Abstract
Dysregulation of lipid homeostasis is a precipitating event in the pathogenesis and progression of hepatosteatosis and metabolic syndrome. These conditions are highly prevalent in developed societies and currently have limited options for diagnostic and therapeutic intervention. Here, using a proteomic and lipidomic-wide systems genetic approach, we interrogated lipid regulatory networks in 107 genetically distinct mouse strains to reveal key insights into the control and network structure of mammalian lipid metabolism. These include the identification of plasma lipid signatures that predict pathological lipid abundance in the liver of mice and humans, defining subcellular localization and functionality of lipid-related proteins, and revealing functional protein and genetic variants that are predicted to modulate lipid abundance. Trans-omic analyses using these datasets facilitated the identification and validation of PSMD9 as a previously unknown lipid regulatory protein. Collectively, our study serves as a rich resource for probing mammalian lipid metabolism and provides opportunities for the discovery of therapeutic agents and biomarkers in the setting of hepatic lipotoxicity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All proteomics raw and processed data associated with the paper have been deposited in PRIDE proteomeXchange (http://www.proteomexchange.org/) using the following credentials: HMDP proteomics project accession = PXD010818 (reviewer username/password = reviewer09277@ebi.ac.uk/hyEjKh9x). ACAD11 AP–MS project accession = PXD010788 (reviewer username/password = reviewer14659@ebi.ac.uk/fg65Cfuf). Proteomics of PSMD9 ASO project accession PXD010759 (reviewer username/password = reviewer88344@ebi.ac.uk/7bATnY6j). All HMDP lipidomics raw acquisition files and associated documents are available at https://doi.org/10.6084/m9.figshare.7488854. Any remaining datasets are available in the Supplementary Information files or can be made available upon reasonable request to the corresponding author(s). All biological material will be made available upon reasonable request, with the exception of the ASOs, which are subject to a standing materials transfer agreement between the listed academic institutions and Ionis Therapeutics.
References
Younossi, Z. M. et al. Global epidemiology of nonalcoholic fatty liver disease-Meta-analytic assessment of prevalence, incidence, and outcomes. Hepatology 64, 73–84 (2016).
Sookoian, S. & Pirola, C. J. The genetic epidemiology of nonalcoholic fatty liver disease: toward a personalized medicine. Clin. Liver Dis. 16, 467–485 (2012).
Sookoian, S. & Pirola, C. J. Genetic predisposition in nonalcoholic fatty liver disease. Clin. Mol. Hepatol. 23, 1–12 (2017).
Mackay, T. F. et al. The Drosophila melanogaster Genetic Reference Panel. Nature 482, 173–178 (2012).
Peirce, J. L., Lu, L., Gu, J., Silver, L. M. & Williams, R. W. A new set of BXD recombinant inbred lines from advanced intercross populations in mice. BMC Genet. 5, 7 (2004).
Churchill, G. A. et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat. Genet. 36, 1133–1137 (2004).
Bennett, B. J. et al. A high-resolution association mapping panel for the dissection of complex traits in mice. Genome Res. 20, 281–290 (2010).
Churchill, G. A., Gatti, D. M., Munger, S. C. & Svenson, K. L. The Diversity Outbred mouse population. Mamm. Genome 23, 713–718 (2012).
Wu, Y. et al. Multilayered genetic and omics dissection of mitochondrial activity in a mouse reference population. Cell 158, 1415–1430 (2014).
Williams, E. G. et al. Systems proteomics of liver mitochondria function. Science 352, aad0189 (2016).
Chick, J. M. et al. Defining the consequences of genetic variation on a proteome-wide scale. Nature 534, 500–505 (2016).
Buscher, K. et al. Natural variation of macrophage activation as disease-relevant phenotype predictive of inflammation and cancer survival. Nat. Commun. 8, 16041 (2017).
Wang, J. J. et al. Genetic dissection of cardiac remodeling in an isoproterenol-induced heart failure mouse model. PLoS Genet. 12, e1006038 (2016).
Hui, S. T. et al. The genetic architecture of diet-induced hepatic fibrosis in mice. Hepatology 68, 2182–2196 (2018).
Jha, P. et al. Systems analyses reveal physiological roles and genetic regulators of liver lipid species. Cell Syst. 6, 722–733 (2018).
