Abstract
The colonic epithelium facilitates host–microorganism interactions to control mucosal immunity, coordinate nutrient recycling and form a mucus barrier. Breakdown of the epithelial barrier underpins inflammatory bowel disease (IBD). However, the specific contributions of each epithelial-cell subtype to this process are unknown. Here we profile single colonic epithelial cells from patients with IBD and unaffected controls. We identify previously unknown cellular subtypes, including gradients of progenitor cells, colonocytes and goblet cells within intestinal crypts. At the top of the crypts, we find a previously unknown absorptive cell, expressing the proton channel OTOP2 and the satiety peptide uroguanylin, that senses pH and is dysregulated in inflammation and cancer. In IBD, we observe a positional remodelling of goblet cells that coincides with downregulation of WFDC2—an antiprotease molecule that we find to be expressed by goblet cells and that inhibits bacterial growth. In vivo, WFDC2 preserves the integrity of tight junctions between epithelial cells and prevents invasion by commensal bacteria and mucosal inflammation. We delineate markers and transcriptional states, identify a colonic epithelial cell and uncover fundamental determinants of barrier breakdown in IBD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw and processed sequencing data files are available under the GEO accession number GSE116222. The source code for analyses has been deposited at https://github.com/agneantanaviciute/colonicepithelium. Proteomics data have been deposited at the ProteomeXchange Consortium (http://www.proteomexchange.org) via the PRIDE69 partner repository with the dataset identifiers PXD011655 and 10.6019/PXD011655.
References
Peterson, L. W. & Artis, D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 14, 141–153 (2014)
McCauley, H. A. & Guasch, G. Three cheers for the goblet cell: maintaining homeostasis in mucosal epithelia. Trends Mol. Med. 21, 492–503 (2015).
Kabat, A. M., Pott, J. & Maloy, K. J. The mucosal immune system and its regulation by autophagy. Front. Immunol. 7, 240 (2016).
Hooper, K. M., Barlow, P. G., Henderson, P. & Stevens, C. Interactions between autophagy and the unfolded protein response: implications for inflammatory bowel disease. Inflamm. Bowel Dis. https://doi.org/10.1093/ibd/izy380 (2018).
Iyer, S. S. et al. Dietary and microbial oxazoles induce intestinal inflammation by modulating aryl hydrocarbon receptor responses. Cell 173, 1123–1134 (2018).
Rathinam, V. A. K. & Chan, F. K.-M. Inflammasome, inflammation, and tissue homeostasis. Trends Mol. Med. 24, 304–318 (2018).
McDole, J. R. et al. Goblet cells deliver luminal antigen to CD103+ dendritic cells in the small intestine. Nature 483, 345–349 (2012).
Johansson, M. E. & Hansson, G. C. Immunological aspects of intestinal mucus and mucins. Nat. Rev. Immunol. 16, 639–649 (2016).
Ayabe, T. et al. Secretion of microbicidal α-defensins by intestinal Paneth cells in response to bacteria. Nat. Immunol. 1, 113–118 (2000).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Ito, G. et al. Lineage-specific expression of bestrophin-2 and bestrophin-4 in human intestinal epithelial cells. PLoS ONE 8, e79693 (2013).
Tu, Y. H. et al. An evolutionarily conserved gene family encodes proton-selective ion channels. Science 359, 1047–1050 (2018).
Ikpa, P. T. et al. Guanylin and uroguanylin are produced by mouse intestinal epithelial cells of columnar and secretory lineage. Histochem. Cell Biol. 146, 445–455 (2016).
Sato, M. & Bremner, I. Oxygen free radicals and metallothionein. Free Radic. Biol. Med. 14, 325–337 (1993).
Gao, S. et al. Tracing the temporal-spatial transcriptome landscapes of the human fetal digestive tract using single-cell RNA-sequencing. Nat. Cell Biol. 20, 721–734 (2018); erratum 20, 1227 (2018).
Mojica, W. & Hawthorn, L. Normal colon epithelium: a dataset for the analysis of gene expression and alternative splicing events in colon disease. BMC Genomics 11, 5 (2010).
