Abstract
Broadly neutralizing monoclonal antibodies protect against infection with HIV-1 in animal models, suggesting that a vaccine that elicits these antibodies would be protective in humans. However, it has not yet been possible to induce adequate serological responses by vaccination. Here, to activate B cells that express precursors of broadly neutralizing antibodies within polyclonal repertoires, we developed an immunogen, RC1, that facilitates the recognition of the variable loop 3 (V3)-glycan patch on the envelope protein of HIV-1. RC1 conceals non-conserved immunodominant regions by the addition of glycans and/or multimerization on virus-like particles. Immunization of mice, rabbits and rhesus macaques with RC1 elicited serological responses that targeted the V3-glycan patch. Antibody cloning and cryo-electron microscopy structures of antibody–envelope complexes confirmed that immunization with RC1 expands clones of B cells that carry the anti-V3-glycan patch antibodies, which resemble precursors of human broadly neutralizing antibodies. Thus, RC1 may be a suitable priming immunogen for sequential vaccination strategies in the context of polyclonal repertoires.
Similar content being viewed by others
Main
Single-cell antibody cloning from human donors who are infected with HIV-1 revealed that broadly neutralizing antibodies (bNAbs) have undergone unusually extensive somatic mutations1,2,3,4. Moreover, the high degree of somatic mutations is essential for binding to the native HIV-1 envelope (Env) spike and for the neutralizing activity of bNAbs5. The accumulation of large numbers of mutations suggests that bNAbs evolve in response to iterative rounds of somatic hypermutation and selection in germinal centres6. Studies in humans revealed that this occurs in response to viral escape variants that arise from antibody pressure4. Together, these observations suggest that vaccination to elicit bNAbs requires a series of sequential immunogens starting with an immunogen that induces the expansion7 of B lymphocytes expressing appropriate germline precursors8.
Sequential immunization to guide the development of bNAbs was demonstrated in genetically modified mice that carry inferred germline precursors of human bNAbs8,9. However, the priming immunogens that were used to initiate the response failed to activate and expand B cells that expressed inferred bNAb precursors in animals with polyclonal antibody repertoires. Thus, a goal of HIV-1 vaccine development has been to design immunogens that recruit B cells that express bNAb precursors into germinalcentre reactions in animals with polyclonal repertoires.
The germline-targeting approach to immunogen design focuses on producing immunogens that bind with high affinity to specific bNAb precursors—the rationale being that the recruitment of B cells to germinal centres is in part dependent on receptor affinity for antigen10,11,12,13,14. However, this methodology effectively limits the repertoire of recruited B cells qualitatively and quantitatively. Moreover, it fails to account for the findings that each germinal centre accommodates different founder B cells with a wide range of affinities, and that entry into the germinal centre is limited by competition and not absolute affinity7,10.
Here we describe RC1, an immunogen designed to recruit and expand diverse V3-glycan-specific B cells by improving accessibility of the V3-glycan patch epitope, which includes a group of high-mannose and complex-type N-glycans that surround V3 (Asn residues in gp120: N133, N137, N156, N295, N301, N332, N339, N385 and N392)15. bNAbs that target this site, including PGT12116, 10-107417 and BG1818, reach through these glycans using elongated heavy-chain complementarity-determining region (CDR) 3 (CDRH3) loops and portions of light-chain CDRs 1 and 3 (CDRL1 and CDRL3) to contact the highly conserved GDIR (G324-D325-I326-R327) motif at the base of V319. Here we show that RC1 activates and expands a diverse group of B cells expressing antibodies that resemble human V3-glycan patch bNAb precursors in mice, rabbits and rhesus macaques.
RC1 facilitates antibody binding to the V3-glycan patch
RC1 was designed using 11MUTB20, a modified native-like Env trimer (SOSIP.664) derived from the clade A/E BG505 Env21, as a template. Compared to BG505, 11MUTB includes substitutions in V1 and lacks potential N-linked glycosylation sites (PNGSs) at N133 and N13720 (Fig. 1a). We reasoned that removal of the N156 PNGS (N156Q) to create RC1 would facilitate recognition of the V3-glycan patch by increasing accessibility of V1 residues that interact with V3-glycan bNAbs22,23. Consistent with this idea, the absence of the N156 PNGS enhances neutralization by PGT121 and 10-1074, whereas the absence of other glycans, such as N301 or N137, reduces neutralization (Extended Data Fig. 1a). In addition, we hypothesized that removal of the N156 glycan, which includes negatively charged terminal sialic acids22,24, would produce a more electrostatically neutral Env surface that could facilitate the binding of the largely neutral precursor of PGT121 and 10-1074 (inferred germline of PGT121/10-1074)25.
a, N-glycans (coloured spheres) and GDIR motifs (red surfaces) mapped onto BG505 (Protein Data Bank (PDB) code 5T3Z) (N137 glycan from PDB 5FYL) in the top-down orientation. b, Left and right, side views of structures of BG505 and RC1 complexed with 10-1074 (glycan atoms are coloured spheres). Middle, superimposition of the boxed regions with protein in cartoon representations. Dark and light purple, 10-1074 VH and VL, respectively; red, GDIR; wheat, other portions of RC1; grey, BG505; orange spheres, N156 glycan. Regions of V1 showing displacement (gp120 residues 139–140) are indicated by dots and an arrow. c, Surface plasmon resonance (SPR) data for PGT121/10-1074 binding to Env trimers. NB, no binding above background; RU, response units. Representative plot from three independent experiments.
To characterize RC1, we compared its antigenic properties to BG505 (Extended Data Fig. 1b) and solved a 4.0-Å single-particle cryo-electron microscopy (cryo-EM) structure of RC1 complexed with 10-1074, comparing it to a BG505–10-1074 structure22 (Fig. 1b, Extended Data Fig. 2, Extended Data Table 1). Both structures showed three 10-1074 antigen-binding fragments (Fabs) bound to the V3-glycan patch epitopes of a closed Env trimer (Fig. 1b). Compared with BG505, the V1 loop in RC1 included more ordered residues and was shifted towards the 10-1074 CDRH3, allowing for increased interactions between RC1 and 10-1074 (Fig. 1b).
Despite structural changes in V1—which resulted from deletion of the N156 glycan (Fig. 1b)—the inferred germline of PGT121/10-107417 bound to RC1 and 11MUTB with similar affinities (dissociation constant (KD) values of approximately 50 μM; Fig. 1c), and priming immunizations with RC1 and 11MUTB elicited comparable V3-glycan-specific serological responses in knock-in mice that carried genes encoding the inferred germline of PGT121/10-10749 (Fig. 2a–c). Thus, RC1 exhibited structural changes resulting from N156 glycan deletion that did not affect its affinity for the inferred germline of PGT121/10-1074.
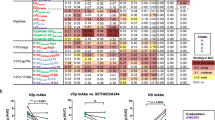
a, Immunization protocol. b, d, f, h, Representative ELISAs showing serum binding to indicated immunogens. Controls include naive serum (red), purified PGT121 (green) and inferred germline (iGL) of PGT121/10-1074 (black). OD405, optical density at 405 nm. b, The inferred germline of PGT121 knock-in (KI) mice9. d, f, h, Wild-type (WT) mice. c, e, Area under the curve (AUC) for ELISAs in b and d, respectively, but combined results from two experiments using three or four mice each. Each dot represents serum from one mouse. f, Binding to RC1 and RC1-glycanKO. g, Ratio of the AUC for RC1 and RC1-glycanKO ELISAs from seven experiments with two or three mice immunized with RC1. Red dot corresponds to mouse WT 4 in f. i, Ratio of the AUC for RC1 and RC1-glycanKO ELISAs for wild-type mice immunized with RC1 (seven experiments) or RC1-4fill (five experiments). j, Pie charts show clonal expansion of RC1-binding B cells in the germinal centre. Coloured slices are proportional to the number of clonal relatives. White indicates single IgVH sequences. The number of heavy chains analysed is indicated in the centre. k, IgH nucleotide (nt) mutations from naive and RC1 immunized mice in j. l, ELISA binding of representative monoclonal antibodies from RC1-immunized mice to RC1 and RC1-glycanKO. m, ELISA binding of Ab275MUR and Ab276MUR to indicated Env proteins. c, e, i, Unpaired t-tests. c, e, g, i, k, Data are mean and each dot is an individual mouse (c, e, g, i) or an individual sequence (k).
