Abstract
Pancreatic ductal adenocarcinoma (PDAC) remains recalcitrant to all forms of cancer treatment and carries a five-year survival rate of only 8%1. Inhibition of oncogenic KRAS (hereafter KRAS*), the earliest lesion in disease development that is present in more than 90% of PDACs, and its signalling surrogates has yielded encouraging preclinical results with experimental agents2,3,4. However, KRAS*-independent disease recurrence following genetic extinction of Kras* in mouse models anticipates the need for co-extinction strategies5,6. Multiple oncogenic processes are initiated at the cell surface, where KRAS* physically and functionally interacts to direct signalling that is essential for malignant transformation and tumour maintenance. Insights into the complexity of the functional cell-surface-protein repertoire (surfaceome) have been technologically limited until recently and—in the case of PDAC—the genetic control of the function and composition of the PDAC surfaceome in the context of KRAS* signalling remains largely unknown. Here we develop an unbiased, functional target-discovery platform to query KRAS*-dependent changes of the PDAC surfaceome, which reveals syndecan 1 (SDC1, also known as CD138) as a protein that is upregulated at the cell surface by KRAS*. Localization of SDC1 at the cell surface—where it regulates macropinocytosis, an essential metabolic pathway that fuels PDAC cell growth—is essential for disease maintenance and progression. Thus, our study forges a mechanistic link between KRAS* signalling and a targetable molecule driving nutrient salvage pathways in PDAC and validates oncogene-driven surfaceome annotation as a strategy to identify cancer-specific vulnerabilities.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available from the corresponding author upon reasonable request.
Code availability
Code for screen hit analysis are available upon request from the authors.
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
Mann, K. M., Ying, H., Juan, J., Jenkins, N. A. & Copeland, N. G. KRAS-related proteins in pancreatic cancer. Pharmacol. Ther. 168, 29–42 (2016).
Pettazzoni, P. et al. Genetic events that limit the efficacy of MEK and RTK inhibitor therapies in a mouse model of KRAS-driven pancreatic cancer. Cancer Res. 75, 1091–1101 (2015).
Shimizu, T. et al. The clinical effect of the dual-targeting strategy involving PI3K/AKT/mTOR and RAS/MEK/ERK pathways in patients with advanced cancer. Clin. Cancer Res. 18, 2316–2325 (2012).
Kapoor, A. et al. Yap1 activation enables bypass of oncogenic Kras addiction in pancreatic cancer. Cell 158, 185–197 (2014).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
da Cunha, J. P. et al. Bioinformatics construction of the human cell surfaceome. Proc. Natl Acad. Sci. USA 106, 16752–16757 (2009).
Binder, J. X. et al. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database 2014, bau012 (2014).
Biankin, A. V. et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491, 399–405 (2012).
Capello, M. et al. Carboxylesterase 2 as a determinant of response to irinotecan and neoadjuvant FOLFIRINOX therapy in pancreatic ductal adenocarcinoma. J. Natl. Cancer Inst. 107, djv132 (2015).
Carugo, A. et al. In vivo functional platform targeting patient-derived xenografts identifies WDR5-Myc association as a critical determinant of pancreatic cancer. Cell Reports 16, 133–147 (2016).
Alexander, C. M. et al. Syndecan-1 is required for Wnt-1-induced mammary tumorigenesis in mice. Nat. Genet. 25, 329–332 (2000).
Subramanian, S. V., Fitzgerald, M. L. & Bernfield, M. Regulated shedding of syndecan-1 and -4 ectodomains by thrombin and growth factor receptor activation. J. Biol. Chem. 272, 14713–14720 (1997).
Chen, K. & Williams, K. J. Molecular mediators for raft-dependent endocytosis of syndecan-1, a highly conserved, multifunctional receptor. J. Biol. Chem. 288, 13988–13999 (2013).
Zimmermann, P. et al. Syndecan recycling is controlled by syntenin–PIP2 interaction and Arf6. Dev. Cell 9, 377–388 (2005).
Robertson, S. E. et al. Extracellular signal-regulated kinase regulates clathrin-independent endosomal trafficking. Mol. Biol. Cell 17, 645–657 (2006).
Honda, A. et al. Phosphatidylinositol 4-phosphate 5-kinase α is a downstream effector of the small G protein ARF6 in membrane ruffle formation. Cell 99, 521–532 (1999).
