Abstract
The invasion of mammalian cytoplasm by microbial DNA from infectious pathogens or by self DNA from the nucleus or mitochondria represents a danger signal that alerts the host immune system1. Cyclic GMP–AMP synthase (cGAS) is a sensor of cytoplasmic DNA that activates the type-I interferon pathway2. On binding to DNA, cGAS is activated to catalyse the synthesis of cyclic GMP–AMP (cGAMP) from GTP and ATP3. cGAMP functions as a second messenger that binds to and activates stimulator of interferon genes (STING)3,4,5,6,7,8,9. STING then recruits and activates tank-binding kinase 1 (TBK1), which phosphorylates STING and the transcription factor IRF3 to induce type-I interferons and other cytokines10,11. However, how cGAMP-bound STING activates TBK1 and IRF3 is not understood. Here we present the cryo-electron microscopy structure of human TBK1 in complex with cGAMP-bound, full-length chicken STING. The structure reveals that the C-terminal tail of STING adopts a β-strand-like conformation and inserts into a groove between the kinase domain of one TBK1 subunit and the scaffold and dimerization domain of the second subunit in the TBK1 dimer. In this binding mode, the phosphorylation site Ser366 in the STING tail cannot reach the kinase-domain active site of bound TBK1, which suggests that STING phosphorylation by TBK1 requires the oligomerization of both proteins. Mutational analyses validate the interaction mode between TBK1 and STING and support a model in which high-order oligomerization of STING and TBK1, induced by cGAMP, leads to STING phosphorylation by TBK1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Li, T. & Chen, Z. J. The cGAS–cGAMP–STING pathway connects DNA damage to inflammation, senescence, and cancer. J. Exp. Med. 215, 1287–1299 (2018).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP–AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Wu, J. et al. Cyclic GMP–AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Saitoh, T. et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc. Natl Acad. Sci. USA 106, 20842–20846 (2009).
Jin, L. et al. MPYS, a novel membrane tetraspanner, is associated with major histocompatibility complex class II and mediates transduction of apoptotic signals. Mol. Cell. Biol. 28, 5014–5026 (2008).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011).
Zhang, X. et al. Cyclic GMP–AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 51, 226–235 (2013).
Cai, X., Chiu, Y. H. & Chen, Z. J. The cGAS–cGAMP–STING pathway of cytosolic DNA sensing and signaling. Mol. Cell 54, 289–296 (2014).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
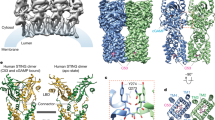
Shang, G., Zhang, C., Chen, Z. J., Bai, X.-c. & Zhang, X. Cryo-EM structures of STING reveal its mechanism of activation by cyclic GMP–AMP. Nature https://doi.org/10.1038/s41586-019-0998-5 (2019).
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Larabi, A. et al. Crystal structure and mechanism of activation of TANK-binding kinase 1. Cell Reports 3, 734–746 (2013).
Tu, D. et al. Structure and ubiquitination-dependent activation of TANK-binding kinase 1. Cell Reports 3, 747–758 (2013).
Shu, C. et al. Structural insights into the functions of TBK1 in innate antimicrobial immunity. Structure 21, 1137–1148 (2013).
Zhao, B. et al. Structural basis for concerted recruitment and activation of IRF-3 by innate immune adaptor proteins. Proc. Natl Acad. Sci. USA 113, E3403–E3412 (2016).
Lu, D. et al. Structural insights into the T6SS effector protein Tse3 and the Tse3-Tsi3 complex from Pseudomonas aeruginosa reveal a calcium-dependent membrane-binding mechanism. Mol. Microbiol. 92, 1092–1112 (2014).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Gouet, P., Courcelle, E., Stuart, D. I. & Metoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
Acknowledgements
We thank H. Yu and R. Hibbs for sharing instruments and reagents, and X. Tan for contributing to the analysis of some STING mutants. Cryo-EM data were collected at the University of Texas Southwestern Medical Center (UTSW) Cryo-Electron Microscopy Facility, which is funded by the Cancer Prevention and Research Institute of Texas (CPRIT) Core Facility Support Award RP170644. We thank D. Nicastro for facility access and data acquisition. This work is supported in part by the Howard Hughes Medical Institute (Z.J.C.), grants from the National Institutes of Health (GM088197 and R35GM130289 to X.Z), grants from the Welch foundation (I-1389 to Z.J.C.; I-1702 to X.Z; I-1944 to X.-c.B) and grants from CPRIT (RP150498 to Z.J.C.; RP160082 to X.-c.B.). X.-c.B. and X.Z. are Virginia Murchison Linthicum Scholars in Medical Research at UTSW. Z.J.C. is an investigator of the Howard Hughes Medical Institute.
Reviewer information
Nature thanks Andrea Ablasser, Philip Kranzusch and Osamu Nureki for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
All authors participated in research design, data analyses and manuscript preparation; G.S. and C.Z. prepared the protein samples for cryo-EM; X.-c.B., G.S. and X.Z. performed data acquisition, image processing, structure determination and analyses; C.Z. and X.G. did functional assays under the supervision of Z.J.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification of STING and TBK1, and characterization of their interaction.