RC1 elicits V3-glycan antibodies in wild-type mice
To determine whether RC1 can activate B cells carrying antibodies that are specific to the V3-glycan patch in wild-type mice, we immunized C57BL/6J mice once with RC1 or 11MUTB (Fig. 2a). 11MUTB did not produce a measurable serological response, but RC1-immunized mice showed reproducible anti-V3-glycan-patch-specific serological responses, as shown by enzyme-linked immunosorbent assays (ELISAs) in which we compared binding to RC1 and a mutant variant of RC1 (RC1-glycanKO) that lacks two V3 PNGSs (N301 and N332) that are critical for human V3-glycan patch bNAbs (Fig. 2d–g, Extended Data Table 2). Moreover, serum from the RC1-immunized mice cross-reacted with 11MUTB but not with the more native-like Env 10MUT20 or with BG505 (Extended Data Fig. 3a). The improved immunogenicity of the V3-glycan patch epitope of RC1 results from specific removal of the N156 glycan from 11MUTB, because removal of a nearby glycan at N301 that is also part of the glycan patch (11MUTBΔ301) (Extended Data Table 2) failed to induce detectable serological responses (Fig. 2h). We conclude that, in contrast to 11MUTB and 11MUTBΔ301, RC1 elicits V3-glycan-specific serological responses in wild-type mice.
To reduce antibody responses to off-target epitopes26,27,28,29 and further focus responses on the V3-glycan patch, we produced an RC1 variant, RC1-4fill, by introducing PNGSs to add glycans to gp120 positions 230, 241, 289 and 344 (Extended Data Fig. 4). Compared with RC1, RC1-4fill elicited serological responses that were more focused on the V3-glycan patch in wild-type mice (Fig. 2i). We conclude that RC1-4fill focuses antibody responses to the V3-glycan patch.
Expansion of V3-glycan-specific B cells in mice
To further characterize the humoral responses elicited by RC1 and RC1-4fill, we sequenced antibody genes from single B cells from the germinal centre that bound to RC1 but not RC1-glycanKO (Extended Data Fig. 3b). All analysed RC1- and RC1-4fill-immunized mice showed expansion of B cell clones in the germinal centre (Fig. 2a, j). The expanded clones predominantly expressed heavy-chain V genes VH5-6, VH9-3 and VH2-9, and κ light-chain genes VK3-4 and VK14-111 (Fig. 2j, Extended Data Table 4 and Supplementary Tables 1, 2). The CDRH3 sequences in expanded clones showed similarities to human V3-glycan patch bNAbs, such as Tyr-rich or RXY motifs and longer-than-average CDRH3 sequences (Extended Data Table 4 and Supplementary Table 1). Consistent with a single immunization, the VH genes of the expanded clones had an average of 3.2 nucleotide mutations (Fig. 2k and Supplementary Table 1).
We mapped the target sites of these antibodies by ELISA against RC1 and RC1 mutant proteins. A diverse group of monoclonal antibodies showed V3-glycan-patch-specific binding (Fig. 2l). Further characterization of two mouse (MUR) antibodies, Ab275MUR and Ab276MUR, showed that they bound to the V3-glycan patch of RC1 (Ab275MUR, KD ≈ 30 nM) in a GDIR- and N301-glycan-dependent manner, and both retained binding to 11MUTB (Ab275MUR, KD ≈ 230 nM)—demonstrating accommodation of the N156 glycan—whereas neither antibody bound to BG505 or a peptide that covers the crown of the V3 loop (Fig. 2m, Extended Data Table 2, Extended Data Fig. 3c, d). Acquired mutations were essential for binding, because RC1 did not bind to the Ab276MUR-reverted inferred germline (Extended Data Fig. 3e). Consistent with a single immunization, neither Ab275MUR nor Ab276MUR showed detectable neutralizing activity against a panel of tier 1B and tier 2 HIV-1 isolates in assays using the TZM-bl reporter cell line. We conclude that RC1 and RC1-4fill expand mouse B cell clones expressing antibodies that target the V3-glycan patch.
V3-glycan antibody responses in rabbits and macaques
To enhance potential avidity effects and limit exposure of off-target epitopes at the Env base, we multimerized RC1-4fill on virus-like particles (VLPs) using the SpyTag–SpyCatcher system30,31 (Fig. 3a, b). VLPs were used to prime rabbits and rhesus macaques. Single immunizations of 4 rabbits and 16 macaques with VLP-RC1-4fill elicited serological responses that were partially specific for the V3-glycan patch in all animals (Fig. 3c–f, Extended Data Fig. 5a). Serum from macaques primed with VLP-RC1-4fill showed sequentially reduced binding to the more native-like immunogens 11MUTB and 10MUT20 (Extended Data Fig. 5b) and no neutralizing activity against a small panel of HIV-1 isolates that included fully glycosylated tier 2 and glycan-deleted viruses (Extended Data Table 3). Thus, VLP-RC1-4fill elicited robust serological responses that mapped in part to the V3-glycan patch in rabbits and rhesus macaques.
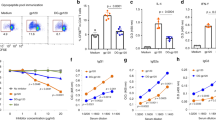
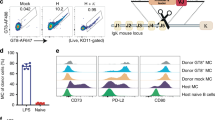
a, Model of VLP-RC1-4fill: RC1-4fill (wheat and pink), SpyTag (gold), SpyCatcher (cyan) and bacteriophage AP205 (green). b, Negative-stain electron microscopy images comparing VLPs (top) and VLP-RC1 (bottom). Arrows indicate the VLP surface (black) and RC1 (red). Scale bars, 50 nm. Representative image from three independent experiments. c, d, Immunization protocols for rabbits (c) and non-human primates (d). LN, lymph node. e, f, AUC for ELISAs with serum from four rabbits (e) and eight non-human primates (f) primed with VLP-RC1-4fill against RC1 (black) and RC1-glycanKO (grey). g, h, Flow cytometry plots showing frequency of B cells in the germinal centre that bind to RC1 but not to RC1-glycanKO. g, Representative flow cytometry plots. h, Quantification. n = 4 naive and n = 4 immunized non-human primates. i, Pie charts showing clonal expansion of RC1-binding B cells in the germinal centre (see legend in Fig. 2j). j, IgVH mutations for the sequences of clones shown in i (Supplementary Table 3). k, Logo plots comparing CDRL3 of inferred germline of PGT121/10-1074 and all IgL from B cells in the germinal centre shown in i. l, Fraction of CDRL3 sequences from i that show a DSS-like motif. h, l, Unpaired t-test. h, j, l, Data are mean and individual values.
To further characterize responses elicited by VLP-RC1-4fill in macaques, we purified germinal centre B cells from the draining lymph nodes that bound to RC1 but not to RC1-glycanKO by flow cytometry (RC1+RC1-glycanKO−). Whereas RC1+ cells were absent from germinal centres of naive macaques, RC1+RC1-glycanKO− germinal centre B cells were found at an average frequency of 0.4% of germinal centre B cells in the lymph nodes in the four macaques analysed (Fig. 3g, h).
Antibody cloning from four immunized macaques revealed expanded B cell clones that used a variety of VH genes, as found for human V3-glycan patch bNAbs32, with an average of 5.6 nucleotide somatic mutations (Fig. 3i, j and Supplementary Table 3). Most characterized human V3-glycan patch bNAbs contain a λ light chain18,33. Analysis of λ genes revealed that macaque RC1-binding cells preferentially used gene segments VL132 (91% nucleotide sequence identity to VL2-8 germline gene segments in PGT125–PGT128 and PGT130–PGT131) and VL124 (94% nucleotide sequence identity to the VL3-21 germline in PGT121–PGT123/10-1074) (Fig. 3k). In addition, 86% of the λ light chains had CDRL3 sequences that included an Asp-Ser-Ser (DSS) motif present in the inferred germlines of PGT121–123 and 10-1074/PGT12417 (Fig. 3l and Supplementary Table 4). This motif mutates to Asp-Ser-Arg (DSR) in the mature bNAbs, which is critical for the neutralization activity of PGT12134. Thus, there is congruence between the sequence of precursors of human V3-glycan patch bNAbs and the antibodies expressed by macaque B cell clones that are elicited by priming with VLP-RC1-4fill.
