Abstract
The GABAB (γ-aminobutyric acid type B) receptor is one of the principal inhibitory neurotransmitter receptors in the brain, and it signals through heterotrimeric G proteins to activate a variety of effectors, including G-protein-coupled inwardly rectifying potassium channels (GIRKs)1,2. GABAB-receptor signalling is tightly regulated by auxiliary subunits called KCTDs, which control the kinetics of GIRK activation and desensitization3,4,5. However, the mechanistic basis for KCTD modulation of GABAB signalling remains incompletely understood. Here, using a combination of X-ray crystallography, electron microscopy, and functional and biochemical experiments, we reveal the molecular details of KCTD binding to both GABAB receptors and G-protein βγ subunits. KCTDs associate with the receptor by forming an asymmetric pentameric ring around a region of the receptor carboxy-terminal tail, while a second KCTD domain, H1, engages in a symmetric interaction with five copies of Gβγ in which the G-protein subunits also interact directly with one another. We further show that KCTD binding to Gβγ is highly cooperative, defining a model in which KCTD proteins cooperatively strip G proteins from GIRK channels to induce rapid desensitization following receptor activation. These results provide a framework for understanding the molecular basis for the precise temporal control of GABAB signalling by KCTD proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The refined coordinates and structure factors for the KCTD16BTB/GABAB2 peptide complex and KCTD12H1/Gβ1γ2 complex have been deposited in the Protein Data Bank under the accession codes 6M8R and 6M8S, respectively.
References
Pin, J. P. & Bettler, B. Organization and functions of mGlu and GABAB receptor complexes. Nature 540, 60–68 (2016).
Froestl, W. Chemistry and pharmacology of GABAB receptor ligands. Adv. Pharmacol. 58, 19–62 (2010).
Schwenk, J. et al. Native GABAB receptors are heteromultimers with a family of auxiliary subunits. Nature 465, 231–235 (2010).
Turecek, R. et al. Auxiliary GABAB receptor subunits uncouple G protein βγ subunits from effector channels to induce desensitization. Neuron 82, 1032–1044 (2014).
Ivankova, K. et al. Up-regulation of GABAB receptor signaling by constitutive assembly with the K+ channel tetramerization domain-containing protein 12 (KCTD12). J. Biol. Chem. 288, 24848–24856 (2013).
Marshall, F. H., Jones, K. A., Kaupmann, K. & Bettler, B. GABAB receptors—the first 7TM heterodimers. Trends Pharmacol. Sci. 20, 396–399 (1999).
Geng, Y. et al. Structure and functional interaction of the extracellular domain of human GABAB receptor GBR2. Nat. Neurosci. 15, 970–978 (2012).
Geng, Y., Bush, M., Mosyak, L., Wang, F. & Fan, Q. R. Structural mechanism of ligand activation in human GABAB receptor. Nature 504, 254–259 (2013).
Bettler, B. & Tiao, J. Y. Molecular diversity, trafficking and subcellular localization of GABAB receptors. Pharmacol. Ther. 110, 533–543 (2006).
Gassmann, M. & Bettler, B. Regulation of neuronal GABAB receptor functions by subunit composition. Nat. Rev. Neurosci. 13, 380–394 (2012).
Padgett, C. L. & Slesinger, P. A. GABAB receptor coupling to G-proteins and ion channels. Adv. Pharmacol. 58, 123–147 (2010).
Huang, C. L., Slesinger, P. A., Casey, P. J., Jan, Y. N. & Jan, L. Y. Evidence that direct binding of Gβγ to the GIRK1 G protein-gated inwardly rectifying K+ channel is important for channel activation. Neuron 15, 1133–1143 (1995).
Correale, S. et al. A biophysical characterization of the folded domains of KCTD12: insights into interaction with the GABAB2 receptor. J. Mol. Recog. 26, 488–495 (2013).
Lee, M. T. et al. Genome-wide association study of bipolar I disorder in the Han Chinese population. Mol. Psychiatry 16, 548–556 (2011).
Metz, M., Gassmann, M., Fakler, B., Schaeren-Wiemers, N. & Bettler, B. Distribution of the auxiliary GABAB receptor subunits KCTD8, 12, 12b, and 16 in the mouse brain. J. Comp. Neurol. 519, 1435–1454 (2011).
Schwenk, J. et al. Modular composition and dynamics of native GABAB receptors identified by high-resolution proteomics. Nat. Neurosci. 19, 233–242 (2016).
Pinkas, D. M. et al. Structural complexity in the KCTD family of Cullin3-dependent E3 ubiquitin ligases. Biochem. J. 474, 3747–3761 (2017).
Seddik, R. et al. Opposite effects of KCTD subunit domains on GABAB receptor-mediated desensitization. J. Biol. Chem. 287, 39869–39877 (2012).
