Abstract
Polymorphonuclear myeloid-derived suppressor cells (PMN-MDSCs) are pathologically activated neutrophils that are crucial for the regulation of immune responses in cancer. These cells contribute to the failure of cancer therapies and are associated with poor clinical outcomes. Despite recent advances in the understanding of PMN-MDSC biology, the mechanisms responsible for the pathological activation of neutrophils are not well defined, and this limits the selective targeting of these cells. Here we report that mouse and human PMN-MDSCs exclusively upregulate fatty acid transport protein 2 (FATP2). Overexpression of FATP2 in PMN-MDSCs was controlled by granulocyte–macrophage colony-stimulating factor, through the activation of the STAT5 transcription factor. Deletion of FATP2 abrogated the suppressive activity of PMN-MDSCs. The main mechanism of FATP2-mediated suppressive activity involved the uptake of arachidonic acid and the synthesis of prostaglandin E2. The selective pharmacological inhibition of FATP2 abrogated the activity of PMN-MDSCs and substantially delayed tumour progression. In combination with checkpoint inhibitors, FATP2 inhibition blocked tumour progression in mice. Thus, FATP2 mediates the acquisition of immunosuppressive activity by PMN-MDSCs and represents a target to inhibit the functions of PMN-MDSCs selectively and to improve the efficiency of cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq data are deposited to the Gene Expression Omnibus (GEO) data repository with accession number GSE126885. Source Data for each figure are provided. Other data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Zhou, J., Nefedova, Y., Lei, A. & Gabrilovich, D. Neutrophils and PMN-MDSC: their biological role and interaction with stromal cells. Semin. Immunol. 35, 19–28 (2018).
Veglia, F., Perego, M. & Gabrilovich, D. Myeloid-derived suppressor cells coming of age. Nat. Immunol. 19, 108–119 (2018).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Kumar, V., Patel, S., Tcyganov, E. & Gabrilovich, D. I. The nature of myeloid-derived suppressor cells in the tumor microenvironment. Trends Immunol. 37, 208–220 (2016).
Moore, K. J., Sheedy, F. J. & Fisher, E. A. Macrophages in atherosclerosis: a dynamic balance. Nat. Rev. Immunol. 13, 709–721 (2013).
O’Neill, L. A. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23 (2016).
Hubler, M. J. & Kennedy, A. J. Role of lipids in the metabolism and activation of immune cells. J. Nutr. Biochem. 34, 1–7 (2016).
Veglia, F. et al. Lipid bodies containing oxidatively truncated lipids block antigen cross-presentation by dendritic cells in cancer. Nat. Commun. 8, 2122 (2017).
Ramakrishnan, R. et al. Oxidized lipids block antigen cross-presentation by dendritic cells in cancer. J. Immunol. 192, 2920–2931 (2014).
Herber, D. L. et al. Lipid accumulation and dendritic cell dysfunction in cancer. Nat. Med. 16, 880–886 (2010).
Cubillos-Ruiz, J. R. et al. ER stress sensor XBP1 controls anti-tumor immunity by disrupting dendritic cell homeostasis. Cell 161, 1527–1538 (2015).
Al-Khami, A. A. et al. Exogenous lipid uptake induces metabolic and functional reprogramming of tumor-associated myeloid-derived suppressor cells. OncoImmunology 6, e1344804 (2017).
den Brok, M. H., Raaijmakers, T. K., Collado-Camps, E. & Adema, G. J. Lipid droplets as immune modulators in myeloid cells. Trends Immunol. 39, 380–392 (2018).
Black, P. N. & DiRusso, C. C. Yeast acyl-CoA synthetases at the crossroads of fatty acid metabolism and regulation. Biochim. Biophys. Acta 1771, 286–298 (2007).
Black, P. N., Ahowesso, C., Montefusco, D., Saini, N. & DiRusso, C. C. Fatty acid transport proteins: targeting FATP2 as a gatekeeper involved in the transport of exogenous fatty Acids. MedChemComm 7, 612–622 (2016).