We expressed 38 macaque germinal centre antibodies with CDRL3 sequences that resembled the CDRL3 sequences of inferred germline V3-glycan patch bNAbs (Supplementary Table 5). The CDRL3 sequences of 33 out of 38 antibodies contained a DSS motif and a Gln at position 89 (QXXDSS motif), which was also found in the CDRL3 sequence of the inferred germlines of PGT121–PGT123, 10-1074, PGT124 and BG1817,18 (Extended Data Table 5). In addition, 5 out of 38 antibodies contained a Ser-Tyr-Ala-Gly (SYAG) motif, which is present in the CDRL3 sequences of the inferred germlines of PGT125–PGT128, PGT130 and PGT131 (Extended Data Table 5). In total, 30 out of 33 QXXDSS motif-containing antibodies and 2 out of 5 SYAG motif-containing antibodies bound to the V3-glycan patch epitope, as determined by ELISA using RC1 and RC1-glycanKO with additional mutations in the GDIR sequence (RC1-glycanKO–GAIA) (Fig. 4a and Supplementary Table 5). In addition, the CDRL1 of all 38 macaque antibodies contained an Asn-Ile-Gly (NIG)-like motif present in the PGT121/10-1074 antibody (33 with a NIG motif, 4 with a DIG (Asp-Ile-Gly) motif and 1 with a NLG (Asn-Leu-Gly) motif; Supplementary Table 5). The CDRH3 lengths of the 38 antibodies were relatively long (11–21 residues; average = 15.5 residues) (Fig. 4b). Longer CDRH3 sequences were enriched in Tyr and/or Phe residues, similar to the long CDRH3 sequences found in human V3-glycan patch bNAbs16,17,18 (Supplementary Table 5). The antibody VH and VL genes included an average of 4.9 and 3.3 nucleotide mutations, respectively (Fig. 4c). Consistent with recruitment of antibodies with a range of affinities to germinal centres and their subsequent affinity maturation7,10, the inferred germline versions of the macaque antibodies showed lower affinities for RC1 than their mutated counterparts, ranging from levels that were below quantification to KD values in the micromolar range (Extended Data Fig. 5c). Similarity between the macaque antibodies and PGT121/10-1074 was corroborated by ELISAs using an anti-idiotypic antibody specific for the PGT121/10-1074 inferred germline antibody (Extended Data Fig. 5d). The inferred germline-reverted versions of 5 out of 11 macaque antibodies were recognized by the anti-PGT121/10-1074 antibody (Extended Data Fig. 5d).
a, ELISA binding of representative macaque monoclonal antibodies to RC1 and RC1-glycanKO–GAIA. b, CDRH3 length of 32 V3-glycan-patch-specific monoclonal antibodies. AA, amino acids. c, Nucleotide mutations in IgVH and IgVL of 32 V3-glycan-patch-specific monoclonal antibodies. d, e, AUCs for ELISA binding of monoclonal antibodies to indicated proteins. Each AUC value corresponds to one ELISA curve. b, c, Data are mean and each dot is an individual sequence.
To further characterize the target site of the macaque antibodies, we performed ELISAs against additional proteins: RC1-glycanKO, RC1–GAIA, RC1-glycanKO–GAIA, 11MUTBΔ301, RC1Δ301, RC1Δ332, 11MUTB20 and BG505 (Fig. 4d, e, Extended Data Table 2). The ELISAs suggested four distinct RC1-binding patterns among antibodies that contained a CDRL3 QXXDSS motif (Fig. 4d) and another pattern among antibodies that contained a SYAG motif (Fig. 4e). Whereas all antibodies were glycan dependent as determined by no binding to RC1-glycanKO, they differed in binding to 11MUTB or 10MUT and dependence on GDIR and N301, N332 and N156 glycans (Fig. 4a, d, e). Although none of the antibodies recognized BG505, Ab933NHP (where NHP denotes antibodies elicited in non-human primates), Ab936NHP and Ab1170NHP bound to 11MUTB, indicating that they can accommodate the N156 glycan (Fig. 4d, e). Consistent with no binding to BG505, none of the antibodies exhibited neutralizing activity, and removal of the N133, N137 and N156 glycans did not render the BG505/T332N and JRCSF.JB viruses sensitive to neutralization (Extended Data Table 3b, c), suggesting that the lack of neutralization is not due to clashes with those glycans. We conclude that macaque immunization with VLP-RC1-4fill elicits V3-glycan-patch-specific antibodies that resemble precursors of human bNAbs that target this site.
Cryo-EM structures of antibodies bound to RC1
We determined structures of one mouse and two macaque Fabs complexed with RC1 using single-particle cryo-EM. Ab275MUR (4.4-Å resolution) and Ab874NHP (3.9 Å) (derived from the same clone as Ab876NHP) bound RC1 similarly to each other, consistent with their 69% VH domain amino acid sequence identity, whereas Ab897NHP (4.4 Å) (related by 48% and 54% VH sequence identity to Ab275MUR and Ab874NHP, respectively) adopted a distinct angle of approach (Fig. 5a, Extended Data Fig. 5e).
a, Top, VH–VL domains of 10-1074 and elicited antibodies bound to one protomer of RC1 (GDIR residues are red; glycans are coloured spheres). Bottom, antibody combining sites (CDRs shown as loops) mapped onto gp120 (glycans as coloured spheres; GDIR in red). b, Comparisons of interactions of the GDIR motif with 10-1074 and elicited antibodies (colours as in a).
All three Fabs in the RC1 complexes bound to the V3-glycan patch epitope and contacted the GDIR motif, but with different orientations and footprints from each other and from V3-glycan patch bNAbs (Fig. 5a, Extended Data Fig. 5e). 10-1074 contacts the conserved GDIR motif using CDRH3, CDRL1 and CDRL322 (Figs. 1c, 5a); Ab874NHP and Ab275MUR made GDIR contacts using CDRH2; whereas Ab897NHP used CDRL1 and CDRL3 (Fig. 5a, b). In addition, Ab874NHP and Ab897NHP contain the conserved CDRL3 QXXDSS motif, which makes contact with conserved regions of the V3-glycan epitope in mature bNAbs22,35. Similar to mature V3-glycan patch bNAbs, Ab897NHP contains a substitution (Ser93Asn) within the QXXDSS motif that enables contacts with gp120GDIR, and also uses its conserved CDRL1 NIG motif to contact the V1 loop (Extended Data Fig. 5f).
Ab275MUR and Ab874NHP also interacted with the N332 glycan, consistent with mature V3-glycan bNAbs (Fig. 5a, Extended Data Fig. 5e). However, in contrast to 10-1074—which interacts with the N332 glycan via its CDRL1, light-chain framework region 3 (FRWL3), CDRH2 and CDRH322—Ab275MUR made contacts using CDRH2, whereas Ab874NHP engaged the N332 glycan with CDRH2 and heavy-chain framework region 3 (FRWH3). We did not observe N332 glycan interactions in the Ab897NHP–RC1 structure. Despite reduced binding of Ab275MUR, Ab876NHP (same clone as Ab874NHP) and Ab897NHP to RC1Δ301 (Fig. 2l), none of these Fabs showed interactions with the N301 glycan in our structures, suggesting that glycan heterogeneity obscures this interaction, and/or conformational heterogeneity in a V3-glycan patch that lacks this glycan diminishes binding36. We conclude that RC1 elicits V3-glycan-patch-targeting antibodies with distinct binding modes in animals with polyclonal antibody repertoires.
Conclusions
HIV-1 bNAbs develop in infected humans by sequential rounds of somatic mutation in response to a rapidly evolving pathogen4. Vaccination with a series of related antigens can reproduce this progression of events in genetically engineered mice that carry supraphysiological numbers of B lymphocytes that express the inferred germline precursors of bNAbs9. An important goal of HIV-1 vaccine design is to develop immunogens that initiate this response in organisms with polyclonal immune systems and then reproduce these responses in humans.
HIV-1 immunogen design has focused on increasing the affinity of candidate immunogens for specific inferred germline precursors of bNAbs, with the objective of recruiting a specific group of rare precursors into the germinal centre1. This approach typically fails to account for increases in apparent affinity produced by interactions between multimerized antigen and clusters of bivalent antigen receptors on the surface of a B cell. Moreover, entry into the germinal centre is primarily limited by competition7,10,11,14. Thus, the importance of affinity is relative, as shown by the observation that B cells bearing low-affinity receptors are frequently found in germinal centres under physiological conditions10,37, and by our finding that inferred germline precursors of macaque antibodies elicited by RC1 showed relatively low affinity for the immunogen.
