an equal and opposite force between neighbouring turns. For outside decoration, the surface normal points outward. The resulting forces would constrict a right-handed helix and expand a left-handed helix. For inside decoration, the surface normal points inward, reversing the sign of the power-stroke torque. This reverses the resultant forces on the filament, which would expand a right-handed helix and constrict a left-handed helix. See also Supplementary Video 1. c, Modelling an example helical Mgm1 filament on an inner-tube surface. Although the Mgm1 tetramer on the inside lattice observed by cryo-ET resembled the crystal tetramer closely, formation of a continuous filament on the inside of a narrow tube would require curvature changes in the tetramer relative to the crystal structure. Using an all-atom structure-based model, we explore how the tetramer structure might change as part of a tight filament. The modelling parameters ensured that a short filament (4 dimers) fits within the steric constraints of a 30-nm-radius tube, and that the pitch results in a 1-start helix (left-handed pitch angle of 3.6°). Otherwise, the shape of the tetramer is free to find its optimal shape. Changes in the interface bending angles result in a transition from positive curvature (θ2 > θ1) to negative curvature (θ2 < θ1) (Extended Data Fig. 7d). d, Comparison of the constrained tetramer shown in c (central dimers) with the crystal structure. Minor changes in interface-1 and larger changes in interface-2 (with minimal changes to atomic packing, see insets) enable a conformational switch within the tetramer from binding to a concave surface (as in the crystal packing geometry) to binding to a convex surface. In this case, θ1 = 128° and θ2 = 117°. See also Extended Data Fig. 7g for comparison to explicit solvent simulations. e, Schematic overview of mitochondrial inner membrane remodelling. f–h, Models of mitochondrial membrane remodelling by Mgm1 and OPA1 filaments. f, During inner-membrane fusion, Mgm1 or OPA1 filaments may assemble on opposing membrane buds to stabilize the membrane curvature at the fusion site, as previously proposed62. g, On the inner surface of cristae, Mgm1 or OPA1 filaments may assemble into left-handed helical filaments to constrict the crista junction in a GTPase-dependent fashion. Alternatively, they may assemble into right-handed helical filaments that expand the crista volume to prevent their collapse. In this way, Mgm1 filaments may counteract the membrane-constricting activity of the ATPase synthase dimers63 or the MICOS complex64,65,66,67 to pull lipids into cristae and enable the dynamic transition from a tight crista state with reduced oxidative phosphorylation to an expanded active state with high oxidative phosphorylation activity. In agreement with this model, cristae have been shown to collapse when a GTPase-deficient OPA1 variant is expressed14. h, Similar to dynamin assemblies at the neck of clathrin-coated pits, Mgm1 or OPA1 may assemble in a right-handed helix around the neck of an inner membrane junction, resulting in constriction and membrane scission upon GTP hydrolysis. The assembly geometry of the Mgm1 or OPA1 filaments may depend on lipid composition, interaction partners or the specific Mgm1 or OPA1 isoform. Consistent with the latter assumption, inner membrane fusion requires the long form of OPA1, but the short OPA1 isoforms are sufficient for stabilizing crista membranes68.
Supplementary information
Supplementary Figure 1
Sequence alignment of Mgm1 proteins.
Supplementary Figure 2
Uncropped SDS PAGE gels and Western blots.
Supplementary Information
This zipped folder contains source data for Extended Data Fig 4.
Supplementary Video 1
Membrane remodelling models of Mgm1 filaments Effects of handedness and assembly geometry of the Mgm1 helix for a GTPase-driven power stroke.
Rights and permissions
About this article
Cite this article
Faelber, K., Dietrich, L., Noel, J.K. et al. Structure and assembly of the mitochondrial membrane remodelling GTPase Mgm1. Nature 571, 429–433 (2019). https://doi.org/10.1038/s41586-019-1372-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1372-3
This article is cited by
-
In situ architecture of Opa1-dependent mitochondrial cristae remodeling
The EMBO Journal (2024)
-
Stay in touch with the endoplasmic reticulum
Science China Life Sciences (2024)
-
Structural mechanism of mitochondrial membrane remodelling by human OPA1
Nature (2023)
-
OPA1 helical structures give perspective to mitochondrial dysfunction
Nature (2023)
-
Structural insights into the mechanism of GTP initiation of microtubule assembly
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.