Melton, E. M., Cerny, R. L., DiRusso, C. C. & Black, P. N. Overexpression of human fatty acid transport protein 2/very long chain acyl-CoA synthetase 1 (FATP2/Acsvl1) reveals distinct patterns of trafficking of exogenous fatty acids. Biochem. Biophys. Res. 440, 743–748 (2013).
Youn, J. I., Collazo, M., Shalova, I. N., Biswas, S. K. & Gabrilovich, D. I. Characterization of the nature of granulocytic myeloid-derived suppressor cells in tumor-bearing mice. J. Leukoc. Biol. 91, 167–181 (2012).
Rodriguez, P. C. et al. Arginase I in myeloid suppressor cells is induced by COX-2 in lung carcinoma. J. Exp. Med. 202, 931–939 (2005).
Sinha, P., Clements, V. K., Fulton, A. M. & Ostrand-Rosenberg, S. Prostaglandin E2 promotes tumor progression by inducing myeloid-derived suppressor cells. Cancer Res. 67, 4507–4513 (2007).
Li, Y. et al. Myeloid-derived suppressor cells as a potential therapy for experimental autoimmune myasthenia gravis. J. Immunol. 193, 2127–2134 (2014).
Obermajer, N., Muthuswamy, R., Lesnock, J., Edwards, R. P. & Kalinski, P. Positive feedback between PGE2 and COX2 redirects the differentiation of human dendritic cells toward stable myeloid-derived suppressor cells. Blood 118, 5498–5505 (2011).
He, Y.-M. et al. Transitory presence of myeloid-derived suppressor cells in neonates is critical for control of inflammation. Nat. Med. 24, 224–231 (2018).
Casbon, A. J. et al. Invasive breast cancer reprograms early myeloid differentiation in the bone marrow to generate immunosuppressive neutrophils. Proc. Natl Acad. Sci. USA 112, E566–E575 (2015).
Condamine, T. et al. Lectin-type oxidized LDL receptor-1 distinguishes population of human polymorphonuclear myeloid-derived suppressor cells in cancer patients. Sci. Immunol. 1, aaf8943 (2016).
Ahowesso, C. et al. Chemical inhibition of fatty acid absorption and cellular uptake limits lipotoxic cell death. Biochem. Pharmacol. 98, 167–181 (2015).
Kumar, V. et al. Cancer-associated fibroblasts neutralize the anti-tumor effect of CSF1 receptor blockade by inducing PMN-MDSC infiltration of tumors. Cancer Cell 32, 654–668 (2017).
Zelenay, S. et al. Cyclooxygenase-dependent tumor growth through evasion of immunity. Cell 162, 1257–1270 (2015).
Fujita, M. et al. COX-2 blockade suppresses gliomagenesis by inhibiting myeloid-derived suppressor cells. Cancer Res. 71, 2664–2674 (2011).
Veltman, J. D. et al. COX-2 inhibition improves immunotherapy and is associated with decreased numbers of myeloid-derived suppressor cells in mesothelioma. Celecoxib influences MDSC function. BMC Cancer 10, 464 (2010).
Henao-Mejia, J. et al. Generation of genetically modified mice using the CRISPR–Cas9 genome-editing system. Cold Spring Harb. Protoc. 2016, pdb.prot090704 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Chen, H. S. et al. BET-inhibitors disrupt Rad21-dependent conformational control of KSHV latency. PLoS Pathog. 13, e1006100 (2017).
Michelini, Z. et al. Development and use of SIV-based Integrase defective lentiviral vector for immunization. Vaccine 27, 4622–4629 (2009).
Negri, D. et al. Immunization with an SIV-based IDLV expressing HIV-1 Env 1086 clade c elicits durable humoral and cellular responses in rhesus macaques. Mol. Ther. 24, 2021–2032 (2016).