The principles we used to produce RC1 did not take affinity for a germline B cell receptor into account. Instead, RC1 was designed to increase the number of bNAb progenitors that compete for entry into germinal centres by making the antigenic target site more available and facilitating binding to electrostatically neutral inferred germline precursors25. In addition, VLP-RC1-4fill incorporates the idea that masking competing off-target epitopes26,29 by addition of glycans27 and tethering the bottom of the trimer to a VLP minimizes competition for entry into the germinal centre.
RC1 differs from other HIV-1 vaccine candidates in that it induces B cells that express antibodies against a targeted epitope to undergo clonal expansion in germinal centres in animals with a fully polyclonal B cell repertoire. In macaques, these B cells express antibodies that show sequence and structural similarities to inferred germline precursors of bNAbs that target the V3-glycan patch. Like the precursors of human bNAbs, they do not bind to wild-type Env or neutralize HIV-15. Notably, biochemical and structural results showed that antibodies with distinct mechanisms of targeting the V3-glycan patch were elicited by RC1, increasing the probability that one or more might develop breadth and potency after boosting9. Thus, VLP-RC1-4fill is a suitable candidate immunogen for further evaluation in sequential vaccination strategies to elicit bNAbs.
Methods
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Envelope proteins
Env trimers were expressed as soluble native-like gp140 trimers21. The newly engineered Env SOSIP trimers, RC1, RC1-4fill, RC1–Avitag, RC1–SpyTag, RC1-glycanKO, RC1-glycanKO–Avitag, RC1-glycanKO–GAIA and RC1–GAIA, BG50521 and the BG505 variants 11MUTB, 10MUT, 7MUT and 5MUT20 were cloned in the pPPPI4 expression vector using synthetic gene fragments (Integrated DNA Technologies (IDT)). The glycan variants RC1Δ301, RC1Δ332 and 11MUTBΔ301 were produced by site-directed mutagenesis (QuikChange Lightning Multi-site directed mutagenesis kit, Agilent Technologies). Specific modifications of each protein are listed in Extended Data Table 2.
Soluble Env trimers were expressed by transient transfection in HEK293-6E cells (National Research Council of Canada) or Expi293 cells (Life Technologies) and purified from cell supernatants by 2G12 or NIH45-46 immunoaffinity chromatography and size-exclusion chromatography (SEC) as previously described38. Proteins were stored at 4 °C in 20 mM Tris pH 8.0 and 150 mM sodium chloride (TBS buffer). SpyTagged immunogens were buffer-exchanged into 20 mM sodium phosphate pH 7.5, 150 mM NaCl. Cell lines were not tested for mycoplasma contamination.
VLP production and conjugation
A C-terminal SpyTag sequence (13 residues) was added to RC1-4fill to form an irreversible isopeptide bond to SpyCatcher protein31. We produced and purified SpyCatcher–AP20539 VLPs as described30 and separated conjugated VLPs from free Env trimers by SEC on a Superdex 200 column. Conjugation of Env trimers was verified by negative-stain electron microscopy and/or SDS–PAGE (Fig. 3, Extended Data Fig. 4), and immunogen concentrations were estimated by comparing to known amounts of free immunogen run on the same SDS–PAGE gel. Conjugated and unconjugated VLPs were compared by negative-stain electron microscopy on a FEI Tecnai T12 transmission electron microscope at 120 keV using a Gatan Ultrascan 2k × 2k CCD detector.
Mass spectrometry
The glycosylation profiles of RC1 and RC1-4fill were determined as previously described40. In brief, samples were denatured with Lys-C (Promega), Arg-C (Promega), Glu-C (Promega) and chymotrypsin (Promega). Following digestion, the samples were deglycosylated by Endo H (Promega) and PNGase F (Glyko, Prozyme) in the presence of 18O water (Cambridge Isotope Laboratories). The resulting peptides were separated on an Acclaim PepMap RSLC C18 column (75 μm × 15 cm) and analysed using an Orbitrap Fusion Lumos Tribrid mass spectrometer (Thermo Fisher Scientific) with a 240-min linear gradient consisting of 1–100% solvent B over 180 min at a flow rate of 200 nl min−1. Full mass spectrometry scans were acquired using the Fusion instrument software (v.2.0, Thermo Fisher Scientific), and the resulting spectra were analysed and filtered using SEQUEST (Proteome Discoverer 1.4, Thermo Fisher Scientific) and ProteoIQ (v.2.7, Premier Biosoft). Site occupancy was calculated using spectral counts assigned to the 18O-Asp-containing (PNGase-F-cleaved) and/or HexNAc-modified (Endo-H-cleaved) peptides and their unmodified counterparts.
Animals
Mice carrying the inferred germline IgH and IgL human PGT121 and 10-1074 bNAbs (GLHL121 knock-in mice) were previously described9. Six-to-eight-week-old C57BL/6J male mice from The Jackson Laboratory were used for immunizations. All animal procedures were performed in accordance to protocols approved by the Rockefeller University Institutional Animal Care and Use Committee (IACUC). Male and female GLHL121 knock-in mice or C57BL/6J wild-type mice were equally distributed in groups and immunized intraperitoneally with 10 μg of soluble SOSIP Env trimer in Ribi adjuvant (Sigma-Aldrich) (1:1).
Six-month-old New Zealand white rabbits (Covance) were used for immunizations. Rabbits were immunized subcutaneously with approximately 22 μg of RC1-4fill SOSIP Env trimer conjugated to VLP (VLP-RC1-4fill) in an ISCOMs-like saponin adjuvant (see below). Serum samples were collected from mice and rabbits on weeks 0 and 2 after immunization. All procedures in rabbits were approved by the Denver PA IACUC.
Sixteen rhesus macaques (Macaca mulatta) of Indian genetic origin, two-to-four years of age, were housed and cared for in accordance with Guide for Care and Use of Laboratory Animals Report number NIH 82-53 (Department of Health and Human Services, Bethesda, 1985) in a biosafety level 2 NIH facility. All animal procedures and experiments were performed according to protocols approved by the IACUC of NIAID, NIH.
Macaques were immunized subcutaneously in the medial inner forelegs and hind legs (total of four sites per animal) with approximately 200 μg (experiment 1; Fig. 3f) or 100 μg (experiment 2; Extended Data Fig. 5a) of RC1-4fill SOSIP trimer conjugated to VLP (RC1-4fill VLP) adjuvated in IscoMPLA. Blood and lymph node biopsies were obtained from naive macaques and from the immunized macaques three weeks after immunization.
Adjuvant synthesis
ISCOM-like saponin adjuvant was prepared as described41. Final adjuvant concentrations were determined by cholesterol quantification (Sigma-Aldrich, MAK043).
ELISA
ELISAs with SOSIP Env trimers 11MUTB, RC1, 11MUTBΔ301, RC1Δ301, RC1–GAIA, RC1-glycanKO, RC1-glycanKO–GAIA, RC1Δ332, BG505, 10MUT, 7MUT, 5MUT or the V3 loop-consensus C peptide (KGKGKGKGKGCTRPNNNTRKSIRIGPGQTFYATGDIIGDIRQAHC) were performed as described9. Serum samples were assayed at a 1:100 or 1:30 starting dilution and seven additional threefold serial dilutions. Mouse and human IgGs or human Fabs were evaluated at concentrations specified in the main text.
Alternatively, 96-well plates were directly coated with 50 μl of a solution of Fab at 20 μg ml−1 in 1× PBS overnight at 4 °C, washed and blocked as above and incubated in 50 μl of a solution of RC1 or RC1-glycanKO–GAIA at 2 μg ml−1 in blocking buffer for 1 h at room temperature. Plates were washed as above and developed using a chimeric version (human Fabs and mouse crystallizable fragment (Fc)) of the CD4-binding site bNAb 3BNC6042 at threefold serial dilutions starting at 5 μg ml−1 followed by anti-mouse IgG secondary antibody conjugated to HRP (Jackson ImmunoResearch, 115-035-071).
For anti-idiotype ELISAs, 96-well plates were coated with 50 μl of a solution of IgG at 10 μg ml−1 in 1× PBS overnight at 4 °C, washed and blocked as above and incubated with biotinylated anti-inferred germline PGT121 idiotypic antibody. Plates were developed with streptavidin conjugated to HRP.