Bar-Sagi, D. & Feramisco, J. R. Induction of membrane ruffling and fluid-phase pinocytosis in quiescent fibroblasts by ras proteins. Science 233, 1061–1068 (1986).
Commisso, C. et al. Macropinocytosis of protein is an amino acid supply route in Ras-transformed cells. Nature 497, 633–637 (2013).
Brown, F. D., Rozelle, A. L., Yin, H. L., Balla, T. & Donaldson, J. G. Phosphatidylinositol 4,5-bisphosphate and Arf6-regulated membrane traffic. J. Cell Biol. 154, 1007–1017 (2001).
Radhakrishna, H., Klausner, R. D. & Donaldson, J. G. Aluminum fluoride stimulates surface protrusions in cells overexpressing the ARF6 GTPase. J. Cell Biol. 134, 935–947 (1996).
Fujii, M., Kawai, K., Egami, Y. & Araki, N. Dissecting the roles of Rac1 activation and deactivation in macropinocytosis using microscopic photo-manipulation. Sci. Rep. 3, 2385 (2013).
Su, G., Blaine, S. A., Qiao, D. & Friedl, A. Shedding of syndecan-1 by stromal fibroblasts stimulates human breast cancer cell proliferation via FGF2 activation. J. Biol. Chem. 282, 14906–14915 (2007).
Langford, J. K., Yang, Y., Kieber-Emmons, T. & Sanderson, R. D. Identification of an invasion regulatory domain within the core protein of syndecan-1. J. Biol. Chem. 280, 3467–3473 (2005).
Beekman, J. M. & Coffer, P. J. The ins and outs of syntenin, a multifunctional intracellular adaptor protein. J. Cell Sci. 121, 1349–1355 (2008).
Taguchi, A. et al. Lung cancer signatures in plasma based on proteome profiling of mouse tumor models. Cancer Cell 20, 289–299 (2011).
Aguirre, A. J. et al. Activated Kras and Ink4a/Arf deficiency cooperate to produce metastatic pancreatic ductal adenocarcinoma. Genes Dev. 17, 3112–3126 (2003).
Burbach, B. J., Friedl, A., Mundhenke, C. & Rapraeger, A. C. Syndecan-1 accumulates in lysosomes of poorly differentiated breast carcinoma cells. Matrix Biol. 22, 163–177 (2003).
Commisso, C., Flinn, R. J. & Bar-Sagi, D. Determining the macropinocytic index of cells through a quantitative image-based assay. Nat. Protoc. 9, 182–192 (2014).
Acknowledgements
We thank D. Bar-Sagi and C. Ramirez (New York University) for their suggestions and constructive feedback; R. Sanderson at University of Alabama for sharing SDC1 constructs; T. Tieu, M. Peoples, J. Ren and Q. Chang for technical assistance; D. Aten for help with the graphical abstract; and our colleagues at the Institute for Applied Cancer Science (IACS), the Flow Cytometry and Cellular Imaging Core, the Sequencing and Microarray Facility, the Department of Veterinary Medicine, Medical Graphics and Photography at The MD Anderson Cancer Center (MDACC) (Cancer Center Support Grant, CA016672). We thank all members of the laboratories of G.F.D., R.A.D., H.Y. and S.H. for discussions and reagents. The research was supported by the Odyssey Postdoctoral Fellowship at MDACC, the PanCAN-AACR Pathway to Leadership Grant (16-70-25-YAO) and 2017 Hirshberg foundation for pancreatic cancer research to W.Y.; Pancreatic Cancer Moon Shot Program at MDACC, CPRIT (RP160471), DOD (W81XWH-11-1-0418), and Harrington Discovery Institute Grant to G.F.D.; P01 Grant (P01CA117969 12, NIH) to H.W., A.M., R.A.D., G.F.D. and H.Y.