Chu, C. M. et al. Gene expression profiling of colorectal tumors and normal mucosa by microarrays meta-analysis using prediction analysis of microarray, artificial neural network, classification, and regression trees. Dis. Markers 2014, 634123 (2014).
Ding, L. et al. Claudin-7 indirectly regulates the integrin/FAK signaling pathway in human colon cancer tissue. J. Hum. Genet. 61, 711–720 (2016).
The Cancer Genome Atlas Research Network et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Vanhove, W. et al. Strong upregulation of AIM2 and IFI16 inflammasomes in the mucosa of patients with active inflammatory bowel disease. Inflamm. Bowel Dis. 21, 2673–2682 (2015).
Li, H. et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49, 708–718 (2017); correction 50, 1754 (2018).
Sasaki, N. et al. Reg4+ deep crypt secretory cells function as epithelial niche for Lgr5+ stem cells in colon. Proc. Natl Acad. Sci. USA 113, E5399–E5407 (2016).
Chen, C.-L., Yang, J., James, I. O. A., Zhang, H. Y. & Besner, G. E. Heparin-binding epidermal growth factor-like growth factor restores Wnt/β-catenin signaling in intestinal stem cells exposed to ischemia/reperfusion injury. Surgery 155, 1069–1080 (2014).
Slowikowski, K., Hu, X. & Raychaudhuri, S. SNPsea: an algorithm to identify cell types, tissues and pathways affected by risk loci. Bioinformatics 30, 2496–2497 (2014).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386 (2018).
Anderson, C. A. et al. Meta-analysis identifies 29 additional ulcerative colitis risk loci, increasing the number of confirmed associations to 47. Nat. Genet. 43, 246–252 (2011); erratum 43, 919 (2011).
Leiper, J. M. The DDAH-ADMA-NOS pathway. Ther. Drug Monit. 27, 744–746 (2005).
Ellinghaus, D. et al. Analysis of five chronic inflammatory diseases identifies 27 new associations and highlights disease-specific patterns at shared loci. Nat. Genet. 48, 510–518 (2016).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Spenlé, C. et al. Dysregulation of laminins in intestinal inflammation. Pathol. Biol. 60, 41–47 (2012).
Chhikara, N. et al. Human epididymis protein-4 (HE-4): a novel cross-class protease inhibitor. PLoS ONE 7, e47672 (2012).
Behrens, I., Stenberg, P., Artursson, P. & Kissel, T. Transport of lipophilic drug molecules in a new mucus-secreting cell culture model based on HT29-MTX cells. Pharm. Res. 18, 1138–1145 (2001).
O’Sullivan, S., Gilmer, J. F. & Medina, C. Matrix metalloproteinases in inflammatory bowel disease: an update. Mediators Inflamm. 2015, 964131 (2015).
Johansson, M. E. V. et al. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc. Natl Acad. Sci. USA 105, 15064–15069 (2008).
Porter, E. M., van Dam, E., Valore, E. V. & Ganz, T. Broad-spectrum antimicrobial activity of human intestinal defensin 5. Infect. Immun. 65, 2396–2401 (1997).
Cash, H. L., Whitham, C. V., Behrendt, C. L. & Hooper, L. V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313, 1126–1130 (2006).
Johansson, M. E. V., Larsson, J. M. H. & Hansson, G. C. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc. Natl Acad. Sci. USA 108 (Suppl 1), 4659–4665 (2011).
Waldman, S. A. & Camilleri, M. Guanylate cyclase-C as a therapeutic target in gastrointestinal disorders. Gut 67, 1543–1552 (2018).
Johansson, M. E. et al. Bacteria penetrate the normally impenetrable inner colon mucus layer in both murine colitis models and patients with ulcerative colitis. Gut 63, 281–291 (2014).
Johansson, M. E. Fast renewal of the distal colonic mucus layers by the surface goblet cells as measured by in vivo labeling of mucin glycoproteins. PLoS ONE 7, e41009 (2012).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171-181 (2014).
Sielaff, M. et al. Evaluation of FASP, SP3, and iST protocols for proteomic sample preparation in the low microgram range. J. Proteome Res. 16, 4060–4072 (2017).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Johansson, M. E. V. et al. Bacteria penetrate the inner mucus layer before inflammation in the dextran sulfate colitis model. PLoS ONE 5, e12238 (2010).