Flow cytometry and single B cell sorting
Single-cell suspensions were obtained from the draining lymph nodes and spleens of immunized mice, and mature B cells were isolated by negative selection using anti-CD43 magnetic beads (MACS) following the manufacturer’s instructions.
Frozen peripheral blood mononuclear cells or cells from lymph node biopsies obtained from the naive and immunized macaques were thawed and washed in RPMI 1640 medium (1×) (Gibco, 11875-093). Mouse or macaque cells were incubated with 100 μl of FACS buffer (1× PBS with 2% fetal bovine serum and 1 mM EDTA) with mouse (BD Biosciences, 553142) or human (BD Biosciences, 564219) Fc Block, respectively, at a 1:500 dilution for 30 min on ice.
RC1 and RC1-glycanKO (RC1+RC1-glycanKO−) tetramers were prepared by incubating 5 μg of Avitagged and biotinylated RC1 (RC1–AviBio) or Avitagged and biotinylated RC1-glycanKO (RC1-glycanKO–AviBio) with fluorophore-conjugated streptavidin at a 1:200 dilution in 1× PBS for 30 min on ice.
RC1+RC1-glycanKO− mouse B cells were isolated using RC1–AviBio conjugated to streptavidin BV711 (BD Biosciences, 563262) and RC1-glycanKO–AviBio conjugated to streptavidin–PE (BD Biosciences, 554061) as baits. RC1+RC1-glycanKO− macaque B cells were isolated using RC1–AviBio conjugated with streptavidin–PE and streptavidin–AF647 and RC1-glycanKO–AviBio conjugated with streptavidin–BV605 (BD Biosciences, 563260). Tetramers were mixed with the human or mouse antibody cocktails indicated below to a final concentration of 5 μg ml−1 each.
Mouse cells were stained with anti-CD4–APC-eFluor780 (Invitrogen, 47-0042-82), anti-CD8–APC-eFluor780 (Invitrogen, 47-0081-82), anti-F4/80–APC-eFluor780 (Invitrogen, 47-4801-82), anti-NK1.1–APC-eFluor780 (Invitrogen, 47-5941-82), anti-CD11b–APC-eFluor780 (eBioscience, 47-0112-82), anti-CD11c–APC-eFluor780 (eBioscience, 47-0114-82), anti-Gr-1–APC-eFluor780 (Invitrogen, 47-5931-82), anti-B220–APC (Biolegend, 103212), anti-GL7–FITC (BD Biosciences, 553666) and anti-CD95–BV421 (BD Biosciences, 562633) antibodies at 1:200 dilution and the live/dead marker Zombie NIR (Biolegend, 77184) at a 1:400 dilution in FACS buffer. Macaque cells were stained with anti-CD16–APC-eFluor780 (Invitrogen, 47-0168-41), anti-CD8a–APC-eFluor780 (Invitrogen, 47-0086-42), anti-CD3–APC-eFluor780 (Invitrogen, 47-0037-41), anti-CD14–APC-eFluor780 (eBiosciences, 47-0149-41), anti-CD20–PE-Cy7 (BD, 335793), anti-CD38–FITC (Stem Cell Technologies, 60131FI), anti-IgG–BV421 (BD Biosciences, 562581) and anti-IgM–PerCP-Cy5.5 (BD Biosciences, 561285) antibodies at a 1:200 dilution and the live/dead marker Zombie NIR at a 1:400 dilution in FACS buffer.
Zombie NIR−CD4−CD8−F4/80−NK1.1−CD11b−CD11c−B220+GL7+CD95+RC1+RC1-glycanKO− single cells were isolated from the mouse cell homogenates and Zombie NIR−CD16−CD8a−CD3−CD14−CD20+CD38+IgG+/−double RC1+RC1-glycanKO− single cells were isolated from the macaque cell homogenates using a FACS Aria III (Becton Dickinson).
Single cells were sorted into individual wells of a 96-well plate containing 5 μl of lysis buffer (TCL buffer (Qiagen, 1031576) with 1% of 2-β-mercaptoethanol). Plates were immediately frozen on dry ice and stored at −80 °C.
Antibody sequencing and cloning
Single-cell RNA was purified using magnetic beads (RNAClean XP, A63987, Beckman Coulter). RNA was eluted from the magnetic beads with 11 μl of a solution containing 14.5 ng μl−1 of random primers (Invitrogen, 48190-011), 0.5% of tergitol (type NP-40, 70% in H2O, Sigma-Aldrich, NP40S-100ML) and 0.6 U μl−1 of RNase inhibitor (Promega, N2615) in nuclease-free water (Qiagen), and incubated at 65 °C for 3 min. cDNA was synthesized by reverse transcription (SuperScript III Reverse Transcriptase, Invitrogen, 18080-044, 10,000 U)43. cDNA was stored at −80 °C or used for antibody gene amplification by nested polymerase chain reaction (PCR) after addition of 10 μl of nuclease-free water.
Mouse and macaque antibody genes were cloned43 using the primers in Supplementary Table 6. PCR protocols were as follows for annealing (°C)/elongation (s)/number of cycles: first PCR (IgG IgH and Igλ): 46/55/50; second PCR (IgG IgH and Igλ): 50/55/50.
Inferred germline macaque IgGs and Fabs were produced by reverting all nucleotide mutations in the V(D)J antibody genes to their corresponding inferred germline sequences while conserving the N nucleotides from the V(D)J junctions found in the mutated antibodies.
Antibody production and purification
Immunoglobulins were purified from 200 μl of mouse or macaque serum using Ab Spin Trap Protein G Sepharose columns (GE Healthcare, 28-4083-47). Immunoglobulin-containing fractions were buffer-exchanged with PBS by overnight dialysis at 4 °C (dialysis cassettes 20000 MWCO, Thermo Fisher Scientific, 66005).
For structural studies, mouse IgGs and macaque His6-tagged Fabs were expressed by transient transfection in HEK293-6E or Expi293 cells and purified from cell supernatants using protein A or G (GE Healthcare) (for IgGs) or Ni-NTA (GE Healthcare) or Ni Sepharose 6 Fast Flow (GE Healthcare) (for Fabs) chromatography and SEC44. Mouse Fab was obtained by digesting IgG at 1−5 mg ml−1 with ficin (Sigma-Aldrich). Fab was purified by protein G (GE Healthcare) and SEC chromatography45, followed by monoQ 5/50 (GE Healthcare) ion-exchange chromatography. The common inferred germline of the PGT121 and 10-1074 bNAbs17 was expressed as a His6-tagged Fab.
In vitro neutralization assay
TZM-bl assays were performed as previously described46. In brief, neutralization activity was calculated as a function of the reduction in Tat-induced luciferase expression in the TZM-bl reporter cell line after a single round of virus infection with Env pseudoviruses.
SPR and OCTET binding studies
SPR experiments were performed using a T200 (Biacore). For measuring the affinity for PGT121/10-1074 inferred germline Fab, protein A was immobilized on a CM5 chip by primary amine chemistry (Biacore manual) and 200 nM 8ANC195G52K5 IgG or a non-HIV Env-binding IgG (mG053) was injected as previously described44. Then, 1 μM human Fc was injected to block remaining protein A sites. After capturing 10 μM RC1, 11MUTB or 10MUT, a concentration series of PGT121/10-1074 inferred germline Fab (fourfold dilutions from a top concentration of 160 μM for 10MUT, and twofold dilutions from a top concentration of 150 μM for 11MUTB and RC1) was injected, and binding reactions were allowed to reach equilibrium. KD values were derived by nonlinear regression analysis of plots of Req (equilibrium binding response) versus the log of the injected protein concentration and the data were fitted to a 1:1 binding model47. To measure the affinity of Ab275MUR, a concentration series of Fab was injected over immobilized RC1 or 11MUTB (fourfold dilutions from a top concentration of 50 μM). KD values were calculated from the on/off rates (ka/kd), which were derived using a 1:1 binding model from seven concentrations of Ab275MUR Fab (3.125 μM to 0.763 nM). Flow cells were regenerated as previously described44 with 1 M guanidine HCl and/or 10 mM glycine pH 2.0 at a flow rate of 90 μl min−1.