Reviewer information
Nature thanks Channing Der, Harald Alfred Stenmark and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
W.Y. and G.F.D. designed the studies, interpreted the data and wrote the manuscript. Y.A.W. and R.A.D. provided valuable suggestions and intellectual input. W.Y., A.K., A.T. and S.H. performed the proteomic study and analysis. W.Y. and J.L.R carried out the loss-of-function screen. W.Y., W.W., H.J., J.L., L.Y., A.K., P.H., Z.C., Z.T., L.N., Q.W., V.R., and G.J.M. conducted other experiments. S.S., J.L., J.Y., and Z.T. were responsible for bioinformatics analysis. Z.X., S.J., P. Deng, P. Den, B.H., P.F.P., A.C., S.X.Z., I.L.H, N.F. and T.P.H. provided technical assistance. H.W. and A.M. provided assistance with pathology. J.B.F. provided patient-derived xenograft cells. Y.A.W., A.V., H.Y., A.K.D. and R.A.D. provided valuable intellectual input and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.A.D. is a co-founder, advisor and director of Tvardi Therapeutics. G.F.D. reports personal fees from and stock ownership in Karyopharm Therapeutics, Forma Therapeutics, Metabomed, BiovelocITA, Nurix and Orionis Biosciences; and personal fees from Blueprint Medicines, Taiho Pharmaceutical, Symphogen and Helsinn Ventures.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Interrogation of surfaceome changes upon KRAS* signalling by SILAC-based proteomic analysis.
a, Western blot for p-ERK and KRAS in iKras* p53L/+ cells in the presence (ON) or absence (OFF) of doxycycline for 24 h. The experiment was repeated twice with similar results. b, c, Venn diagram showing the total number of proteins identified (b) and quantified (c) by SILAC-based proteomic analysis upon Kras* inactivation in three independent iKras* p53L/+ cells. d, e, Venn diagram showing the number of decreased (KRAS* ON/OFF ratio > 1.2) (d) or increased (KRAS* ON/OFF ratio < 0.83) (e) proteins quantified with normalized MS2 counts >2 from the SILAC-based proteomic analysis on Kras* inactivation in three independent iKras* p53L/+ cell cultures. f, Immunohistochemistry for EPHA2 and CD9 in orthotopic xenograft tumours from iKras* p53L/+ model in the presence (ON) or absence (OFF) of doxycycline for 24 h (scale bar, 100 μm). The experiment was repeated twice with similar results. g, Top-20 surfaceome genes preferentially regulated by KRAS*.
Extended Data Fig. 2 Functional surfaceome analysis identified SDC1 as a KRAS*-dependent surface candidate.
a, Normalized counts showing the distribution of reference and tumour barcodes to establish library coverage in vivo in three independent iKras* p53L/+ tumour cell cultures, showing similar results across expreiments. In vivo screens were conducted in 3–5 mice for for each cell culture. b, Correlation matrix among replicates of orthotopic xenograft-derived AK192, AK196 and AK10965 tumours screened with the surfacome-targeting shRNA library (barcode-level fold-change comparison, Pearson correlation coefficient). c, Positive (Psma and Rpl30) and negative (Luc) controls were plotted applying the RSP/logP score. d, Venn diagram showing the number of hits identified from three independent iKras* p53L/+ tumour cell cultures.
Extended Data Fig. 3 SDC1 membrane expression is elevated in KRAS*-driven PDAC.
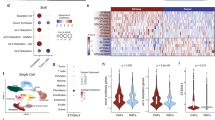
a, iKras* p53L/+ tumour cells were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h and subjected to immunofluorescence for SDC1 (red) and DAPI (blue) (scale bar, 20 µm). For ReON samples, cells were grown in the absence of doxycycline for 48 h followed by doxycycline treatment for 24 h. Representative images are shown; experiments were repeated three times with similar results. b, iKras* p53L/+ tumour cells were grown in the presence (ON) or absence (OFF) of doxycycline for indicated time periods or treated with MEK inhibitor (AZD8330, 100 nM) for 16–18 h and cell-surface levels of CTNNA1 and PMCA were measured by FACS analysis. Quantification of fluorescence intensity is shown. Representative figures and data from biological duplicates are shown. c, iKras* p53L/+ tumour cells were grown in the presence (ON) or absence (OFF) of doxycycline for 24 h and Sdc1 mRNA level was measured by quantitative PCR with reverse transcription (RT–qPCR) (n = 3, Data are mean + s.d.). Experiments were repeated twice with similar results. d, iKras* p53L/+ tumour cells stably overexpressing SDC1 (Sdc1-OE) or empty vector (Vec) and a stable single clone with double nickase-mediated SDC1 deletion (SC1) derived from iKras* p53L/+ tumour