Skarnes, W. C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011).
Bradley, A. et al. The mammalian gene function resource: the International Knockout Mouse Consortium. Mamm. Genome 23, 580–586 (2012).
Pettitt, S. J. et al. Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat. Methods 6, 493–495 (2009).
Kojouharoff, G. et al. Neutralization of tumour necrosis factor (TNF) but not of IL-1 reduces inflammation in chronic dextran sulphate sodium-induced colitis in mice. Clin. Exp. Immunol. 107, 353–358 (1997).
Lesuffleur, T. et al. Differential expression of the human mucin genes MUC1 to MUC5 in relation to growth and differentiation of different mucus-secreting HT-29 cell subpopulations. J. Cell Sci. 106, 771–783 (1993).
Berger, G. et al. A simple, versatile and efficient method to genetically modify human monocyte-derived dendritic cells with HIV-1-derived lentiviral vectors. Nat. Protocols 6, 806–816 (2011).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Kuhn, R. M., Haussler, D. & Kent, W. J. The UCSC genome browser and associated tools. Brief. Bioinform. 14, 144–161 (2013).
Karolchik, D. et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 32, D493–D496 (2004).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Scialdone, A. et al. Computational assignment of cell-cycle stage from single-cell transcriptome data. Methods 85, 54–61 (2015).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Bacher, R. et al. SCnorm: robust normalization of single-cell RNA-seq data. Nat. Methods 14, 584–586 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
MacArthur, J. et al. The new NHGRI-EBI catalog of published genome-wide association studies (GWAS catalog). Nucleic Acids Res. 45 (D1), D896–D901 (2017).
The 1000 Genome Projects Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44 (D1), D447–D456 (2016).
Acknowledgements
We thank all of the patients who contributed to this study, our endoscopy teams and our clinical research nurses led by S. Fourie, who made this work possible. We acknowledge the support of the Wolfson Imaging Centre, the WIMM flow-cytometry facility, the Discovery Proteomics Facility, R. Dhaliwal for preparing TEM samples, the Oxford NIHR Biomedical Research Centre, the NIHR Clinical Research Network (CRN) Thames Valley, and the Oxford Single Cell Consortium. This work was supported by an NIHR Research Professorship and a Wellcome Investigator Award (to A.S.); the MRC (H.K. and A.S.); Abbvie (K.P.); and Celgene (A. Antanaviciute and H.H.C.). D.F.-C. was supported by a Royal College of Surgeons of England/British Association of Paediatric Surgeons Research Fellowship, an Oxford Wellcome Clinical Training Fellowship and by OHSRC, part of Oxford Hospitals charity. Further acknowledgements are given in the Supplementary Information.
Reviewer information
Nature thanks Richard Blumberg, Louis Vermeulen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
K.P., D.F-C., A. Antanaviciute and A.S. conceptualized the study. K.P., D.F-C. and M.J. performed and analysed experiments. A. Aulicino, H.H.C., N.A., S.D., J.L., R.S.A and E.B. performed wet laboratory experiments. C.L., N.K.A. and E.J. assisted with all microscopy-related experiments and analysis. R.G. and L.B. assisted with pathology and scoring. P.H. assisted with genetic experiments. A. Antanaviciute, H.K and J.K. performed computational analysis and design. S.D., R.F. and B.M.K. performed proteomic experiments. Writing and editing were carried out by K.P., D.F-C., A. Antanaviciute, H.K. and A.S. H.K. co-supervised and A.S. conceived the study, obtained funding and supervised.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification and validation of epithelial-cell subpopulations.