Jha, P. et al. Genetic regulation of plasma lipid species and their association with metabolic phenotypes. Cell Syst. 6, 709–721 (2018).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Alshehry, Z. H. et al. An efficient single phase method for the extraction of plasma Lipids. Metabolites 5, 389–403 (2015).
Meikle, P. J. & Summers, S. A. Sphingolipids and phospholipids in insulin resistance and related metabolic disorders. Nat. Rev. Endocrinol. 13, 79–91 (2017).
Samuel, V. T. & Shulman, G. I. Nonalcoholic fatty liver disease as a nexus of metabolic and hepatic diseases. Cell Metab. 27, 22–41 (2018).
Alshehry, Z. H. et al. Plasma lipidomic profiles improve on traditional risk factors for the prediction of cardiovascular events in type 2 diabetes mellitus. Circulation 134, 1637–1650 (2016).
Tonks, K. T. et al. Skeletal muscle and plasma lipidomic signatures of insulin resistance and overweight/obesity in humans. Obesity 24, 908–916 (2016).
Mundra, P. A. et al. Large-scale plasma lipidomic profiling identifies lipids that predict cardiovascular events in secondary prevention. JCI Insight 3, 121326 (2018).
Yang, P., Zhang, Z., Zhou, B. B. & Zomaya, A. Y. A clustering based hybrid system for biomarker selection and sample classification of mass spectrometry data. Neurocomputing 73, 2317–2331 (2010).
Peng, K. Y. et al. Mitochondrial dysfunction-related lipid changes occur in nonalcoholic fatty liver disease progression. J. Lipid Res. 59, 1977–1986 (2018).
Ryan, C. J., Kennedy, S., Bajrami, I., Matallanas, D. & Lord, C. J. A compendium of co-regulated protein complexes in breast cancer reveals collateral loss events. Cell Syst. 5, 399–409.e395 (2017).
Ruepp, A. et al. CORUM: the comprehensive resource of mammalian protein complexes—2009. Nucleic Acids Res. 38, D497–D501 (2010).
Huttlin, E. L. et al. Architecture of the human interactome defines protein communities and disease networks. Nature 545, 505–509 (2017).
Hein, M. Y. et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163, 712–723 (2015).
Das, J. & Yu, H. HINT: High-quality protein interactomes and their applications in understanding human disease. BMC Syst. Biol. 6, 92 (2012).
Chatr-Aryamontri, A. et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45 (D1), D369–D379 (2017).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Waterham, H. R., Ferdinandusse, S. & Wanders, R. J. Human disorders of peroxisome metabolism and biogenesis. Biochim. Biophys. Acta 1863, 922–933 (2016).
Van Veldhoven, P. P. Biochemistry and genetics of inherited disorders of peroxisomal fatty acid metabolism. J. Lipid Res. 51, 2863–2895 (2010).
Spanos, C. et al. Proteomic identification and characterization of hepatic glyoxalase 1 dysregulation in non-alcoholic fatty liver disease. Proteome Sci. 16, 4 (2018).
Mäkinen, V. P. et al. Integrative genomics reveals novel molecular pathways and gene networks for coronary artery disease. PLoS Genet. 10, e1004502 (2014).
Watanabe, T. K. et al. cDNA cloning and characterization of a human proteasomal modulator subunit, p27 (PSMD9). Genomics 50, 241–250 (1998).
Gragnoli, C. PSMD9 gene in the NIDDM2 locus is linked to type 2 diabetes in Italians. J. Cell. Physiol. 222, 265–267 (2010).
Gragnoli, C. Overweight condition and waist circumference and a candidate gene within the 12q24 locus. Cardiovasc. Diabetol. 12, 2 (2013).
Gragnoli, C. & Cronsell, J. PSMD9 gene variants within NIDDM2 may rarely contribute to type 2 diabetes. J. Cell. Physiol. 212, 568–571 (2007).
Parks, B. W. et al. Genetic control of obesity and gut microbiota composition in response to high-fat, high-sucrose diet in mice. Cell Metab. 17, 141–152 (2013).
Parks, B. W. et al. Genetic architecture of insulin resistance in the mouse. Cell Metab. 21, 334–347 (2015).