OCTET experiments were performed using the OCTET Red96 system to determine affinities of inferred germline and mutated macaque Fabs for RC1. Biotinylated RC1–Avitag was immobilized on high-precision streptavidin (SAX) biosensors (FORTÉBIO) using a solution of biotinylated RC1–Avitag at 400 nM in dilution buffer (FORTÉBIO). Four serial dilutions of each macaque Fab, one irrelevant Fab and 3BNC60 Fab were prepared in dilution buffer (FORTÉBIO). The binding experiment was performed at 30 °C using the following protocol: baseline 1 (60 s), load RC1 (300 s), baseline 2 (200 s), Fab association (300 s) and Fab dissociation (600 s). Analysis was performed using OCTET software Data Analysis HT 10.0 (FORTÉBIO).
Cryo-EM sample and grid preparation
RC1 complexed with 10-1074 was prepared by incubating purified RC1 with 10-1074 Fab and a CD4-binding site (CD4bs) Fab at a 1:3:3 molar ratio (gp140 protomer:10-1074 Fab:CD4bs Fab) overnight at room temperature. The RC1–Fab complex was isolated by SEC in TBS (20 mM Tris pH 8.0, 100 mM NaCl) using a Superdex-200 Increase 10/300 column (GE Healthcare). RC1–mouse/macaque Fab complexes were prepared by incubating purified RC1 with a mouse or macaque Fab and with 8ANC195 Fab42 at a 1:1.3:1.3 molar ratio as above and used without SEC purification. RC1–Fab complexes were diluted to 0.75–1.4 mg ml−1 in TBS, and 3 μl was added to Quantifoil R1.2/1.3 300 mesh copper grids (Electron Microscopy Services) that had been freshly glow-discharged using a PELCO easiGlow (Ted Pella). Samples were vitrified in 100% liquid ethane using a Mark IV Vitrobot (Thermo Fisher Scientific). Sample preparation conditions are summarized in Extended Data Table 1.
Cryo-EM data collection
RC1–Fab complexes were collected on a 200 kV Thermo Fisher Talos Arctica electron microscope using EPU automated image acquisition software48. Movies were collected on a Falcon 3EC direct electron detector (Thermo Fisher Scientific) operating in counting mode at a nominal magnification of 73,000× (1.436 Å per pixel) using a defocus range of −1.4 μm to −3.0 μm or −0.8 μm to −2.5 μm. Data for the RC1–10-1074 complex were collected across two separate sessions and combined during data processing. Microscope conditions are summarized in Extended Data Table 1.
Cryo-EM data processing
Movies were motion-corrected and dose-weighted using the MotionCor249 frame alignment program in RELION-350. Dose-weighted summed images were used for CTF determination using Gctf51, and reference-free particle picking from each micrograph was achieved using Laplacian-of-Gaussian filtering in RELION-350. Unbinned extracted particles were imported into cryoSPARC v.252 and subjected to reference-free two-dimensional classification using a 240-Å circular mask. Particles from the best two-dimensional classes were selected for heterogeneous ab initio model generation (two models). The best model exhibited C3 symmetry and was used as an initial model for homogenous three-dimensional auto-refinement in cryoSPARC v.252. Resolutions were estimated using the gold standard Fourier shell correlation (FSC = 0.143)53, and maps were auto-sharpened in cryoSPARC52. For interpreting N-linked glycans, maps were generated with overall B-factors ranging from −150 to −400 Å2 to improve local features and map connectivity54. See Extended Data Fig. 2 and Extended Data Table 1.
Model building
Initial coordinates were generated by docking reference models into the maps using UCSF Chimera55. For the RC1–10-1074 complex, BG505 Env and 10-1074 Fab (PDB code 53TZ) were docked into the density maps. For RC1 complexes with mouse or macaque Fabs, BG505 Env and PGT121/10-1074 inferred germline (PDB codes 5CEZ and 4FQQ) coordinates were docked into density maps. Initial models were refined into electron microscopy maps using rigid body refinement55. Models were built using Fab and RC1 sequences following iterative rounds of real-space refinement in Coot and PHENIX56,57. Coordinates for glycans were added as Man9 and then trimmed to fit the maps at σ = 5. Model validation was done using MolProbity58 and Privateer59. Superposition calculations and molecular representations were generated with PyMOL (v.1.5.0.4 Schrodinger), UCSF Chimera55 and ResMap60.
Analysis
MacVector v.15.5.3 was used for sequence analysis and graphs were created using R language. Flow cytometry data were processed using FlowJo v.10.5.0. GraphPad Prism 7 was used for data analysis. Immunoglobulin gene sequence AB1 files were converted to FASTQ format using the Biopython package. FASTQ files were trimmed by quality using cutadapt v.1.18 software. Igblast v.1.9.0 was used for VDJ assignment and clone analysis was performed using Change-O software v.0.3.7. For macaques, a custom VDJ database was created using previously reported immunoglobulin gene sequences61.
Quantification and statistical analysis
Statistical information, including n, mean and statistical significance values, is indicated in the text or the figure legends. GraphPad Prism 7 was used for statistical analysis by unpaired Student’s t-test. Data were considered statistically significant at *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The atomic models and cryo-EM density maps generated during the current study have been deposited in the Protein Data Bank and Electron Microscopy Data Bank with accession numbers 6ORN and EMD-20175 (RC1–10-1074), 6ORQ and EMD-20178 (RC1–Ab275MUR), 6ORO and EMD-20176 (RC1–Ab874NHP), and 6ORP and EMD-20177 (RC1–Ab897NHP). Sequence datasets generated and analysed during the current study are available from the corresponding authors upon reasonable request.
References
McCoy, L. E. & Burton, D. R. Identification and specificity of broadly neutralizing antibodies against HIV. Immunol. Rev. 275, 11–20 (2017).
West, A. P. Jr et al. Structural insights on the role of antibodies in HIV-1 vaccine and therapy. Cell 156, 633–648 (2014).
Kwong, P. D. & Mascola, J. R. HIV-1 vaccines based on antibody identification, B cell ontogeny, and epitope structure. Immunity 48, 855–871 (2018).
Bonsignori, M. et al. Antibody-virus co-evolution in HIV infection: paths for HIV vaccine development. Immunol. Rev. 275, 145–160 (2017).
Klein, F. et al. Somatic mutations of the immunoglobulin framework are generally required for broad and potent HIV-1 neutralization. Cell 153, 126–138 (2013).
Victora, G. D. & Nussenzweig, M. C. Germinal centers. Annu. Rev. Immunol. 30, 429–457 (2012).
Schwickert, T. A. et al. A dynamic T cell-limited checkpoint regulates affinity-dependent B cell entry into the germinal center. J. Exp. Med. 208, 1243–1252 (2011).
Escolano, A., Dosenovic, P. & Nussenzweig, M. C. Progress toward active or passive HIV-1 vaccination. J. Exp. Med. 214, 3–16 (2017).
Escolano, A. et al. Sequential immunization elicits broadly neutralizing anti-HIV-1 Antibodies in Ig knockin mice. Cell 166, 1445–1458 (2016).
Tas, J. M. et al. Visualizing antibody affinity maturation in germinal centers. Science 351, 1048–1054 (2016).
Dal Porto, J. M., Haberman, A. M., Shlomchik, M. J. & Kelsoe, G. Antigen drives very low affinity B cells to become plasmacytes and enter germinal centers. J. Immunol. 161, 5373–5381 (1998).
Abbott, R. K. et al. Precursor frequency and affinity determine B cell competitive fitness in germinal centers, tested with germline-targeting HIV vaccine immunogens. Immunity 48, 133–146 (2018).
Dosenovic, P. et al. Anti-HIV-1 B cell responses are dependent on B cell precursor frequency and antigen-binding affinity. Proc. Natl Acad. Sci. USA 115, 4743–4748 (2018).
Shih, T.-A. Y., Meffre, E., Roederer, M. & Nussenzweig, M. C. Role of BCR affinity in T cell dependent antibody responses in vivo. Nat. Immunol. 3, 570–575 (2002).
Kong, L. et al. Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nat. Struct. Mol. Biol. 20, 796–803 (2013).
Walker, L. M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011).
Mouquet, H. et al. Complex-type N-glycan recognition by potent broadly neutralizing HIV antibodies. Proc. Natl Acad. Sci. USA 109, E3268–E3277 (2012).
Freund, N. T. et al. Coexistence of potent HIV-1 broadly neutralizing antibodies and antibody-sensitive viruses in a viremic controller. Sci. Transl. Med. 9, eaal2144 (2017).