cells were blotted to validate SDC1 expression. Some samples were also treated with heparinase (Hepa) and chondroitinase (Cho) before western blot analysis. Experiments were repeated twice with similar results. e, iKras* p53L/+ tumour cells grown in the presence (ON) or absence (OFF) of doxycycline for indicated time periods were processed for western blot analysis to detect SDC1, vinculin, p-ERK and KRAS. Experiments were repeated twice with similar results. f, Representative immunohistochemistry for SDC1 from the LSL-KRAS PDAC mouse model showing membrane SDC1 level in normal pancreas, acinar-to-ductal metaplasia (ADM) and pancreatic intraepithelial neoplasia (PanIN). Experiments were repeated twice with similar results. Scale bar, 200 µm. g, Left, representative immunohistochemistry for SDC1 in normal pancreas, tumour adjacent pancreatitis (TAP), PanINs and invasive tumours (PDA) from the human PDAC TMA. Right, quantification of the TMA scores. Scale bar, 50 µm. h, Representative H&E staining and SDC1 immunohistochemistry from the TMA analysis. Representative images of SDC1 immunohistochemistry showing staining classified as low (score 1), intermediate (score 2) and high (score 3). Scale bar, 200 µm. i, mRNA expression of SDC1 in public microarray datasets. Data are mean ± s.d.; P values were determined by unpaired two-sided Student’s t-test. j, Top, outline of experimental design for sequential cerulein and doxycycline treatment in iKras* mice (top). Chronic pancreatitis was induced in iKras* mice by cerulein injection for six weeks. The mice were then treated with doxycycline for indicated times. Bottom, pancreatic or tumour tissues were subjected to H&E staining or immunohistochemistry for CK19 and SDC1. Experiments were repeated twice with similar results. Scale bar, 100 µm.
Extended Data Fig. 4 SDC1 is required for tumorigenic activity of KRAS*-driven PDAC.
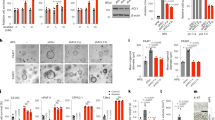
a, Representative images of clonogenic assay for two independent iKras* p53L/+ PDAC cells infected with scrambled shRNA or shRNA against Sdc1. Experiments were repeated three times with similar results. b, c, Validation of Sdc1 knockdown by RT–qPCR (b) and FACS analysis with anti-SDC1 antibody (c) in iKras* p53L/+ PDAC cells infected with scrambled shRNA or shRNA against Sdc1. In b, n = 3; data are mean ± s.d.; P values were determined by unpaired two-sided Student’s t-test. In c, experiments were repeated three times with similar results. d, Two independent iKras* p53L/+ tumour cells stably expressing SDC1 or empty vector were infected with scrambled shRNA or shRNA against Sdc1. Quantification of clonogenic assay is shown (n = 4 biological replicates; data are mean ± s.d.). P values were determined by unpaired two-sided Student’s t-test. e, Digital photographs of dissected tumours from mice implanted with iKras* p53L/+ tumour cells containing scrambled shRNA or shRNA against Sdc1. f, Validation of double nickase-mediated Sdc1 deletion in iKras* p53L/+ PDAC cells using 281-2 anti-SDC1 antibody and FACS analysis. Experiments were repeated twice with similar results. g, Representative figure of clonogenic assay for iKras* p53L/+ PDAC cells with wild-type SDC1 or double nickase-mediated Sdc1 deletion. Experiments were repeated twice with similar results. h, Digital photographs of dissected tumours from mice implanted with iKras* p53L/+ tumour cells with wild-type SDC1 or double nickase-mediated Sdc1 deletion. i, Representative figure of clonogenic assay for human PDAC cells infected with scrambled shRNA or shRNA against SDC1. Experiments were repeated twice with similar results. j, Validation of SDC1 knockdown by shRNA in human PDAC cell lines from (i) using DL-101 anti-SDC1 antibody and FACS analysis. Experiments were repeated twice with similar results. k, Digital photographs of dissected tumours from subcutaneous xenografts of mice implanted with PATC69 cells infected with scrambled shRNA or shRNA against SDC1.
Extended Data Fig. 5 SDC1 loss leads to changes in tumour microenvironment.
a, Validation of SDC1 level by FACS analysis in primary cultures derived from the PDAC mouse model with indicated Sdc1 genotypes. Experiments were repeated twice with similar results. b, c, Quantification (a) and representative images (b) from immuno-profiling of SDC1 wild-type and knockout (KO) tumours from the GEMM model by immunohistochemistry or immunofluorescence staining (n = 20 random images from four biological replicates). P values were determined by unpaired two-sided Student’s t-test.
