Abstract
The evolution of female mating preferences for harmful male traits is a central paradox of sexual selection1,2,3,4,5,6,7,8,9. Two dominant explanations for this paradox8,10 are Fisher’s runaway process, which is based on genetic correlations between preference and trait1,3,4, and Zahavi’s handicap principle, in which the trait is an honest costly signal of male quality2,6,8,11. However, both of these explanations require the exogenous initial spread of female preferences before harmful male traits can evolve1,2,3,4,6,8,11. Here I present a mechanism for the evolution of female mating preferences for harmful male traits that is based on the selfish evolutionary interests of sex chromosomes. I demonstrate that female-biased genetic elements—such as the W and X sex chromosomes—will evolve mating preferences for males who display traits that reduce their fitness and/or that of their male offspring, but increase fitness in female offspring. In particular, W-linked preferences can cause nearly lethal male traits to sweep to fixation. Sex-linked preferences can drive the evolution of traits such as ornamental handicaps and male parental care, and can explain variation in ornamentation and behaviour across taxa with divergent sex-determining mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
No datasets were generated or analysed in this study.
Code availability
Simulation code is available upon request.
References
Fisher, R. A. The Genetical Theory of Natural Selection (Oxford Univ. Press, 1930).
Zahavi, A. Mate selection—a selection for a handicap. J. Theor. Biol. 53, 205–214 (1975).
Lande, R. Models of speciation by sexual selection on polygenic traits. Proc. Natl Acad. Sci. USA 78, 3721–3725 (1981).
Kirkpatrick, M. Sexual selection and the evolution of female choice. Evolution 36, 1–12 (1982).
Trivers, R. Social Evolution (Benjamin Cummings, 1985).
Grafen, A. Biological signals as handicaps. J. Theor. Biol. 144, 517–546 (1990).
Andersson, M. B. Sexual Selection (Princeton Univ. Press, 1994).
Pomiankowski, A. N. in Oxford Surveys in Evolutionary Biology (eds Harvey, P. H. & Partridge, L.) 136–184 (Oxford Univ. Press, 1988).
Albert, A. Y. K. & Otto, S. P. Sexual selection can resolve sex-linked sexual antagonism. Science 310, 119–121 (2005).
Kokko, H., Brooks, R., McNamara, J. M. & Houston, A. I. The sexual selection continuum. Proc. R. Soc. Lond. B 269, 1331–1340 (2002).
Pomiankowski, A. Sexual selection: the handicap principle does work–sometimes. Proc. R. Soc. Lond. B 231, 123–145 (1987).
Burt, A. & Trivers, R. Genes in Conflict : The Biology of Selfish Genetic Elements (Belknap, 2006).
Haig, D., Úbeda, F. & Patten, M. M. Specialists and generalists: the sexual ecology of the genome. Cold Spring Harb. Perspect. Biol. 6, a017525 (2014).
Bachtrog, D. et al. Sex determination: why so many ways of doing it? PLoS Biol. 12, e1001899 (2014).
Rice, W. R. Sex chromosomes and the evolution of sexual dimorphism. Evolution 38, 735–742 (1984).
van Doorn, G. S. Intralocus sexual conflict. Ann. NY Acad. Sci. 1168, 52–71 (2009).
Cox, R. M. & Calsbeek, R. Sexually antagonistic selection, sexual dimorphism, and the resolution of intralocus sexual conflict. Am. Nat. 173, 176–187 (2009).
Innocenti, P. & Morrow, E. H. The sexually antagonistic genes of Drosophila melanogaster. PLoS Biol. 8, e1000335 (2010).
Cheng, C. & Kirkpatrick, M. Sex-specific selection and sex-biased gene expression in humans and flies. PLoS Genet. 12, e1006170 (2016).
Seger, J. & Trivers, R. Asymmetry in the evolution of female mating preferences. Nature 319, 771–773 (1986).
Kirkpatrick, M. & Hall, D. W. Sexual selection and sex linkage. Evolution 58, 683–691 (2004).
Hastings, I. M. Manifestations of sexual selection may depend on the genetic basis of sex determination. Proc. R. Soc. Lond. B 258, 83–87 (1994).
Reeve, H. K. & Pfennig, D. W. Genetic biases for showy males: are some genetic systems especially conducive to sexual selection? Proc. Natl Acad. Sci. USA 100, 1089–1094 (2003).
Clutton-Brock, T. H. The Evolution of Parental Care (Princeton Univ. Press, 1991).
Reeve, H. K. & Shellman-Reeve, J. S. The general protected invasion theory: sex biases in parental and alloparental care. Evol. Ecol. 11, 357–370 (1997).
Spence, M. Job market signaling. Q. J. Econ. 87, 355–374 (1973).
Berset-Brändli, L., Jaquiéry, J., Broquet, T., Ulrich, Y. & Perrin, N. Extreme heterochiasmy and nascent sex chromosomes in European tree frogs. Proc. R. Soc. Lond. B 275, 1577–1585 (2008).
Otto, S. P. et al. About PAR: the distinct evolutionary dynamics of the pseudoautosomal region. Trends Genet. 27, 358–367 (2011).
Werren, J. H., Baldo, L. & Clark, M. E. Wolbachia: master manipulators of invertebrate biology. Nat. Rev. Microbiol. 6, 741–751 (2008).
Funkhouser, L. J. & Bordenstein, S. R. Mom knows best: the universality of maternal microbial transmission. PLoS Biol. 11, e1001631 (2013).
Ezenwa, V. O., Gerardo, N. M., Inouye, D. W., Medina, M. & Xavier, J. B. Animal behavior and the microbiome. Science 338, 198–199 (2012).
Edward, D. A. & Chapman, T. The evolution and significance of male mate choice. Trends Ecol. Evol. 26, 647–654 (2011).