Sok, D. et al. A prominent site of antibody vulnerability on HIV envelope incorporates a motif associated with CCR5 binding and its camouflaging glycans. Immunity 45, 31–45 (2016).
Steichen, J. M. et al. HIV vaccine design to target germline precursors of glycan-dependent broadly neutralizing antibodies. Immunity 45, 483–496 (2016).
Sanders, R. W. et al. A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013).
Gristick, H. B. et al. Natively glycosylated HIV-1 Env structure reveals new mode for antibody recognition of the CD4-binding site. Nat. Struct. Mol. Biol. 23, 906–915 (2016).
Garces, F. et al. Structural evolution of glycan recognition by a family of potent HIV antibodies. Cell 159, 69–79 (2014).
Andrabi, R. et al. Glycans function as anchors for antibodies and help drive HIV broadly neutralizing antibody development. Immunity 47, 524–537 (2017).
Scharf, L. et al. Structural basis for germline antibody recognition of HIV-1 immunogens. eLife 5, e13783 (2016).
McCoy, L. E. et al. Holes in the glycan shield of the native HIV envelope are a target of trimer-elicited neutralizing antibodies. Cell Rep. 16, 2327–2338 (2016).
Duan, H. et al. Glycan masking focuses immune responses to the HIV-1 CD4-binding site and enhances elicitation of VRC01-class precursor antibodies. Immunity 49, 301–311 (2018).
Garrity, R. R. et al. Refocusing neutralizing antibody response by targeted dampening of an immunodominant epitope. J. Immunol. 159, 279–289 (1997).
Klasse, P. J. et al. Epitopes for neutralizing antibodies induced by HIV-1 envelope glycoprotein BG505 SOSIP trimers in rabbits and macaques. PLoS Pathog. 14, e1006913 (2018).
Brune, K. D. et al. Plug-and-Display: decoration of virus-like particles via isopeptide bonds for modular immunization. Sci. Rep. 6, 19234 (2016).
Zakeri, B. et al. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc. Natl Acad. Sci. USA 109, E690–E697 (2012).
Longo, N. S. et al. Multiple antibody lineages in one donor target the glycan-V3 supersite of the HIV-1 envelope glycoprotein and display a preference for quaternary binding. J. Virol. 90, 10574–10586 (2016).
Burton, D. R. & Hangartner, L. Broadly neutralizing antibodies to HIV and their role in vaccine design. Annu. Rev. Immunol. 34, 635–659 (2016).
Sok, D. et al. The effects of somatic hypermutation on neutralization and binding in the PGT121 family of broadly neutralizing HIV antibodies. PLoS Pathog. 9, e1003754 (2013).
Lee, J. H., de Val, N., Lyumkis, D. & Ward, A. B. Model building and refinement of a natively glycosylated HIV-1 Env protein by high-resolution cryoelectron microscopy. Structure 23, 1943–1951 (2015).
Zolla-Pazner, S. et al. Structure/function studies involving the V3 region of the HIV-1 envelope delineate multiple factors that affect neutralization sensitivity. J. Virol. 90, 636–649 (2015).
Murugan, R. et al. Clonal selection drives protective memory B cell responses in controlled human malaria infection. Sci. Immunol. 3, eaap8029 (2018).
Wang, H. et al. Asymmetric recognition of HIV-1 envelope trimer by V1V2 loop-targeting antibodies. eLife 6, e27389 (2017).
Tissot, A. C. et al. Versatile virus-like particle carrier for epitope based vaccines. PLoS ONE 5, e9809 (2010).
Duan, H. et al. Glycan masking focuses immune responses to the HIV-1 CD4-binding site and enhances elicitation of VRC01-class precursor antibodies. Immunity 49, 301–311 (2018).
Lövgren-Bengtsson, K. & Morein, B. in Vaccine Adjuvants: Preparation Methods and Research Protocols (ed. O’Hagan, D.) 239–258 (Humana, 2000).
Scheid, J. F. et al. Sequence and structural convergence of broad and potent HIV antibodies that mimic CD4 binding. Science 333, 1633–1637 (2011).
von Boehmer, L. et al. Sequencing and cloning of antigen-specific antibodies from mouse memory B cells. Nat. Protocols 11, 1908–1923 (2016).
Scharf, L. et al. Broadly neutralizing antibody 8ANC195 recognizes closed and open states of HIV-1 Env. Cell 162, 1379–1390 (2015).
Diskin, R., Marcovecchio, P. M. & Bjorkman, P. J. Structure of a clade C HIV-1 gp120 bound to CD4 and CD4-induced antibody reveals anti-CD4 polyreactivity. Nat. Struct. Mol. Biol. 17, 608–613 (2010).
Montefiori, D. C. Measuring HIV neutralization in a luciferase reporter gene assay. Methods Mol. Biol. 485, 395–405 (2009).
Vaughn, D. E. & Bjorkman, P. J. High-affinity binding of the neonatal Fc receptor to its IgG ligand requires receptor immobilization. Biochemistry 36, 9374–9380 (1997).
Tan, Y. Z., Cheng, A., Potter, C. S. & Carragher, B. Automated data collection in single particle electron microscopy. Microscopy 65, 43–56 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Terwilliger, T. C., Sobolev, O. V., Afonine, P. V. & Adams, P. D. Automated map sharpening by maximization of detail and connectivity. Acta Crystallogr. 74, 545–559 (2018).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Corcoran, M. M. et al. Production of individualized V gene databases reveals high levels of immunoglobulin genetic diversity. Nat. Commun. 7, 13642 (2016).
Acknowledgements
We thank members of the Bjorkman, Martin and Nussenzweig laboratories for discussions, T. Eisenreich and S. Tittley for animal husbandry, K. Gordon for flow cytometry, S. Zolla-Pazner for providing the V3-consensus C peptide and anti-V3 monoclonal antibodies, M. Howarth for providing plasmids and advice for VLP expression and purification and A. Malyutin for help with cryo-EM data collection. Cryo-EM was done in the Beckman Institute Resource Center for Transmission Electron Microscopy at Caltech. This work was supported by the National Institute of Allergy and Infectious Diseases (NIAID) of the National Institutes of Health (NIH) Grant HIVRAD P01 AI100148 (to P.J.B. and M.C.N.), NIH Grant P50 GM082545-06 (to P.J.B.), NIH Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery (CHAVI-ID) 1UM1 AI100663-01 (to M.C.N.), the Intramural Research Program of the NIAID (to M.A.M.), the National Center for Biomedical Glycomics P41GM103490 and NIGMS R01GM130915 (to L.W.), Gates CAVD grant OPP1146996 (to M.S.S. and D.C.M.), NIH/NIAID P01 AI138212 (to L.S., A.T.M. and M.C.N.) and the Robertson Fund of the Rockefeller University (M.C.N.). Additional support included an NSF GRFP (to M.E.A.), an EMBO fellowship (to J.M.), the HHMI Hanna Gray Fellowship and the Postdoctoral Enrichment Program from the Burroughs Welcome Fund (to C.O.B.). M.C.N. and D.J.I. are HHMI investigators.
Author information
Authors and Affiliations
Contributions
A.E., H.B.G., P.J.B. and M.C.N. designed the research. A.E., H.B.G., M.E.A., J.M., R.G., C.O.B., A.A.C., H.W., J.G., D.Y., J.R.K., Z.W., P.Z., L.N., K.-H.Y., J.B., H.G., A.V.V. and M.S. performed the research. A.E., H.B.G., M.E.A., J.M., R.G., T.Y.O., J.P., A.P.W., P.J.B. and M.C.N. analysed the data. D.C.M. and M.S.S. supervised in vitro neutralization assays. A.G. supervised antibody production. M.A.M. planned and supervised the immunization experiments in macaques. D.J.I. planned and supervised adjuvant production. A.T.M. and L.S. produced anti-idiotypic antibodies. L.W. planned and supervised mass spectrometry experiments. A.E., H.B.G., M.E.A., P.J.B. and M.C.N. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
There are patents on 3BNC117 and 10-1074 on which M.C.N. and P.J.B. are inventors. M.C.N. is a member of the Scientific Advisory Boards of Celldex and Frontier Biosciences.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 RC1 characterization.
a, Comparison of geometric mean half-maximum inhibitory concentrations (IC50) for V3-glycan patch bNAbs (10-1074 and PGT121) and an N156 glycan-dependent V1V2 bNAb (BG1) evaluated against HIV-1 strains that either contained or did not contain a PNGS at the indicated positions (the number of HIV-1 strains is indicated in the parentheses). Values of IC50 greater than 50 μg ml−1 were set to 50 μg ml−1 for geometric mean calculations. Whereas V3-glycan patch bNAbs showed enhanced neutralization upon removal of the N156 glycan, removal of nearby glycans (N137 or N301) diminished or had little effect on neutralization. b, ELISA data showing the binding of different classes of bNAbs to RC1, RC1-4fill and BG505. bNAbs were evaluated at 5 μg ml−1 and seven additional threefold dilutions. n = 2. RC1 and RC1-4fill show similar binding patterns for V3-glycan patch bNAbs, CD4-binding site bNAbs (CD4bs) and gp120–gp41 interface bNAbs, but reduced binding to BG1, a V1V2 bNAb that interacts with the N156 glycan (see a).