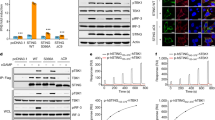
a, Binding between purified human STING and TBK1. Tsi3-tagged TBK1 was captured by Tse3-conjugated beads. Pull-down of STING by TBK1 was assessed by western blot. IB, immunoblotting; STING-FL, full-length STING; STING-Δtail, STING(1–343). b, Both human and chicken STING are able to induce phosphorylation of human TBK1 in cells. HeLa-C9 cells with undetectable endogenous STING were used to generate cell lines that stably expressed human STING–Flag or chicken STING–Flag. Cells were stimulated with cGAMP (1 μM) and analysed for TBK1 phosphorylation by immunoblotting. c, Gel filtration chromatography of the hybrid complex between chicken STING and human TBK1. Data are representative of two independent experiments.
Extended Data Figure 2 Flow chart of cryo-EM image processing for the complex between chicken STING and human TBK1.
a, Representative micrograph. b, Representative 2D classes of the intact complex. c, Representative 2D classes from TBK1-focused image processing. n > 3. d, f, Final reconstructions of the intact STING–TBK1 complex (d) and from the TBK1-focused refinement (f), with colours based on local resolution. e, g, Gold-standard FSC curves of the final 3D reconstructions of the intact complex and from the TBK1-focused refinement. h, Image-processing procedure.
Extended Data Fig. 3 Sample density maps.
Sample density maps are shown for the C-terminal tail of chicken STING and various parts of human TBK1.
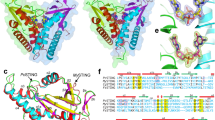
Extended Data Fig. 4 Structural comparison of apo TBK1 (PDB code 4IM0) and TBK1 bound to STING.
a, Overall TBK1 structures. b, A zoomed-in view of TBK1 regions bound to the STING C-terminal tail.
Extended Data Fig. 5 Sequence conservation of TBK1 from human and chicken.
Residues that are identical in the TBK1 of both species are coloured grey. Non-conserved residues are coloured red; non-identical, but similar, residues are coloured pink.
Extended Data Fig. 6 The binding and phosphorylation of STING by TBK1 relies on the interface between TBK1 and the STING C-terminal tail, and on the oligomerization of STING.
a, Mutations of TBK1-binding residues in the STING tail diminish cGAMP-induced phosphorylation of both TBK1 and STING. The S1 post-nuclear supernatant from HEK293T cells that expressed either the STING wild type or mutants was incubated with ATP in the presence or absence of cGAMP, and subjected to immunoblotting analyses for pTBK1, pSTING and STING. b, c, Mutations of TBK1-binding residues in the STING tail diminish cGAMP-induced STING phosphorylation (b) but not STING oligomerization (c). The same samples as in a were resolved by native gels, and analysed by immunoblotting. d–f, Mutations at the oligomerization interface of STING reduce cGAMP-induced oligomerization of STING, as well as phosphorylation of TBK1 and STING. The mutants are based on the accompanying paper on the structures of full-length STING12. The analyses in d, e and f were conducted in the same manner as those in a, b and c, respectively. Data shown here are representative of at least three independent biological replicates.
Extended Data Fig. 7 Cartoon model of STING-mediated activation of TBK1 and the downstream signalling pathway.
The cGAMP-induced oligomerization of STING leads to TBK1 clustering and trans-autophosphorylation. Activated TBK1 phosphorylates STING C-terminal tails that are not bound to the SDD–kinase domain groove in TBK1. Phosphorylated tails of STING recruit IRF3, which is phosphorylated by TBK1. Phosphorylated IRF3 forms a dimer and translocates to the nucleus to initiate the transcription of IFN genes.
Extended Data Fig. 8 Data collection and model statistics.
a, Data collection and model refinement statistics. b, FSC curves between the maps and model.
Supplementary information
Supplementary Information
This file contains Supplementary Results: Functional conservation between human and chicken STING.
Supplementary Figures
Supplementary Figure 1: Original uncropped images of gels or blots.
Video 1
Zoomed-in view of the interface between the STING C-terminal tail and TBK1.
Rights and permissions
About this article
Cite this article
Zhang, C., Shang, G., Gui, X. et al. Structural basis of STING binding with and phosphorylation by TBK1. Nature 567, 394–398 (2019). https://doi.org/10.1038/s41586-019-1000-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1000-2
This article is cited by
-
O-GlcNAc of STING mediates antiviral innate immunity
Cell Communication and Signaling (2024)
-
The combination of Tanshinone IIA and Astragaloside IV attenuates myocardial ischemia–reperfusion injury by inhibiting the STING pathway
Chinese Medicine (2024)
-
Nanomaterial-encapsulated STING agonists for immune modulation in cancer therapy
Biomarker Research (2024)
-
Universal STING mimic boosts antitumour immunity via preferential activation of tumour control signalling pathways
Nature Nanotechnology (2024)
-
IKKε and TBK1 prevent RIPK1 dependent and independent inflammation
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.