Mank, J. E., Hall, D. W., Kirkpatrick, M. & Avise, J. C. Sex chromosomes and male ornaments: a comparative evaluation in ray-finned fishes. Proc. R. Soc. Lond. B 273, 233–236 (2006).
Acknowledgements
I thank C. Veller for research assistance and comments on the manuscript. I am grateful to D. Haig, R. Trivers and J. Losos for comments on the manuscript, S. Otto, M. Nowak, M. Zuk, L. Hadany and J. Boyle for helpful discussions, and C. Noble for help with figure preparation. The simulations in this paper were run on the Odyssey cluster supported by the FAS Division of Science, Research Computing Group at Harvard University. I am supported by an NSF graduate research fellowship.
Reviewer information
Nature thanks Andrew Pomiankowski and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Long-term frequencies of sex-linked preferences and traits.
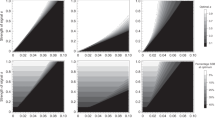
Frequencies of a W-linked, Z-linked and X-linked mutant P allele and an autosomal mutant T allele after 106 generations, each having started at 1% frequency, in Hardy–Weinberg and linkage equilibrium. The strength of the preference is α = 1.5 (top) or α = 5 (bottom). hT is the dominance of the T allele with respect to the wild-type t allele; hP is the dominance of the P allele with respect to the wild-type p allele. In the case of an X-linked preference, I assume that hP = hT; in the case of W-linked and Z-linked preferences, hP is not applicable, as both W- and Z-linked preferences are hemizygous in females. Note that in the case of a W-linked preference, the P allele will eventually attain high frequency in parameter regions in which it does not appear to do so here; for example, compare the results here for a W-linked preference of strength α = 1.5 with Fig. 1c, in which frequencies after 5 × 106 generations are reported.
Extended Data Fig. 2 Two equilibria for W-linked preferences for sexually antagonistic traits.
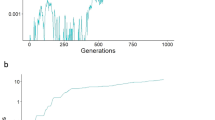
Frequency trajectories of the W-linked P allele and an autosomal, male-costly female-beneficial T allele under two strengths of the preference. The heat maps displayed here are also shown in Fig. 1b (top) and Fig. 1d (bottom), and their details are described in the Methods. a, Sample trajectories of P and T alleles when the preference is weak and the cost of the trait to males is large. The P allele fixes but the T allele remains at a low-frequency mutation–selection balance: the equilibrium is one in which all females prefer males that display the costly trait, but very few males display it. b, Sample trajectories of P and T alleles when the preference is strong. The P allele fixes and the T allele attains a very high-frequency mutation–selection balance: the equilibrium is one in which almost all males have low viability, and all females strongly prefer the low-viability males.
Extended Data Fig. 3 Suppression of selfish W-linked preferences for sexually antagonistic traits.
Trajectories of the W-linked P allele, the autosomal, male-costly female-beneficial T allele and an autosomal S allele that suppresses the preference allele, across various fitness effects of the trait. The mutation rate at the trait locus is 10−3 and the T allele is co-dominant (hT = 1/2). In each simulation, at generation 0 the T and P alleles are introduced into the population. After 5 × 105 generations, a mutant S allele appears at an autosomal locus. The simulation is run for an additional 1.5 × 106 generations. Each allele is introduced at frequency 1%, in Hardy–Weinberg equilibrium if at a diploid locus, and in linkage equilibrium with respect to the other loci. The autosomal suppressor locus is unlinked to the trait locus, and the S allele is co-dominant (hS = 1/2), so that a female bearing the P allele and a single S allele has preferences 1, \({\alpha }_{i}^{1/4}\) and \({\alpha }_{i}^{1/2}\) for tt, Tt and TT individuals, respectively, whereas a female with the P allele and no S allele has preferences 1, \({\alpha }_{i}^{1/2}\) and αi for tt, Tt and TT individuals. Suppression is more likely to evolve when the strength of the W-linked preference is weak, and the average fitness cost of the trait across males and females is high.
Extended Data Fig. 4 Arms-race dynamics between W-linked preferences and their suppressors.
A weak preference allele P1 initially invades and fixes, which pushes the sexually antagonistic T allele (sf = 0.01, sm = 0.1) to intermediate frequency. At 5 × 105 generations, a mutant allele that suppresses the action of P1 appears at an unlinked locus. The suppressor invades and fixes, which eliminates the effect of P1 so that the T allele decreases to a low frequency. At 2 × 106 generations, a medium-strength preference allele P2 invades and fixes, which pushes T back to a high frequency. An unlinked suppressor of P2 appears at 2.5 × 106 generations, but immediately goes extinct: the medium-strength preference is evolutionarily resistant to suppression.
Extended Data Fig. 5 Dynamics of preferences in the pseudo-autosomal region.
Trajectories of the pseudo-autosomal mutant P allele and autosomal mutant T allele in a ZW system, for various strengths of the preference, fitness effects of the trait (always costly in males and beneficial in females) and recombination rates between the preference locus and the sex-determining locus. The mutant trait allele is co-dominant (hT = 1/2). In each case, there is some (low) threshold recombination rate, below which the preference and trait can evolve to high frequency and above which they cannot.
Supplementary information
Supplementary Information
This file contains supplementary information S1-S7 which includes Figures S1-S4 and Table S1.
Rights and permissions
About this article
Cite this article
Muralidhar, P. Mating preferences of selfish sex chromosomes. Nature 570, 376–379 (2019). https://doi.org/10.1038/s41586-019-1271-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1271-7
This article is cited by
-
Evolutionary and demographic consequences of temperature-induced masculinization under climate warming: the effects of mate choice
BMC Ecology and Evolution (2021)
-
Sex chromosomes manipulate mate choice
Nature (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.