Extended Data Fig. 6 KRAS* induces SDC1 membrane expression via the MAPK pathway.
a, b, iKras* p53L/+ PDAC cells were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h. ReON cells were grown in the absence of doxycycline for 48 h and then treated with doxycycline for 24 h. Treatment of ON or ReON samples with trametinib (50 nM) or BKM120 (100 nM) for 16–18 h is indicated. a, Cells were stained with anti-SDC1 antibody and surface SDC1 was measured via FACS. Quantification of fluorescence intensity from biological duplicates is shown as mean ± s.d. b, Cell lysates were blotted for phospho-ERK, phospho-AKT and KRAS, with vinculin as loading control. Experiments were repeated twice with similar results. c–f, iKras* p53L/+ PDAC cells were treated with different concentrations of the MEK inhibitor tramatinib (c, d) or AZD8330 (e, f) overnight (16–18h), and the live cells were prepared for FACS analysis for SDC1 (c, e; data are mean ± s.d.); cell lysates were blotted for phospho-ERK, phospho-MEK and KRAS, with vinculin as loading control (d, f). Experiments were repeated twice with similar results. g, Outline of experimental design to measure SDC1 shedding (left). iKras* p53L/+ PDAC cells were grown in the presence (ON) or absence (OFF) of doxycycline for 24 or 48 h. Medium was collected and shed SDC1 was measured by anti-SDC1 enzyme-linked immunosorbent assay (ELISA) (right) (n = 4 biological replicates; data are mean ± s.d.). h, Left, outline of experimental design to measure surface SDC1 internalization. Right, iKras* p53L/+ PDAC cells were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h. Cells were then incubated at 37 °C for indicated times and surface SDC1 was labelled with anti-SDC1 antibody at 4 °C after internalization for indicated times. FACS was performed to detect remaining SDC1 on the cell membrane (n = 2 biological replicates; data are mean ± s.d.). i–j, Outline of experimental design to measure surface SDC1 recycling (i, left). iKras* p53L/+ PDAC cells were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h. Surface SDC1 was labelled with anti-SDC1 at 4 °C. Cells were then incubated at 37 °C for 30 min to allow SDC1 internalization. Cells were then incubated on ice to stop internalization; subsequently, cells were returned to 37 °C for indicated times to allow SDC1 recycling. Recycled SDC1 was measured by FACS (i, right). j, Histograms of FACS analysis, showing recycled cell-surface SDC1 at the indicated time points. Experiments were repeated twice with similar results.
Extended Data Fig. 7 The MAPK–PSD4–ARF6 axis mediates KRAS*-dependent SDC1 membrane localization.
a, Cells were grown in presence (ON) or absence (OFF) of doxycycline or treated with AZD8330 (100 nM) for 16–18 h. Top, ARF6 activity was measured with GGA3–PBD pull-down assay. Bar graph, ARF6 activity was calculated as ratio of captured ARF6:input ARF6/vinculin. b, iKras* p53L/+ tumour cells were grown in the presence (ON) or absence (OFF) of doxycycline, or treated with AZD8330 (100 nM) for 16–18 h. Cell lysates were used for measurement of PIPK activity (n = 3 biological replicates; data are mean ± s.d.). P values were determined by unpaired two-sided Student’s t-test. c, Representative images of morphology change in iKras* p53L/+ tumour cells with dominant negative ARF6 (ARF6(T27N)) or constitutively active ARF6 (ARF6(Q67L)). Experiments were repeated three times with similar results. d, Top and middle, iKras* p53L/+ tumour cells stably expressing ARF6(Q67L) or empty vector were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h and surface SDC1 was measured by FACS using anti-SDC1 antibody. Bottom, fluorescence intensity of surface SDC1 (n = 3 biological replicates; data are mean ± s.d.). P values were determined by paired two-sided Student’s t-test. e, iKras* p53L/+ tumour cells stably expressing ARF6(T27N) or empty vector were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h and surface SDC1 was measured by FACS using anti-SDC1 antibody. Representative histograms (top and middle) and bar chart (bottom) of fluorescence intensity are shown (n = 4 biological replicates; data are mean ± s.d.). P values were determined by paired two-sided