Acknowledgements
This work was supported by NIH grant CA R01CA165065 to D.I.G. and V.E.K., NIH grant AI110485 to M.B., and by animal, genomics and flow cytometry core facilities of the Wistar Institute. We thank R. Schreiber for providing us with F244 cells; L. Joannas and J. Henao-Meja for generating Slc27a2fl/fl mice; W. Stremmel for providing Slc27a4fl/fl mice; and D. Weiner for providing tetramers.
Reviewer information
Nature thanks Andreas Stahl, Judith Varner and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
F.V. participated in research design, performed most of the experiments, and wrote the manuscript. V.A.T. performed lipidomics experiments. M.B. prepared lentiviruses for experiments. A.D.L. performed ChIP assays. L.D. performed mouse treatment experiments. A.V.K. performed the analysis of RNA-seq data. T.K.J.T. performed some immunological experiments. Z.S. performed metabolomics experiments and wrote the manuscript. S.B. performed seahorse experiments. F.W. performed immunohistochemistry experiments. E.R. generated COX2-knockout cells. C.D. generated lipofermata and wrote the manuscript. M.E.M. supervised metabolic experiments and reviewed the manuscript. R.H.V. supervised experiments with KPC mice and reviewed the manuscript. P.M.L. supervised ChIP experiments and reviewed the manuscript. C.M., B.N., N.H., G.M. and M.G. provided clinical samples. C.L. generated mice with PMN-targeted STAT5 deletion and performed in vivo experiments. Y.N. generated mice with PMN-targeted STAT5 deletion, and reviewed the manuscript. P.B. produced lipofermata and reviewed the manuscript. V.E.K. obtained financial support for the study, designed experiments, supervised lipidomics analysis and wrote the manuscript. D.I.G. obtained financial support for the study, designed the overall concept and specific experiments, supervised experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Lipid accumulation and expression of lipid transporters in MDSCs.
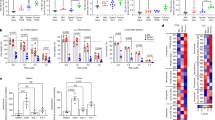
a, Lipid accumulation (BODIPY staining) in PMN-MDSCs isolated from the spleen and tumours of indicated tumour models (n = 4–8 mice per group). Each circle represents an individual mouse. Inset shows confocal image representative of two independent experiments. b, Lipid accumulation (BODIPY staining) in PMNs generated from bone marrow HPCs treated with GM-CSF and tumour explant supernatant (n = 3–5). c, LC–MS analysis of triglycerides in PMNs from control mice and PMN-MDSCs from EL4 tumour-bearing mice (n = 4). d, Lipid accumulation (BODIPY staining) in M-MDSCs isolated from spleen and tumour of indicated tumour models (n = 10). Each circle represents an individual mouse. e, Lipid accumulation (BODIPY staining) in dendritic cells (DCs) and MDSCs generated from CD204-deficient (Msr1−/−) HPCs in the presence of tumour explant supernatant (n = 3). f, Lipid accumulation in PMN-MDSCs from the spleen of wild-type and CD204-knockout (Msr1−/−) tumour-bearing mice (n = 3). g, Suppression of T cell proliferation of PMN-MDSCs from the spleen of wild-type and CD204-knockout tumour-bearing mice. Representative of four experiments, each performed in triplicate. h, Expression of Msr1, Fabp1, Fabp3, Fabp4, Fabp5, Slc27a1, Slc27a4, Slc27a5 and Cd36 in control PMNs and PMN-MDSCs isolated from the spleen and tumours of EL4 tumour-bearing mice (n = 4–5). Data are mean ± s.d. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, unpaired two-sided Student’s t-test.
Extended Data Fig. 2 Expression of gene involved in lipogenesis in PMN-MDSCs.
