Abstract
Experimental models of the human brain are needed for basic understanding of its development and disease1. Human brain organoids hold unprecedented promise for this purpose; however, they are plagued by high organoid-to-organoid variability2,3. This has raised doubts as to whether developmental processes of the human brain can occur outside the context of embryogenesis with a degree of reproducibility that is comparable to the endogenous tissue. Here we show that an organoid model of the dorsal forebrain can reliably generate a rich diversity of cell types appropriate for the human cerebral cortex. We performed single-cell RNA-sequencing analysis of 166,242 cells isolated from 21 individual organoids, finding that 95% of the organoids generate a virtually indistinguishable compendium of cell types, following similar developmental trajectories and with a degree of organoid-to-organoid variability comparable to that of individual endogenous brains. Furthermore, organoids derived from different stem cell lines show consistent reproducibility in the cell types produced. The data demonstrate that reproducible development of the complex cellular diversity of the central nervous system does not require the context of the embryo, and that establishment of terminal cell identity is a highly constrained process that can emerge from diverse stem cell origins and growth environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
scRNA-seq data that support the findings of this study have been deposited at Gene Expression Omnibus with accession number GSE129519, and at the Single Cell Portal (https://portals.broadinstitute.org/single_cell/study/reproducible-brain-organoids).
Code availability
The code used for data analysis is available on GitHub at https://github.com/AmandaKedaigle/BrainOrganoidsReproducibility.
References
Silbereis, J. C., Pochareddy, S., Zhu, Y., Li, M. & Sestan, N. The cellular and molecular landscapes of the developing human central nervous system. Neuron 89, 248–268 (2016).
Quadrato, G., Brown, J. & Arlotta, P. The promises and challenges of human brain organoids as models of neuropsychiatric disease. Nat. Med. 22, 1220–1228 (2016).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53 (2017).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl Acad. Sci. USA 110, 20284–20289 (2013).
Yoon, S. J. et al. Reliability of human cortical organoid generation. Nat. Methods 16, 75–78 (2019).
Pollen, A. A. et al. Establishing cerebral organoids as models of human-specific brain evolution. Cell 176, 743–756 (2019).
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Lancaster, M. A. et al. Guided self-organization and cortical plate formation in human brain organoids. Nat. Biotechnol. 35, 659–666 (2017).
Bagley, J. A., Reumann, D., Bian, S., Lévi-Strauss, J. & Knoblich, J. A. Fused cerebral organoids model interactions between brain regions. Nat. Methods 14, 743–751 (2017).
Xiang, Y. et al. Fusion of regionally-specified hPSC-derived organoids models human brain development and interneuron migration. Cell Stem Cell 21, 383–398 (2017).
Birey, F. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017).
Bershteyn, M. et al. Human iPSC-derived cerebral organoids model cellular features of lissencephaly and reveal prolonged mitosis of outer radial glia. Cell Stem Cell 20, 435–449 (2017).
Rigamonti, A. et al. Large-scale production of mature neurons from human pluripotent stem cells in a three-dimensional suspension culture system. Stem Cell Reports 6, 993–1008 (2016).
Church, G. M. The personal genome project. Mol. Syst. Biol. 1, 0030 (2005).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierachies of the human cortex. Science 358, 1318–1323 (2017).
Molyneaux, B. J. et al. Novel subtype-specific genes identify distinct subpopulations of callosal projection neurons. J. Neurosci. 29, 12343–12354 (2009).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Lodato, S. & Arlotta, P. Generating neuronal diversity in the mammalian cerebral cortex. Annu. Rev. Cell Dev. Biol. 31, 699–720 (2015).
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Kohwi, M. et al. A subpopulation of olfactory bulb GABAergic interneurons is derived from Emx1- and Dlx5/6-expressing progenitors. J. Neurosci. 27, 6878–6891 (2007).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Fan, X. et al. Spatial transcriptomic survey of human embryonic cerebral cortex by single-cell RNA-seq analysis. Cell Res. 28, 730–745 (2018).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030 (2018).
Sheridan, S. D. et al. Epigenetic characterization of the FMR1 gene and aberrant neurodevelopment in human induced pluripotent stem cell models of fragile X syndrome. PLoS ONE 6, e26203 (2011).
Sugathan, A. et al. CHD8 regulates neurodevelopmental pathways associated with autism spectrum disorder in neural progenitors. Proc. Natl Acad. Sci. USA 111, E4468–E4477 (2014).
Velasco, S., Paulsen, B. & Arlotta, P. Highly reproducible human brain organoids recapitulate cerebral cortex cellular diversity. Protocol Exchange https://doi.org/10.21203/rs.2.9542/v1 (2019).
Quadrato, G., Sherwood, J. L. & Arlotta, P. Long term culture and electrophysiological characterization of human brain organoids. Protocol Exchange https://doi.org/10.1038/protex.2017.049 (2017).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Arya, S., Mount, D., Kemp, S. E. & Jefferis, G. RANN: Fast nearest neighbour search (wraps ANN library) using l2 metric. R package version 2.6. (2018).