Palmisano, G. et al. Selective enrichment of sialic acid-containing glycopeptides using titanium dioxide chromatography with analysis by HILIC and mass spectrometry. Nat. Protocols 5, 1974–1982 (2010).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Bern, M., Kil, Y. J. & Becker, C. Byonic: advanced peptide and protein identification software. Curr. Protoc. Bioinformatics 40, 13.20.1–13.20.14 (2012).
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
Käll, L., Storey, J. D. & Noble, W. S. QVALITY: non-parametric estimation of q-values and posterior error probabilities. Bioinformatics 25, 964–966 (2009).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Huynh, K. et al. High-throughput plasma lipidomics: detailed mapping of the associations with cardiometabolic risk factors. Cell Chem. Biol. 26, 71–84.e4 (2018).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protocols 4, 44–57 (2009).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Breitkreutz, B. J., Stark, C. & Tyers, M. The GRID: the General Repository for Interaction Datasets. Genome Biol. 4, R23 (2003).
Ruepp, A. et al. CORUM: the comprehensive resource of mammalian protein complexes. Nucleic Acids Res. 36, D646–D650 (2008).
Ghazalpour, A. et al. Comparative analysis of proteome and transcriptome variation in mouse. PLoS Genet. 7, e1001393 (2011).
Norheim, F. et al. Genetic and hormonal control of hepatic steatosis in female and male mice. J. Lipid Res. 58, 178–187 (2017).
Pamir, N. et al. Proteomic analysis of HDL from inbred mouse strains implicates APOE associated with HDL in reduced cholesterol efflux capacity via the ABCA1 pathway. J. Lipid Res. 57, 246–257 (2016).
Drew, B. G. et al. HSP72 is a mitochondrial stress sensor critical for Parkin action, oxidative metabolism, and insulin sensitivity in skeletal muscle. Diabetes 63, 1488–1505 (2014).
Ribas, V. et al. Skeletal muscle action of estrogen receptor α is critical for the maintenance of mitochondrial function and metabolic homeostasis in females. Sci. Transl. Med. 8, 334ra54 (2016).
Drew, B. G. et al. Estrogen receptor (ER)α-regulated lipocalin 2 expression in adipose tissue links obesity with breast cancer progression. J. Biol. Chem. 290, 5566–5581 (2015).
de Aguiar Vallim, T. Q. et al. MAFG is a transcriptional repressor of bile acid synthesis and metabolism. Cell Metab. 21, 298–311 (2015).
Seth, P. P. et al. Synthesis and biophysical evaluation of 2′,4′-constrained 2′O-methoxyethyl and 2′,4′-constrained 2’O-ethyl nucleic acid analogues. J. Org. Chem. 75, 1569–1581 (2010).
Millard, P., Letisse, F., Sokol, S. & Portais, J. C. IsoCor: correcting MS data in isotope labeling experiments. Bioinformatics 28, 1294–1296 (2012).
Acknowledgements
This work was supported in part by the following: ANZ Victorian Medical Research Trust, Baker Heine Trust, Victorian State Government’s OIS Program and National Heart Foundation of Australia (A.C.C., B.G.D. and E.J.Z.); National Health and Medical Research Council of Australia (NHMRC) grants and fellowships (D.E.J. and B.L.P.); NIH grants HL122677, DK112119, DK102559 (T.Q.d.A.V.), HL028481 (A.J.L., T.Q.d.A.V.), HL118161 and HL136543 (E.J.T.); American Heart Association grant SDG18440015 (T.Q.d.A.V.). Baker Bright Sparks Scholarship and Australian Post-graduate Award (M.F.K.). We thank B. Crossett, S. Cordwell and The Sydney University Mass Spectrometry Facility. We are also grateful for the help and guidance of I. Carmichael and S. Bond and our many internal and external collaborators.