Whorton, M. R. & MacKinnon, R. X-ray structure of the mammalian GIRK2-βγ G-protein complex. Nature 498, 190–197 (2013).
Wang, W., Touhara, K. K., Weir, K., Bean, B. P. & MacKinnon, R. Cooperative regulation by G proteins and Na+ of neuronal GIRK2 K+ channels. eLife 5, e15751 (2016).
Yokogawa, M., Osawa, M., Takeuchi, K., Mase, Y. & Shimada, I. NMR analyses of the Gβγ binding and conformational rearrangements of the cytoplasmic pore of G protein-activated inwardly rectifying potassium channel 1 (GIRK1). J. Biol. Chem. 286, 2215–2223 (2011).
Sarvazyan, N. A., Remmers, A. E. & Neubig, R. R. Determinants of gi1α and βγ binding: measuring high affinity interactions in a lipid environment using flow cytometry. J. Biol. Chem. 273, 7934–7940 (1998).
Dror, R. O. et al. Structural basis for nucleotide exchange in heterotrimeric G proteins. Science 348, 1361–1365 (2015).
Zheng, S. & Ye, K. Purification, crystallization and preliminary X-ray diffraction analysis of Imp3 in complex with an Mpp10 peptide involved in yeast ribosome biogenesis. Acta Crystallogr. F 70, 918–921 (2014).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Lodowski, D. T., Pitcher, J. A., Capel, W. D., Lefkowitz, R. J. & Tesmer, J. J. Keeping G proteins at bay: a complex between G protein-coupled receptor kinase 2 and Gβγ. Science 300, 1256–1262 (2003).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Rees, I., Langley, E., Chiu, W. & Ludtke, S. J. EMEN2: an object oriented database and electronic lab notebook. Microsc. Microanal. 19, 1–10 (2013).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Acknowledgements
We thank K. Arnett and the Harvard Medical School Center for Macromolecular Interactions for assistance with biophysical measurements; M. Chambers at the Harvard Cryo-Electron Microscopy Center for electron microscopy training; and J. Liang and A. Manglik for providing purified Gαi protein. We also thank H. Schmidt for critical reading of the manuscript. Technical support for crystallographic software and computation was provided by SBGrid. Funding was provided by a Klingenstein–Simons Fellowship (A.C.K.), the Smith Family Foundation (A.C.K.), grant R35 GM124731 from the National Institute of General Medicine (J.L.) and a National Science Foundation (NSF) Graduate Research Fellowship (N.A.).
Reviewer information
Nature thanks Diomedes Logothetis and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.Z. performed protein purification, biochemical assays, and structural studies by crystallography and negative-stain electron microscopy, with supervision by A.C.K. Electrophysiology and microscopy experiments were performed by N.A. with supervision by J.L. The manuscript was written by S.Z. and A.C.K. with input from N.A. and J.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mapping the GABAB2-binding region of the KCTD BTB domain.
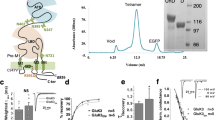
a, KCTD16BTB or KCTD12BTB domains tagged with His6–SMT3 (a SUMO protein) and GABAB2 fragments tagged with His6–GST were coexpressed in E. coli and purified with nickel affinity chromatography. Purified protein was loaded as input for pull-down with glutathione–sepharose beads. Glutathione input (IN) and pull-down (PD) fractions were analysed by SDS–PAGE and Coomassie blue staining. b, His6–GST-tagged GABAB2 fragments and untagged KCTD16BTB were coexpressed in E. coli. Clarified lysate was pulled down with glutathione–sepharose beads and eluate was analysed by SDS–PAGE and Coomassie blue staining. c, Structural superposition of the KCTD16BTB structure as bound to GABAB2 peptide (red; PDB identification 5A15) or free of peptide (grey). d, His6–SMT3-tagged wild-type or mutant (F80A) KCTD16BTB and His6–GST-tagged GABAB2 peptides were coexpressed and purified with nickel affinity. The eluate was treated with the protease ULP1 to cleave the SMT3 tag (IN) before pull-down (PD) using glutathione–sepharose beads. e, Sequence alignment of the BTB domains from human KCTD-family members. Residues with 98%, 80% and 60% similarity are shown in black, grey and light grey, respectively.
Extended Data Fig. 2 Electron-density maps.
a, Composite omit 2Fo − Fc electron-density map, contoured at 1.0σ, for a GABAB2 C-terminal peptide in complex with the KCTD16BTB domain. b, 2Fo − Fc electron-density map, contoured at 1.0σ, for the KCTD12H1 domain in complex with Gβ1γ2.