McDavid, A., Finak, G. & Yajima, M. MAST: model-based analysis of single cell transcriptomics. R package version 1.6.1. (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R News 2, 18–22 (2002).
Acknowledgements
We thank J. R. Brown and C. Gerhardinger (from the P.A. laboratory) for insightful discussions and editing of the manuscript; T. Nguyen, V. Jokhi, A. S. Shetty, J. Stogsdill, D. Di Bella, X. Jin (from the P.A. laboratory) and K. Shekhar (from the A.R. laboratory) for help with scRNA-seq analysis and classification; M. Cuoco, J. Waldman, and D. Dionne (from the A.R. laboratory) for help with scRNA-seq library preparation; the Eggan laboratory for providing the 11a line; the Talkowski laboratory for the GM08330 line; and the Cohen laboratory for the Mito 210 line. This work was supported by grants from the Stanley Center for Psychiatric Research, the Broad Institute of MIT and Harvard, the National Institutes of Health (R01-MH112940 to P.A. and J.Z.L.; P50MH094271 and U01MH115727 to P.A.), the Klarman Cell Observatory to J.Z.L. and A.R., and the Howard Hughes Medical Institute to A.R. This work is dedicated to the memory of Yoshiki Sasai, who was a pioneer of the field of brain organoids and first established the protocol that we build upon in this study.
Author information
Authors and Affiliations
Contributions
P.A., S.V. and A.J.K. conceived the experiments. S.V. generated all organoids. S.V., A.N. and M.R. cultured and characterized all organoids and P.A. supervised their work. S.V. performed scRNA-seq experiments, with help from X.A., G.Q., B.P. and L.N.; A.J.K., S.K.S. and J.Z.L. performed scRNA-seq analysis and J.Z.L. and A.R. supervised their work. P.A., S.V. and A.J.K. wrote the manuscript with contributions from all authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
A.R. is an SAB member of Thermo Fisher Scientific and Syros Pharmaceuticals and a co-founder and equity holder of Celsius Therapeutics. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Comparison of organoids and spheroids generated by different protocols.
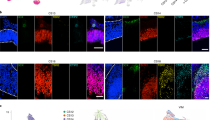
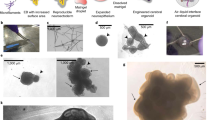
a, From left: self-patterned whole-brain organoids3, dorsally patterned forebrain organoids (protocol modified from ref. 5), and dorsal and ventral forebrain spheroids (protocol modified from ref. 15). All models are generated from the PGP1 line and cultured for six months. b, Immunohistochemistry for neuronal (MAP2), dorsal forebrain progenitor (EMX1 and PAX6), CFuPN (CTIP2), and CPN (SATB2) markers, across a time course from one to six months. Scale bars: whole organoids (Whole Org), 200 μm; others, 50 μm.
Extended Data Fig. 2 Analysis of cell-type-specific markers in organoids derived from different lines.
a, Expression of selected marker genes used in cell-type identification. Violin plots show distribution of normalized expression in cells from CCA-aligned organoids at three months (n = 9 individual organoids from three batches). CFuPNS: n = 15,866 cells; CPNs, n = 18,905 cells; cycling, n = 4,035 cells; immature INs, n = 353 cells; immature PNs, n = 6,727 cells; IPCs, n = 4,276 cells; oRG, n = 5,436 cells; and RG, n = 3,318 cells. Scale: normalized read counts. b, Immunohistochemistry for neuronal (MAP2), dorsal forebrain progenitor (EMX1 and PAX6), CFuPN (CTIP2), CPN (SATB2), radial glia (SOX2) and proliferation (Ki67) markers in PGP1 (b3), 11a, GM08330 and HUES66 organoids at three months. c, RNA in situ hybridization for markers of IPCs (EOMES, also known as TBR2), Cajal–Retzius (RELN) and post-mitotic PNs (TBR1) in three-month PGP1 (b3), 11a, GM08330 and HUES66 organoids. d, Immunohistochemistry for markers of forebrain progenitors (FOXG1), outer radial glia (HOPX), post-mitotic PNs (TBR1) and IPCs (TBR2) in PGP1 (b3) organoids at three and six months. Scale bars, 60 μm (b, c, d).