a, FATP2 protein in control PMNs and PMN-MDSCs from spleen of tumour-bearing mice. Representative of two experiments. b, FATP2 protein in PMNs generated in vitro from bone marrow HPCs. Representative of two experiments. For gel source data, see Supplementary Fig. 1. c, F244 tumour growth in wild-type and Slc27a2−/− (FATP2 knockout) mice on a SV129 background (n = 4). d, Verification of correct targeting of Slc27a2 by RT–qPCR (left) and FATP2 by western blot (right) analysis in PMN-MDSCs isolated from the spleen of Slc27a2fl/fl × S100a8-cre− and Slc27a2fl/fl × S100a8-cre+ tumour-bearing mice. For gel source data, see Supplementary Fig. 1. e, IFNγ production by CD8+ T cells and CD4 and CD8 T cell proliferation (n = 3) in wild-type and Slc27a2−/− mice. f, Suppressive activity of M-MDSCs isolated from wild-type or Slc27a2−/− tumour-bearing mice. Dashed lines show T cell proliferation without MDSCs. Four experiments were performed with similar results. g, Suppression of T cell proliferation of tumour-associated macrophages (TAM) from wild-type or Slc27a2−/− tumour-bearing mice. Dashed line shows T cell proliferation without macrophages. Representative of three independent experiments. h. Growth of EL4 tumours in wild-type and Slc27a2−/− mice (n = 4). Representative of two experiments. i, Suppression of T cell proliferation of PMN-MDSCs isolated from the spleen or tumours of wild-type or Slc27a2−/− mice. Representative of two independent experiments, performed in triplicate. Dotted line shows T cell proliferation without PMN-MDSCs. j, Growth of LLC tumours in wild-type and Cd36−/− mice, depleted of CD8+ T cells (n = 3). k, Lipid accumulation (BODIPY staining) in PMN-MDSCs and M-MDSCs isolated from the spleen and tumours of Cd36−/− mice (n = 3). Data are mean ± s.d. ****P < 0.0001, unpaired two-sided Student’s t-test.
Extended Data Fig. 3 Effect of FATP2 knockdown on mRNA gene expression.
a, Expression heatmap for genes affected by FATP2 depletion by at least fivefold. b, Number of significantly affected genes (FDR < 5%) for different fold change thresholds. c, List of upstream regulators whose targets were found by ingenuity pathway analysis as significantly enriched among genes affected by FATP2 knockdown. n, number of affected targets; p, enrichment P value; Z, activation z scores calculated by IPA represent predicted regulator state based on the known effect on target and direction of mRNA change. Negative activation z scores predict inhibition, and positive z scores denote activation of the regulator in the FATP2-knockout mice.
Extended Data Fig. 4 LC–MS analysis of lipids from wild-type and Slc27a2−/− PMN-MDSCs.
a, Triglycerides (TG) in PMN-MDSCs from spleens of EL4 wild-type (n = 7) and FATP2-knockout (Slc27a2−/−) tumour-bearing mice (n = 6). Triglyceride species containing linoleic acid (18:2), docosapentaenoic acid (22:5), and docosahexaenoic acid (22:6) (n = 7). b, Cholesterol esters in PMN-MDSCs from spleens of LLC wild-type (n = 7) and FATP2-knockout (Slc27a2−/−) tumour-bearing mice (n = 6). Total cholesterol ester (CE) and arachidonoyl-containing (20:4) cholesterol ester. c, Fatty acids in PMN-MDSCs from the spleen of wild-type (n = 12) and Slc27a2−/− (n = 11) tumour-bearing mice. Linoleic acid (18:2), docosapentaenoic acid (22:5), and docosahexaenoic acid (22:6) fatty acids. d, Distribution of major phospholipids in PMN-MDSCs from Slc27a2−/− (FATP2 KO) (n = 10) and wild-type (n = 12) mice. PC, phosphatidylcholine; PE, phosphatidylethanolamine; PI, phosphatidylinositol; PS, phosphatidylserine. Subscript ‘AA’ denotes arachidonoyl-containing. e, Content of phospholipids containing arachidonic acid in phosphatidylethanolamine, phosphatidylcholine, phosphatidylinositol and phosphatidylserine (n = 12 for wild-type and n = 10 for FAT2 KO mice). f, Content of AA-d11-labelled phospholipids (phosphatidylinositol, phosphatidylglycerol (PG), phosphatidic acid (PA) and phosphatidylserine), n = 5. Data are mean ± s.d.; each circle indicates an individual mouse. *P < 0.05, **P < 0.01, unpaired two-sided Student’s t-test.