Student’s t-test. f, mRNA expression of ARF6 GTPase-activating proteins and guanine nucleotide exchange factors in a iKras* p53L/+ tumour cell microarray dataset on KRAS(G12D) inactivation (n = 4 biological replicates). P values were determined by unpaired two-sided Student’s t-test. g, iKras* p53L/+ tumour cells were grown in the presence (ON) or absence (OFF) of doxycycline or treated with AZD8330 (50 nM) for 16–18 h, and Psd4 mRNA level was measured by RT–qPCR (n = 3). P values were determined by unpaired two-sided Student’s t-test. h, MiaPaCa2 cells containing doxycycline-inducible shRNA targeting human KRAS were grown in the absence (OFF) or presence (ON) of doxycycline, or treated with trametinib (50 nM), AZD8330 (50 nM) or BKM120 (100 nM) for 18 h. Cell lysates were blotted for PSD4, phospho-ERK and KRAS. Arrow, PSD4 band. Experiments were repeated twice with similar results. i, iKras* p53L/+ tumour cells stably expressing Psd4 or empty vector were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h. ARF6 activity was measured by GGA3–PBD pull-down assay. Input lysates were immunoblotted to validate expression of ARF6, p-ERK, p-MEK, PSD4 and KRAS. Experiments were repeated twice with similar results. j, iKras* p53L/+ tumour cells stably expressing Psd4 or empty vector were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h and surface SDC1 was measured by FACS using anti-SDC1 antibody. Representative histograms of FACS analysis (top) and bar graph of fluorescence intensity (bottom) of surface SDC1 are shown (n = 3 biological replicates; data are mean ± s.d.). P values were determined by paired two-sided Student’s t-test.
Extended Data Fig. 8 SDC1 mediates macropinocytosis in KRAS*-driven mouse PDAC cells.
a, b, iKras* p53L/+ tumour cells were grown in the presence (ON) or absence (OFF) of doxycycline for 24 h. For a positive control, cells grown in the presence of doxycycline were treated with EIPA (50 µM) for 16 h. Macropinocytosis was visualized with TMR–dextran (scale bar, 20 µm) (a) and quantified (b) (n = 8 random areas for ON/OFF groups, n = 5 for EIPA group; data are mean ± s.d.). Data are representative of three independent experiments with similar results. c, Validation of Sdc1 knockdown by qPCR in iKras* p53L/+ tumour cells (n = 3; data are mean ± s.d.). d, Macropinocytosis index in KPC tumour cells containing scrambled shRNA or shRNA directed against Sdc1 (n = 6 random areas; data are mean ± s.d.). e, f, iKras* p53L/+ tumour cells stably expressing Psd4 or empty vector were grown in the presence (ON) or absence (OFF) of doxycycline for 48 h. Macropinocytosis was visualized with TMR–dextran (f; scale bar, 20 µm) and quantified (e; n = 6 random areas for Vec-ON, PSD4-ON and PSD4-OFF groups, n = 8 for Vec-OFF group; data are mean ± s.d.). Data are representative of two independent experiments with similar results. g, iKras* p53L/+ tumour cells stably expressing Sdc1 or empty vector were grown in the presence (KRAS*-ON) or absence (KRAS*-OFF) of doxycycline for 48 h. Macropinocytosis was visualized with TMR–dextran (scale bar, 20 µm). Experiments were repeated twice with similar results. h, RHOA activity in iKras* p53L/+ tumour cells containing scrambled shRNA or shRNA against Sdc1 was measured by G-LISA activation assay (n = 2 biological replicates; data are mean ± s.d.). i, RAC1 activity in iKras* p53L/+ tumour cells containing RAC1(Q61L) or empty vector was measured by G-LISA activation assay (n = 2 biological replicates; data are mean ± s.d.). j, k, iKras* p53L/+ tumour cells stably expressing RAC1(Q61L) or empty vector were infected with scrambled shRNA or shRNA against Sdc1. Macropinocytosis was visualized with TMR–dextran (scale bar, 20 µm) (j) and quantified (k) (n = 8 random areas; data are mean ± s.d.). Data are representative of two independent experiments with similar results. l, iKras* p53L/+ tumour cells were cultured in medium with serial concentrations of glutamine. Growth rate was measured and the timelapse graph was generated using the Incucyte live-cell analysis system (n = 3 technical replicates; data are mean ± s.d.). Data are representative of two independent experiments with similar results.