Extended Data Fig. 3 Evaluation of apoptosis, hypoxia and doublets in organoid scRNA-seq data.
a, b, t-SNE plots showing average scaled expression of all genes from the apoptosis (a) and hypoxia (b) mSigDB hallmark gene sets in PGP1 (b1) organoids at three (left) and six (right) months. n = 3 individual organoids per time point. c, d, Histograms showing number of cells expressing apoptosis markers (c) and hypoxia markers (d) in PGP1 organoids at three and six months. The x axis indicates average scaled expression of all genes in the corresponding mSigDB hallmark gene set. The similarity in markers of hypoxia and apoptosis between the three- and six-month single-cell data indicates that the growth conditions of this protocol preserve the health of the tissue over many months in culture. e, Immunohistochemistry of a six-month PGP1 organoid for the apoptotic marker cleaved caspase 3, on the perimeter (box 1) and in the centre (box 2) of an organoid. Scale bars, 500 μm (top), 100 μm (bottom). f, Multiplet detection using the Scrublet39 program. Scores represent the probability that the ‘cell’ represents a droplet containing more than one cell. From left, t-SNE plots of PGP1 (two batches: b1, b2; n = 3 individual organoids for b1, and n = 3 individual organoids for b2) and HUES66 (n = 3 individual organoids) organoids at three months; and 11a (n = 3 individual organoids), GM08330 (n = 3 individual organoids) and PGP1 (two batches: b1, b3; n = 3 individual organoids for b1, and n = 2 individual organoids for b3) organoids at six months. Overall, 2–8% of cells were predicted to be multiplets, which is consistent with the expected multiplet rate (~5%) given our loadings of approximately 10,000 cells per channel35.
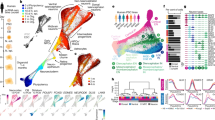
Extended Data Fig. 4 Cell types in individual organoids are generated following a precise and reproducible temporal order and are transcriptionally similar to human fetal cortex cell types.
a, t-SNE plots produced by Monocle2, showing the contribution of cells from individual PGP1 (b1) organoids at three months (n = 2,665, 3,094 and 2,264 cells from organoids 1–3) and six months (n = 3,959, 2,971 and 3,042 cells from organoids 16–18) to plots in Fig. 3a, b. b, Agreement between cell-type classifications in cell populations of 11a, GM08330 and PGP1 organoids (two batches: b1, b3) at six months with cell types described in a previously published scRNA-seq dataset of the human fetal cortex18. Dot size and colour intensity indicate the percentage of organoid cells in each cell cluster assigned to each human cortex cell type by a random forest classifier.
Extended Data Fig. 5 Dorsally patterned forebrain organoids show reproducibility similar to that of endogenous brain, as compared to self-patterned whole-brain organoids.
a, b, Percentage distribution of cell types in individual three-month PGP1 (two batches: b1, b2; n = 3 per batch) and HUES66 (one batch; n = 3) dorsally patterned forebrain organoids (a; left), and individual six-month 11a, GM08330 (one batch each; n = 3 per batch), and PGP1 (two batches: b1, b3, n = 3 per batch) dorsally patterned forebrain organoids (a; right), versus individual six-month self-patterned whole-brain organoids3 (b). c–e, Distribution of cell types as assigned in the original publication across individual samples of fetal human cortex29 (c), adult human cortex samples from distinct individuals28 (d), and adult mouse cortex samples from distinct individuals30 (e). DG, dentate gyrus; CGE, caudal ganglionic eminence; MGE, medial ganglionic eminence.
Supplementary information
Supplementary Information
Supplementary Info Note 1: Correcting for ambient RNA contamination improves co-clustering of organoids in the 3 month HUES66 and PGP1 batch 2 datasets.
Supplementary Information
Supplementary Info Note 2: A step-by-step protocol for organoid culture. A version of this protocol can also be found in Nature Protocol Exchange.
Supplementary Info Table 1
Differentially expressed genes for each cluster of single cells from organoids. Results of running differential expression tests on data from organoids from the PGP1 (two batches: b1, b2; n=3 organoids for each) and HUES66 (n=3 organoids) at 3 months; and organoids from the 11a (n=3 organoids), GM08330 (n=3 organoids), and PGP1 (two batches: b1, b3; n=3 organoids for each) at 6 months. Differential expression was calculated using the MAST algorithm as implemented in the Seurat R package. Adjusted p values are reported after Bonferroni correction based on the total number of genes in the dataset.
Rights and permissions
About this article
Cite this article
Velasco, S., Kedaigle, A.J., Simmons, S.K. et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature 570, 523–527 (2019). https://doi.org/10.1038/s41586-019-1289-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1289-x
This article is cited by
-
Exosome lncRNA IFNG-AS1 derived from mesenchymal stem cells of human adipose ameliorates neurogenesis and ASD-like behavior in BTBR mice
Journal of Nanobiotechnology (2024)
-
A cell fate decision map reveals abundant direct neurogenesis bypassing intermediate progenitors in the human developing neocortex
Nature Cell Biology (2024)
-
BANKSY unifies cell typing and tissue domain segmentation for scalable spatial omics data analysis
Nature Genetics (2024)
-
A patterned human neural tube model using microfluidic gradients
Nature (2024)
-
Comparing stem cells, transdifferentiation and brain organoids as tools for psychiatric research
Translational Psychiatry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.