Extended Data Fig. 3 Representative negative-stain electron micrographs.
a, The full SDS–PAGE gel for the pull-down experiment shown in Fig. 2a. b, Gαi competes with KCTD12 for binding of Gβ1γ2. These pull-down experiments were carried out as for Fig. 2a, except that Gαi was used instead of Gαq. c, Isothermal titration calorimetry affinity measurements for the binding of KCTD12H1 to Gβ1γ2. d, Representative negative-stain electron-microscopy two-dimensional image of KCTD16BTB+H1 bound to the C-terminal GABAB2 domain. e, Representative negative-stain electron-microscopy two-dimensional image of KCTD12H1/Gβ1γ2. f, Representative negative-stain electron-microscopy two-dimensional image of KCTD12BTB+H1/GABAB2Cter/Gβ1γ2.
Extended Data Fig. 4 Three-dimensional negative-stain electron-microscopy reconstruction of full-length KCTD12 in complex with the GABAB2 C-terminal domain and Gβ1γ2.
a, Bottom, three different views of a three-dimensional reconstruction of full-length KCTD12 in complex with Gβ1γ2, with crystal structures of KCTD16BTB/GABAB2 peptide (red) and KCTD12H1/Gβ1γ2 (blue/purple) docked into the negative-stain electron-microscopy envelope. Top, negative-stain electron-microscopy two-dimensional class averages are reproduced from Fig. 2b for reference. b, Fourier shell correlation (FSC) curve as a function of spatial frequency for a negative-stain electron-microscopy map. Resolution is indicated below. c, Structure of the full-length KCTD12 complex. The BTB domain is separated from the H1 domain by 35 Å. Dashed lines represent the linker sequence between the BTB and H1 domains.
Extended Data Fig. 5 Structural and functional analysis of H1 domain.
a, Ribbon representation of the KCTD12H1 domain. The NFLEQ motif, which is important for desensitization, is coloured orange. b, Ribbon representation and topology map of the KCTD12 H1 monomer. Elements of the secondary structure are labelled. c, Ribbon representation of two KCTD12H1 subunits (green and cyan) and Gβ1γ2. d, Subcellular localization of eGFP–KCTD12(R232D) and eGFP–KCTD12(R257D) mutants with or without GABAB receptor.
Extended Data Fig. 6 Binding of Gβ1γ2 to KCTD12 is highly cooperative.
a, Ribbon representation of a complex of GIRK and Gβγ as seen from the cytosolic side. Four isolated Gβγ heterodimers associate with a single GIRK tetramer. b, Ribbon representation of a complex of KCTD12H1 and Gβγ as viewed from the cytosolic side. Gβγ subunits can be seen to interact directly with one another. c, Interactions between two adjacent Gβ subunits (labelled 1 and 2). d, 0.5/5, 1/5 or 5/5 molar ratios of Gβ1γ2 to KCTD12H1 were mixed and subjected to size-exclusion separation. The peaks correspond to 5/5 complexes and H1 domain alone. The x axis shows the elution volume (in ml). e, Left, a stoichiometric amount of Gβ1γ2 carrying a single mutation, R42D (in the Gβ1 subunit), or a double mutation, R42D/R46D, was incubated with KCTD12H1 and analysed by size-exclusion chromatography. Right, fractions from peak 1, peak 2 and peak 3 were analysed by SDS–PAGE and correspond to a 5/5 full complex, a partial complex and free Gβ1γ2, respectively.
Extended Data Fig. 7 A model for GABAB signalling and desensitization.
a, From left, binding of agonist (red star) to the GABAB1 receptor causes the Gβγ heterodimer to dissociate from Gα. Four copies of Gβγ bind to a GIRK channel tetramer, resulting in channel activation and an outflow of K+ ions. Afterwards, KCTD bound to the GABAB2 C terminus strips four copies of Gβγ from the GIRK channel and thereby deactivates the channel. Following nucleotide (GTP) hydrolysis by adenylyl cyclase, GDP-bound Gα binds again to Gβγ, sequestering it from KCTD and priming the system for another signalling cycle. b, Calculated total energy change for the series of progressively tighter binding events is commensurate with the approximate energy released by GTP hydrolysis.
Supplementary information
Rights and permissions
About this article
Cite this article
Zheng, S., Abreu, N., Levitz, J. et al. Structural basis for KCTD-mediated rapid desensitization of GABAB signalling. Nature 567, 127–131 (2019). https://doi.org/10.1038/s41586-019-0990-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0990-0
This article is cited by
-
Profiling the proximal proteome of the activated μ-opioid receptor
Nature Chemical Biology (2024)
-
Identifies KCTD5 as a novel cancer biomarker associated with programmed cell death and chemotherapy drug sensitivity
BMC Cancer (2023)
-
KCTD9 inhibits the Wnt/β-catenin pathway by decreasing the level of β-catenin in colorectal cancer
Cell Death & Disease (2022)
-
Ligand recognition and biased agonism of the D1 dopamine receptor
Nature Communications (2022)
-
The emerging role of the KCTD proteins in cancer
Cell Communication and Signaling (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.