Extended Data Fig. 5 Metabolomic analysis and expression of fatty acid oxidation-related genes in PMN-MDSCs.
a, Oxygen consumption rate (OCR) (left) and basal OCR (right) of wild-type and Slc27a2−/− (FATP2 KO) PMN-MDSCs. Representative of two experiments (left panel; n = 3–4). Cumulative results shown in the right panel, and each circle indicates an individual mouse (n = 7). No statistical differences (P > 0.05) were determined by unpaired two-sided Student’s t-test. b, Extracellular acidification rate (ECAR) (left) and basal ECAR (right) of wild-type and FATP2-knockout PMN-MDSCs. Representative of two independent experiments (left; n = 3–4). Cumulative results shown in the right panel, and each circle indicates an individual mouse (n = 7). *P < 0.05, unpaired two-sided Student’s t-test. c, 13C-labelling of intermediates and associated amino acids of the tricarboxylic acid cycle. Ex vivo MDSCs were cultured in physiological-like medium supplemented with BSA-conjugated 13C16-palmitate and GM-CSF for 18 h. Metabolites were then extracted and analysed by high-resolution LC–MS. 13C isotopologues (M + x) for each metabolite are represented as normalized stacked bars. Representative of three biological replicates. No statistical differences (P > 0.05) were determined by unpaired two-sided Student’s t-test. d, Expression of genes involved in fatty acid oxidation. RT–qPCR analysis of Cpt1a, Acadm and Hadha expression in control PMNs and PMN-MDSCs isolated from the spleen and tumours of tumour-bearing mice. Each group included 3-6 mice. Data are mean ± s.d.
Extended Data Fig. 6 Exchange of nutrients with the media.
Ex vivo MDSCs were cultured in physiological-like medium supplemented with GM-CSF for 18 h. Metabolites were then extracted from the medium and analysed by LC–MS. Upward bars represent efflux from the cells into the media, and downward bars represent uptake (or depletion) from the medium by the cells. Data are normalized to protein content after extraction. Data are mean ± s.d. (n = 3).
Extended Data Fig. 7 Effect of arachidonic acid on production of PGE2 and suppressive activity of PMN-MDSCs.
a, LC–MS analysis of PGE2 in PMNs from control mice and PMN-MDSCs from EL4 and CT26 tumour-bearing mice (n = 3). b, PGE2 release (measured by ELISA) by control PMNs (n = 4) and PMN-MDSCs from wild-type (n = 11), and Slc27a2−/− (FATP2 KO) (n = 8) LLC tumour-bearing mice. c, Expression of Ptges in PMN-MDSCs isolated form the spleen of EL4 (n = 13–15), KPC (n = 3) and RET melanoma (n = 3–6) tumour-bearing mice. d, Expression of Ptgs2 and Ptges (measured by RT–qPCR) in PMN-MDSCs (n = 6). e, Expression of Arg1 and Nos2 (measured by RT–qPCR) in spleen PMN-MDSCs from wild-type and FATP2 KO EL4 tumour-bearing mice (n = 3–5). f, Flow cytometry of myeloid cells differentiated from HPCs cultured in the presence of arachidonic acid. Representative of three experiments. g, Expression of Arg1, Nos2 and Nox2 in PMNs isolated from HPCs cultured in the presence of arachidonic acid. Data are pooled from six independent experiments. h, pSTAT5 expression by flow cytometry at different time points in PMNs isolated from mouse bone marrow treated with different amounts of GM-CSF. Representative of three independent experiments. i, LLC tumour growth (n = 4) in Slc27afl/fl × S100a8-cre− (Cre−) and Slc27afl/fl × S100a8-cre+ (Cre+) mice. j, Slc27a2 expression (measured by RT–qPCR) in PMN-MDSCs from the spleen of wild-type and knockout tumour-bearing mice (n = 4). *P < 0.05, **P < 0.01, unpaired two-sided Student’s t-test. Data are mean ± s.d.