This figure is related to Fig. 1. a, i–iv, Flow-cytometry analysis of cells isolated from biopsies of healthy controls before scRNA-seq (i), measuring epithelial viability (DAPI−), purity (EPCAM+), an immune marker (CD45+) and a stromal marker(CD90+) (n = 4, mean ± s.d.), demonstrating the gating strategy for known epithelial markers (ii), viability (iii) and immune compartment (iv). APC and PE–Cy7 are fluorescent labels. b, i, FACS purification of EpCAM+CD45− isolated epithelial cells (n = 2). ii, iii, Representative images (n = 3) of immunohistochemical validation for LYZ expression in human epithelial tissue sections in small intestine (positive control) (ii) and colon (iii) (images shown at ×20 magnification). c, t-SNE plot of EEC subclusters. Single cells are coloured by cluster annotation. Descriptive cluster labels are shown (n = 3 per group). d, i, ii, Enteroendocrine subsets validated (representative images, n = 3) with double-stain immunohistochemistry for CHGA (blue) and two more novel markers identified from scRNA-seq: PCSK1N (i, brown) and CPE (ii, brown), showing co-localization of both markers in some cells (blue and brown arrowheads) but not in other EECs (blue or brown arrowhead). e, i–iii, Violin plots showing gene expression (y-axis) of different top EEC subcluster markers for enterochromaffin cells (ECs), L-cells (LCs) and a precursor-cell population (PCs) (n = 3; centre bar indicates median value; colour indicates mean expression). f, t-SNE plot visualizing undifferentiated colonic epithelial cell subclusters (n = 3). g, i–iii, Violin plots of gene expression (y-axis) in stem cells (SCs), cell-cycle cluster cells (CC), absorptive progenitor cells (APs), secretory progenitor cells (SPs) and transit-amplifying cells (TAs). The top markers for SCs (i), APs (ii) and SPs (iii) are shown (n = 3; centre bar indicates median value; colour indicates mean expression). h, Crypt-axis score superimposed over the differentiation trajectory captured by Monocle analysis (n = 3). Dimension (Dim) 1 and 2 are on the x- and y-axis respectively. i, Branch-specific expression of selected SC markers, secretory-lineage-specific markers and putative novel lineage-specific transcriptional regulators (n = 3). j, Selected Gene Ontology terms that show significant enrichment among all marker genes for epithelial clusters. The number of markers identified for each cluster is indicated (x-axis). The size of each circle corresponds to the proportion of markers annotated to a given term, while the colour indicates the significance (FDR) (n = 3 biological replicates; hypergeometric test and FDR calculated; Benjamini–Hochberg multiple testing correction).
Extended Data Fig. 2 Validation of the BEST4/OTOP2 cell population.
This figure is related to Fig. 2. a, Cluster distribution along the differentiation trajectory, captured by Monocle. BEST4/OTOP2 cells are highlighted on the left (n = 3). b, t-SNE gene expression overlay of core BEST4/OTOP2 cell markers (n = 3). c, i–v, Representative images (n = 3) of colonic sections stained with key BEST4/OTOP2 cell markers by immunohistochemistry to demonstrate BEST4 staining at high magnification (i; ×100), cathepsin E (CTSE) staining at low (ii; ×40) and high (iii) magnification, and additional stains with smISH for SPIB (iv) and HES4 (v) (each photograph is representative of three samples). d, i, t-SNE visualization of semisupervised clusters of scRNA-seq data identified in a fetal human colon study15 (n = 2). ii, Box plot (with 25th, 50th and 75th quantiles) showing co-localized expression of the core BEST4/OTOP2 cell signature. e, Heat map showing expression of the core BEST4/OTOP2 cell gene signature in TCGA bulk RNA-seq data in patients with colorectal cancer and matched normal tissue. f, i, t-SNE visualization of semisupervised clustering of scRNA-seq data from matched normal samples from a colorectal cancer study21 (n = 10). Only one BEST4/OTOP2 cell was identified in tumour samples (data not shown). ii, Box plot (with 25th, 50th and 75th quantiles) showing localized expression of the core BEST4/OTOP2 cell signature.
Extended Data Fig. 3 Isolation and characterization of the BEST4/OTOP2 cell population.