Extended Data Fig. 9 SDCBP is required for SDC1-mediated macropinocytosis.
a, iKras* p53L/+ tumour cells containing wild-type or indicated mutant constructs of SDC1 or empty vector were infected with scrambled shRNA or shRNA against Sdc1 and surface expression of SDC1 was measured by FACS; representative histograms are shown. Experiments were repeated twice with similar results. b, Shed SDC1 from iKras* p53L/+ tumour cells stably expressing wild-type or soluble mutant SDC1 (Sol), or empty vector was measured by anti-SDC1 ELISA (n = 5 biological replicates; data are mean ± s.d.). c–e, iKras* p53L/+ tumour cells containing wild type or indicated mutant constructs of SDC1 or empty vector were infected with scrambled shRNA or shRNA against Sdc1 to measure RAC1 activity using G-LISA activation assay (c) (n = 4 biological replcates) or macropinocytosis index by visualizing with TMR–dextran (scale bar, 20 µm) and subsequent quantification (d, e). Data are mean ± s.d. Experiments were repeated twice with similar results. f, g, iKras* p53L/+ tumour cells containing wild-type or different mutant constructs of SDC1 or empty vector were infected with scrambled shRNA or shRNA against Sdc1 and subcutaneously injected into nude mice. Photographs of dissected tumours (f) and tumour volume at indicated time points (g) are shown (n = 5 for Vec Scr, Sol-ShSdc1, Ect-ShSdc1, Gag-ShSdc1 and WT-ShSdc1 groups; n = 4 for Vec-ShSdc1 and C30-ShSdc1 groups; data are mean ± s.d.). h–m, iKras* p53L/+ tumour cells were infected with scrambled shRNA or shRNA against Sdcbp. h, Sdcbp knockdown was validated by western blot. Image of ShSdcbp-2 was cropped from the same membrane as Scr and ShSdcbp-1. Experiments were repeated twice with similar results. i, RAC1 activity was measured by G-LISA activation assay (n = 4 biological replicates; data are mean ± s.d.). j, Macropinocytosis was visualized with TMR–dextran (scale bar, 20 µm) and quantified (n = 5 random areas for Scr and ShSdcbp-2; n = 6 random areas for ShSdcbp-1 group; data are mean ± s.d.). Data are representative of two independent experiments with similar results. k, Representative images of clonogenic assay are shown from two independent experiments with similar results. l, Cells were subcutaneously injected into nude mice (n = 4 for Scr and ShSdcbp-1 groups, n = 5 for ShSdcbp-2 group) and tumour size was measured from photographs of dissected tumours shown in m. P values were determined by unpaired two-sided Student’s t-test.
Extended Data Fig. 10 SDC1 is critical for macropinocytosis in KRAS*-dependent human PDAC.
a–c, AsPC1 or PATC69 PDAC cells expressing mouse SDC1 or empty vector were infected with scrambled shRNA or shRNA against SDC1. Macropinocytosis was visualized with TMR–dextran (scale bar, 20 µm) (a) and quantified (b) (data are mean ± s.d.). Data are representative of two independent experiments with similar results. P values were determined by unpaired two-sided Student’s t-test. c, Surface SDC1 evaluated by FACS using human anti-SDC1 antibody DL101 or mouse anti-SDC1 antibody 281-2. Histograms show representative images from two independent experiments with similar results. d, Macropinocytic index of 20 human PDAC cell lines with high (n = 13) or low (n = 7) SDC1 membrane expression, as determined by FACS analysis. P value was determined by Mann–Whitney test. e, Macropinocytic index was quantified in 20 different PDAC cell lines (data are mean ± s.d.). f, Representative images of TMR–dextran (red) staining (scale bar, 20 µm). Data are representative of two independent experiments with similar results.
Supplementary information
Supplementary Figures
This file contains Supplementary Tables 2-4 and Supplementary Figures 1-2.
Supplementary Tables
This file contains Supplementary Tables 1-4.
Source data
Rights and permissions
About this article
Cite this article
Yao, W., Rose, J.L., Wang, W. et al. Syndecan 1 is a critical mediator of macropinocytosis in pancreatic cancer. Nature 568, 410–414 (2019). https://doi.org/10.1038/s41586-019-1062-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1062-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.