Extended Data Fig. 8 Lipid accumulation in MDSCs from patients with cancer.
a, Lipid accumulation (measured by BODIPY staining) in M-MDSCs isolated from the blood of patients with cancer or healthy individuals. Each circle indicates an individual. b, Amount of lipids (BODIPY staining) in M-MDSCs from blood and tumour tissue of patients with cancer. Each circle indicates an individual (n = 5). c, RNA-seq analysis of genes involved in lipid accumulation in human LOX1+ PMN-MDSCs and LOX1− PMNs (n = 4). d, PTGES expression in LOX1+ and LOX1− PMNs from blood of patients with cancer. Fold change compared with LOX1− PMNs (n = 3). e, SLC27A2 expression in M-MDSCs and monocytes isolated from blood of patients with cancer and healthy donors, respectively. Each circle indicates an individual (n = 4–6). Data are mean ± s.d. f, pSTAT5 by flow cytometry at different time points, in human PMNs isolated from the blood of healthy donors and treated with different amounts of GM-CSF. g, FATP2 in PMNs isolated from blood of healthy donors and treated with GM-CSF. Representative of three independent experiments. For gel source data, see Supplementary Fig. 1. h, Content of total phosphatidylethanolamine (PE) and arachidonoyl-containing phosphatidylethanolamine (AA-PE) in PMN-MDSCs isolated from patients with lung cancer (n = 5) or healthy donors (n = 4). Each circle indicates an individual. Data are mean ± s.d. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, unpaired two-sided Student’s t-test.
Extended Data Fig. 9 Effect of lipofermata treatment on tumour-bearing mice.
a, MTT assay after three-day incubation of tumour cells with indicated concentration of lipofermata. b, Percentage and absolute number of tumour-associated antigen (E7-derived peptide)-specific CD8+ T cells in draining lymph nodes of mice bearing TC-1 tumour and treated with lipofermata (n = 3). c, Growth of TC-1 tumours in mice treated with CTLA4 antibody and lipofermata (n = 5). d, CD8+ T cell infiltration of TC-1 tumours in mice treated with CTLA4 antibody and lipofermata. Left, representative staining of two different mice is shown. Scale bars, 50 µm. Right, the number of CD8+ T cells per mm2 (n = 2). e, Growth of TC-1 tumours in mice treated with PD1 antibody and lipofermata (n = 5). Data are mean ± s.d. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, two-sided Student’s t-test (b) or two-way ANOVA test with correction for repeated measurements (e).
Supplementary information
Supplemental Figure 1
Uncut gels presented in main and extended data figures. Each gel is referenced in the figures in the manuscript.
Supplemental information
This file contains a detailed description of lipidomics analysis performed in this study; detailed information on lentiviral vector production and validation; Supplemental Table 1 (Antibodies and reagents used in the study) and Supplemental Table 2 (Sequences of primers using in RT–qPCR).
Source data
Rights and permissions
About this article
Cite this article
Veglia, F., Tyurin, V.A., Blasi, M. et al. Fatty acid transport protein 2 reprograms neutrophils in cancer. Nature 569, 73–78 (2019). https://doi.org/10.1038/s41586-019-1118-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1118-2
This article is cited by
-
Reprogramming of lipid metabolism in the tumor microenvironment: a strategy for tumor immunotherapy
Lipids in Health and Disease (2024)
-
Bladder-cancer-derived exosomal circRNA_0013936 promotes suppressive immunity by up-regulating fatty acid transporter protein 2 and down-regulating receptor-interacting protein kinase 3 in PMN-MDSCs
Molecular Cancer (2024)
-
The application of nanoparticles-based ferroptosis, pyroptosis and autophagy in cancer immunotherapy
Journal of Nanobiotechnology (2024)
-
FFAR2 expressing myeloid-derived suppressor cells drive cancer immunoevasion
Journal of Hematology & Oncology (2024)
-
Oleuropein-driven reprogramming of the myeloid cell compartment to sensitise tumours to PD-1/PD-L1 blockade strategies
British Journal of Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.