This figure is related to Fig. 2. a, Flow-cytometry gating strategy for isolating BEST4+ cells. Cells previously gated as live (DAPI−) singlets were selected as EPCAM+CD45− (i) with concurrent staining of a fluorescence minus one (ii) to allow placement of a BEST4+ gate on fully stained cells (iii). b, One hundred EPCAM+BEST4+ and EPCAM+BEST4− sorted cells (n = 3) processed using microfluidic RT–PCR demonstrate increased expression of markers identified from single-cell data relative to GAPDH. Mean ± s.e.m. values are shown. c, Circos plot showing overlap between the top 200 BEST4/OTOP2 cell markers detected between 10×, Smart-Seq2, quantitative proteomics and semi-supervised clustering of previously published data15,21. d, Over-represented Gene Ontology terms in the significantly upregulated protein set in BEST4/OTOP2 cells as identified by quantitative proteomics (n = 2 BEST4− versus n = 3 BEST4+; hypergeometric test and FDR-calculated Benjamini–Hochberg multiple testing correction).
Extended Data Fig. 4 Gene Ontology enrichment analysis of differentially expressed genes in colonic epithelial-cell clusters.
This figure is related to Figs. 3, 4. a, Dot plot of Gene Ontology biological-process enrichment in upregulated genes (less than 1% FDR), comparing cell clusters in active colitis and health. b, Dot plot of Gene Ontology biological-process enrichment in downregulated genes (less than 1% FDR), comparing cell clusters in active colitis and health. c, Dot plot of Gene Ontology biological-process enrichment in differentially expressed genes (less than 1% FDR) in inactive, but not in active, colitis. Points in each dot plot are coloured by enrichment confidence (−log10(FDR)) and sized by the proportion of all genes within the cluster annotated with the Gene Ontology term. For panels a–c, n = 3 per group; hypergeometric test and FDR-calculated Benjamini–Hochberg multiple testing correction. d, Violin plots showing expression (y-axis) of selected genes that are dysregulated in active colitis (‘inflamed’) compared with healthy samples (‘normal’) in stem cells and/or other undifferentiated populations. n = 3 per group; centre bar indicates median value; colour indicates mean expression. e, i–iv, Representative immunohistochemistry images (n = 3) showing LYZ expression in inflamed and noninflamed colonic tissue sections. i, iii, ×20 magnification; ii, iv, ×40 magnification.
Extended Data Fig. 5 Human colonic epithelium in clinically noninvolved mucosa and ulcerative-colitis-associated GWAS loci analysis.
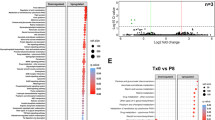
This figure is related to Fig. 3. a, Heat map visualizing the specificity of expression of ulcerative-colitis-associated GWAS loci in immune, epithelial and mesenchymal cell populations. Hierarchical clustering (horizontal) indicates groups of loci with similar expression specificities. b, t-SNE plots of cells in active colitis (n = 3), visualizing selected GWAS ulcerative-colitis-associated gene expression. c, Volcano plot showing the differentially expressed genes detected in a microarray study20, comparing inflamed ulcerative-colitis samples (n = 74) with healthy control colon samples (n = 11). Significantly downregulated core signature genes from BEST4/OTOP2 cells are highlighted (limma linear model empirical Bayes P value and Benjamini–Hochberg multiple testing correction). d, Distribution of cluster sizes in healthy colons (HC) and ulcerative-colitis inflamed (I) and noninflamed (NI) samples (n = 3 per group), shown as bar charts of proportions of total cells captured. Mean ± s.e.m. values are shown. e, t-SNE plot of human colonic epithelium single-cell clusters in noninflamed ulcerative colitis (n = 3).
Extended Data Fig. 6 Human colonic epithelium in clinically involved and noninvolved mucosa.
This figure is related to Fig. 3. a, i–iv, Violin plots visualizing expression (y-axis) of selected differentially expressed genes (less than 1% FDR; two-sided negative binomial likelihood ratio test; Benjamini–Hochberg multiple testing correction) in noninflamed and active ulcerative colitis (n = 3). Centre bar indicates median value; colour indicates mean expression. b, Heat map visualizing relative expression of all differentially expressed genes (less than 1% FDR; two-sided negative binomial likelihood ratio test; Benjamini–Hochberg multiple testing correction) detected in inflamed (red) and noninflamed (green) colitis compared with healthy tissue (blue) (n = 3 per group). c, Venn diagram showing the overlap between differentially expressed genes detected in all clusters in clinically inflamed (I) and clinically noninflamed (NI) colitis, compared with healthy tissue. d, Comparison between MAST generalized linear model coefficients for significant differentially expressed genes in ulcerative colitis inflamed and noninflamed samples with reference to healthy cells. Correlations for goblet and colonocyte cell clusters are shown (n = 3 per group; two-sided Hurdle likelihood ratio test; Benjamini–Hochberg multiple testing correction).