Extended Data Fig. 2 Cryo-EM data collection and processing for RC1 complexes.
a–d, A representative micrograph, selected two-dimensional class averages, orientation distribution summary, GSFSC resolution plot, local resolution (calculated using ResMap) and representative density maps contoured at 7σ for a gp41 helix and antibody CDRH3 are shown. a, The 10-1074–RC1 complex. b, The Ab275MUR–RC1 complex. c, The Ab874NHP–RC1 complex. d, The Ab897NHP–RC1 complex.
Extended Data Fig. 3 Antibody responses in wild-type mice.
a, ELISA cross-reactivity of serum from RC1-immunized wild-type mice to 11MUTB. Binding of the serum from wild-type mice primed with RC1 to RC1, 11MUTB, 10MUT and BG505 is shown in blue. Binding of the human bNAbs 10-1074 (green) and 3BNC117 (red) was evaluated at 5 μg ml−1 as a control. n = 2. b, FACS plots showing the gating strategy to isolate single RC1+RC1-glycanKO− germinal centre B cells (DUMP− (CD4−, CD8−, F4/80−, NK1.1−, CD11b−, CD11c− and Gr-1−) B220+CD95+GL7+RC1-glycanKO−RC1+) from the spleen and draining lymph nodes of wild-type mice primed with RC1 or RC1-4fill. c, Binding of the mouse antibodies Ab275MUR and Ab276MUR to a V3 loop-consensus C peptide (see Methods). Human antibodies 3869 and 3074 were used as positive controls. Antibodies were evaluated at 30 μg ml−1. n = 2. d, Representative sensograms from two independent SPR-binding experiments of Ab275MUR Fab injected over immobilized RC1 (left) or 11MUTB (right). Experimental binding curves (red) are overlaid with predicted curves (black) derived from a 1:1 binding model. Representative of 3 independent experiments. e, Binding of Ab276MUR and its inferred germline version (Ab276MURGL) to RC1 (left) and RC1-glycanKO (right) at 30 μg ml−1. The human monoclonal antibodies PGT121 (green) and 3BNC60 (red) were used as controls at 5 μg ml−1.
Extended Data Fig. 4 Characterization of RC1, RC1-4fill and VLPs.
a, SEC profiles of RC1 and RC1-4fill showing a larger apparent hydrodynamic radius for RC1-4fill compared with RC1, consistent with addition of extra glycans at the introduced PNGSs. b, ELISAs showing comparable binding of PGT122 Fab to RC1 and RC1-4fill. RLU, relative luminescence unit. c, Glycan site occupancy for each PNGS in RC1 and RC1-4fill determined by mass spectrometry. d, SDS–PAGE analysis for RC1 and RC1-4fill under non-reducing (NR), reducing (R) and PNGaseF-treated (PNG) conditions. e, SDS–PAGE analysis for VLP, SpyTagged RC1-4fill and VLP-RC1-4fill under non-reducing and reducing conditions.
Extended Data Fig. 5 Characterization of antibody responses in macaques.
a, Binding of serum from macaques primed with RC1-4fill VLPs. ELISAs of the serum from eight macaques primed with RC1-4fill VLPs and PGT121 to RC1 (black) and RC1-glycanKO (grey) are shown. b, Binding of serum from macaques primed with RC1-4fill VLPs. ELISA of the serum from eight macaques primed with RC1-4fill VLP and one naive macaque to RC1 (black) and the sequentially less modified Env proteins 11MUTB (grey) and 10MUT (white). The human bNAbs PGT121 and 3BNC60 were used as controls at 5 μg ml−1, and the serum was evaluated at a 1:100 dilution and seven additional threefold serial dilutions. c, The affinities (KD) for RC1 of different macaque antibodies isolated after a prime with VLP-RC1-4fill and the corresponding inferred germline-reverted antibodies as determined by bio-layer interferometry (OCTET). d, Binding of an anti-idiotypic antibody that recognizes the inferred germline of PGT121/10-1074 to monoclonal antibodies isolated from macaques primed with VLP-RC1-4fill. The inferred germline (iGL) of PGT121/10-1074, two chimeric antibodies comprising the mutated (MT) heavy chain (HC) and inferred germline light chain (LC) of PGT121 (PGT121 HCMT-LC iGL) or the inferred germline HC and the mutated LC of PGT121 (PGT121HC iGL-LCMT) and different inferred germline bNAbs were used as controls. a, b, d, Results are shown as the area under the ELISA curve (AUC). e, Comparison of binding mode between the vaccine-elicited antibodies (Ab275MUR, Ab874NHP and Ab897NHP) and the V3-glycan patch bNAbs 10-1074, PGT128 and PGT135. RC1 trimer is shown in grey from above and all Fabs are modelled onto the same trimer. For clarity, only one Fab per trimer is shown. f, Interactions between Ab897NHP conserved light chain motifs and RC1 gp120. Lime, DNS motif in CDRL3; red, gp120 GDIR; pink, NIG motif in CDRL1; teal, gp120 V1 loop. Each AUC value corresponds to one ELISA curve.
Supplementary information
Supplementary Table 1
IgH gene sequences from single B-cells isolated from wild-type mice immunized with RC1 or RC1-4fill. Tables show the VH, DH and JH genes, CDRH3 amino acid (AA) sequence and length and the number of nucleotide (nt) mutations in the VH gene. Colors correspond to the expanded clones in Fig. 2i.
Supplementary Table 2
IgK gene sequences from single B-cells from wild-type mice immunized with RC1 or RC1-4fill. Table shows VH and JH genes, CDRL3 amino acid (AA) sequence and length, and the number of nucleotide (nt) mutations in the VK gene. Colors correspond to clusters of light chains.
Supplementary Table 3
IgH gene sequences from single B cells isolated from 4 macaques immunized with RC1-4fill VLPs. Tables show the VH, DH and JH genes, CDRH3 amino acid (AA) sequence and length, and the number of nucleotide (nt) mutations in the VH gene. Colors correspond to the expanded clones in Fig. 3i.
Supplementary Table 4
IgL gene sequences from single B cells isolated from 4 macaques immunized with RC1-4fill VLPs. Tables show the VH and JH genes, CDRL3 amino acid (AA) sequence and length, and the number of nucleotide (nt) mutations in the VL gene. Colors correspond to clusters of light chains.
Supplementary Table 5
Monoclonal antibodies isolated from macaques immunized with RC1-4fill VLPs. The antibodies that targeted the V3-glycan patch epitope of RC1 are indicated in the V3-glycan specificity column (V3-GL SPEC) with a YES. ND=no detectable binding to RC1. AA=Amino acid.
Supplementary Table 6
Primers used for IgH and IgL macaque gene amplification.
Rights and permissions
About this article
Cite this article
Escolano, A., Gristick, H.B., Abernathy, M.E. et al. Immunization expands B cells specific to HIV-1 V3 glycan in mice and macaques. Nature 570, 468–473 (2019). https://doi.org/10.1038/s41586-019-1250-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1250-z
This article is cited by
-
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines
npj Vaccines (2024)
-
Structural basis for breadth development in the HIV-1 V3-glycan targeting DH270 antibody clonal lineage
Nature Communications (2023)
-
Comparing T- and B-cell responses to COVID-19 vaccines across varied immune backgrounds
Signal Transduction and Targeted Therapy (2023)
-
Strategies for HIV-1 vaccines that induce broadly neutralizing antibodies
Nature Reviews Immunology (2023)
-
Infection with wild-type SARS-CoV-2 elicits broadly neutralizing and protective antibodies against omicron subvariants
Nature Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.