Extended Data Fig. 7 Goblet-cell remodelling and WFDC2 dysregulation in inflammation.
This figure is related to Figs. 4, 5. a, i–iv, Violin plots showing cluster gene expression (y-axis) for key marker genes in clusters 1 (i), 2 (ii) and 3 (iii) and common cluster 4 and 5 markers (iv) (n = 3 per group). Centre bar indicates median value; colour indicates mean expression. b, i, Pseudotemporal ordering of goblet-cell clusters. ii, Crypt-axis score superimposed on trajectory analysis. Cells predicted to reside at the top of the crypt are more mature populations, as inferred by pseudotime ordering, and vice versa. n = 3 per group. iii, Expression of MUC2 along the crypt axis. iv, Expression of WFDC2 along the crypt axis. c, i–vi, Gene-expression box plots of selected genes in goblet cells, divided spatially along the crypt axis by binning into four ranges (bottom, mid1, mid2 and top). n = 3 per group; 25th, 50th and 75th percentiles shown. Expression of CD74 (i), LCN2 (ii), REG1A (iii), SPINK1 (iv), SPINK4 (v) and LAMB3 (vi) is shown in health and inflamed ulcerative colitis. d, i–iv, Immunohistochemistry confirms increased expression of REG1A and SPINK4 in inflamed ulcerative-colitis biopsies (ii, iv) as compared with healthy samples (i, iii) (representative images of n = 3 for each). e, Stacked bar chart showing the relative frequency distribution of goblet-cell subclusters (percentage of goblet cells captured) in health and in active (inflamed) and inactive (noninflamed) colitis. f, Violin plots showing expression (y-axis) of WFDC2 in crypt-bottom goblet-cell clusters in healthy colons (HC) and inflammation (I) (n = 3 per group; centre bar indicates median value; colour indicates mean expression). g, Comparison of over-represented (hypergeometric test; Benjamini–Hochberg multiple testing correction) Gene Ontology biological-process terms amongst goblet-cell subcluster markers (n = 3 per group). h, Quantification of WFDC2 and MUC2 expression by immunohistochemistry from patient-matched inflamed and noninflamed sections of 24 patients with ulcerative colitis. Staining intensity was scored from 0 (no staining or weak staining) to 3 (strong staining) by three independent observers. Comparison between WFDC2 inflamed and noninflamed, P = 0.000148773; two-sided Wilcoxon matched-pairs signed-rank test, n = 24 patients. Comparison between MUC2 inflamed and noninflamed is not significant. Mean ± s.d. shown. i, Expression of interferon-induced genes in goblet cells (n = 3 per group): IFI6 (i), ISG15 (ii), IFITM3 (iii) and ISG20 (iv).
Extended Data Fig. 8 In vitro regulation of WFDC2.
This figure is related to Figs. 4, 5. a, i, ii, Untreated (i) and interferon-γ-treated (ii) human colonic organoids in culture. IFNg, interferon-γ. iii, qRT–PCR quantification of WFDC2 expression in interferon-γ-treated and untreated organoids (n = 2 independent experiments; mean values shown). iv, t-SNE plot of inflamed epithelium highlighting localized expression of interferon-γ in intra-epithelial lymphocytes (IELs; n = 3). b, i, ii, Quantification by enzyme-linked immunosorbent assay of WFDC2 secretion into apical (i) or basal (ii) medium of HT29-MTX-E12 cells with and without 100 ng ml−1 of PMA stimulation for 6 h (n = 1). c, i, ii, MMP12 (i) and MMP13 (ii) activity measured in the absence and presence of various concentrations of WFDC2. Data are presented as percentage of activity remaining. n = 3, except for MMP12 + 40 µg ml−1 WFDC2 and untreated MMP13, where n = 2. Mean ± s.d. shown. d, i–iii, WFDC2 knockdown (KD) in HT29-MTX-E12 cell lines (n = 2). i, Immunoblot of WFDC2 on cell lysates from nontransfected (lane 1, wild type, WT), WFDC2 shRNA transfected (lane 2, clone 1; lane 3, clone 2) and scrambled transfected (lane 4; Scr) cells. β-Actin was used as a loading control. I.B., immunoblot. ii, Cell-culture supernatants were tested by immunoblotting for secreted WFDC2. iii, Cells grown on transwells were stained with haematoxylin and eosin and Alcian blue. Arrows indicate the attached mucus layer and mucin-secreting goblet cells.
Extended Data Fig. 9 WFDC2 influences barrier function.
This figure is related to Fig. 5. a, i, ii, Histopathological evaluation of changes in epithelial-cell morphology and mucosal architecture in wild-type (WT; i) and Wfdc2+/− (ii) mice shows bifurcation at the base of the crypt in the Wfdc2+/− mice. iii, Mice were assigned a subjective colitis severity score on the basis of a modification of previously published criteria50. Scores for morphology, ulceration and infiltration were ranked on a scale from 0 (normal or absent) to 4 (severe), which were summed to give an overall score. b, Colonic tissue from Wfdc2+/− mice and wild-type littermates was processed to preserve the mucus layers. Immunohistochemistry for MUC2 in the distal mouse colon reveals mucus-filled goblet cells in the epithelium (e) and secreted mucus. The secreted mucus forms two layers: a stratified inner layer (i) and an outer layer (o). Arrows indicate the inner mucus layer. Higher-magnification images are shown in the bottom panels. n = 4. c, SEM of the colonic surface shows bacteria invading goblet cells in Wfdc2+/− mice. Scale bars, 2 µm. d, i–iv, TEM images of colons of Wfdc2+/− mice show epithelial-cell damage with destruction of microvilli (i), epithelial detachment (ii) and destruction (iii), as well as bacterial aggregates observed over the surface of Wfdc2+/− mice (iv). b–d show representative images; n = 4 animals per group.
Extended Data Fig. 10 Integrated sample analysis and batch distribution.
This figure relates to the Methods. a, Density distribution of cell UMI counts per sample. b, Density distribution of cellular gene-detection rate per condition. c, Density distribution of cellular gene-detection rate per condition per cell-type cluster. d, t-SNE visualization showing integrated clustering analysis of samples across all conditions (n = 3 per group). e, t-SNE visualization of sample batch distribution in the integrated clustering analysis (n = 3 per group). f, Box plots showing entropy of batch mixing for sample batches (n = 9; right); positive controls (Ct), in which clusters were assigned as batches (centre); and negative controls, in which cells were assigned random batch labels in accordance with batch size distribution (left). The entropy of batch mixing for sample batches approaches that of the negative control. Bars show the 25th, 50th and 75th percentiles.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4 and Supplementary Notes with further acknowledgements
Supplementary Data
This file contains Supplementary Data sheets – see contents page for details
Rights and permissions
About this article
Cite this article
Parikh, K., Antanaviciute, A., Fawkner-Corbett, D. et al. Colonic epithelial cell diversity in health and inflammatory bowel disease. Nature 567, 49–55 (2019). https://doi.org/10.1038/s41586-019-0992-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0992-y
This article is cited by
-
DHX9 maintains epithelial homeostasis by restraining R-loop-mediated genomic instability in intestinal stem cells
Nature Communications (2024)
-
Paneth-like cells produced from OLFM4+ stem cells support OLFM4+ stem cell growth in advanced colorectal cancer
Communications Biology (2024)
-
Notoginsenoside R1 promotes Lgr5+ stem cell and epithelium renovation in colitis mice via activating Wnt/β-Catenin signaling
Acta Pharmacologica Sinica (2024)
-
Single-cell RNA sequencing reveals the epithelial cell, fibroblast, and key gene alterations in chronic rhinosinusitis with nasal polyps
Scientific Reports (2024)
-
Tissue-location-specific transcription programs drive tumor dependencies in colon cancer
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.