Reviewer information
Nature thanks Christa Buechler, Steven Munger and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
B.G.D., T.Q.d.A.V., A.C.C. and E.J.T. conceived the original concept. B.G.D., T.Q.d.A.V, A.C.C., B.L.P., M.M.S. and D.E.J. conceptualized the study and designed, performed, oversaw, interpreted and generated data and figures. B.L.P. and D.E.J. generated, analysed and interpreted the proteomic datasets. P.J.M. supervised the generation, analysis and interpretation of lipidomics data. A.J.L. advised on study design and systems genetics analysis, and provided access to data, software and reagents. B.G.D., B.L.P., T.Q.d.A.V., M.M.S., A.C.C., M.F.K., S.C.M., Y.L., E.J.Z., N.A.M., E.J.N., M.L.M., B.L.C., P.M., M.J.W., R.C.R.M., K.-Y.P., R. Lazarus and J.M.W. provided reagents, generated data and contributed to figure production. Specifically, B.L.C., P.M. and M.L.M. performed and analysed in vivo ASO experiments and data, and M.J.W., R.C.R.M. and K.-Y.P. performed and provided data for human plasma lipid signature validations. R. Lee provided access to, and expertise pertaining to ASO generation and delivery. M.M.S., B.L.P., B.G.D., K.J., C.P., R. Lazarus and P.Y. performed bioinformatics analyses. B.G.D., B.L.P., T.Q.d.A.V., A.C.C., M.M.S. and D.E.J. wrote the manuscript. All authors read and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R. Lee holds shares in Ionis Pharmaceuticals. B.G.D., A.C.C., T.Q.d.A.V. and D.E.J. are inventors on patent PCT/AU2019/050033 pertaining to aspects of the PSMD9 work. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Assessment of proteomic and lipidomic data reproducibility.
a, b, Coefficient of variation (CV) analysis of the proteomics (a) and lipidomics (b) data. Box-and-whisker plots (described as in Fig. 3e). c–e, Unsupervised hierarchical clustering of the liver proteomics (c), liver lipidomics (d) and plasma lipidomics (e) data.
Extended Data Fig. 2 Total liver triacylglycerol levels in C57BL/6J mice.
Mice were fed either a normal chow diet (NCD) or a high-fat diet (HFD) for 12 weeks. P value determined by Student’s t-test. Data are mean ± s.e.m., n = 11 chow group; n = 10 HFD group.
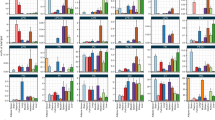
Extended Data Fig. 3 Average strain abundance of designated lipid classes in liver and plasma.
a–e, Abundance is expressed as area under the curve per mg liver protein or per ml of plasma. Liver scale on left, plasma scale on right a, Triacylglycerol. b, Diacylglycerol. c, Ceramide. d, Cholesterol esters. e, PE(P).
Extended Data Fig. 4 Correlation network analysis of the HMDP liver proteome.
a, Protein:protein (P:P) correlations in the HMDP liver proteome, integrated with CORUM-annotated proteins and protein interactions previously identified by AP–MS. Numbers indicate CORUM accessions, orange lines are HMDP P:P correlations; purple lines are correlations observed in both HMDP and CORUM. b–e, P:P correlations of selected CORUM complexes including associations not previously identified by AP–MS (green lines). Biweight midcorrelation analyses performed using ranked Benjamin–Hochberg multiple comparison test. Purple lines are known CORUM interactions, orange lines are HMDP P:P and CORUM interactions, green lines are previously unidentified interactions from HMDP P:P data. A thicker line represents a higher bicor value (q < 0.05, n > 50 strains).
Extended Data Fig. 5 Biweight midcorrelation of 108 liver lipid species against 378 liver proteins mapped onto annotated KEGG pathways.
Highlighted are various correlations (orange is positive, aqua is negative) between individual lipid species and proteins in pathways associated with unsaturated fatty acid metabolism, fatty acid degradation and metabolism, lysosomal degradation, and proteolysis. Only proteins containing more than one significant correlation to a lipid and annotated to the KEGG database are shown (biweight midcorrelation using ranked Benjamin–Hochberg multiple comparison test, q < 0.05, n > 50).
Extended Data Fig. 6 Overexpression of PSMD9 in C57BL/6J and DBA/2J mice.
Adenoviral overexpression of PSMD9 in C57BL/6J and DBA/2J mice (n = 9, 7 days after tail-vein injection of 109 plaque-forming units). a, b, Western blot (a) and densitometry (b) of PSMD9 and PDI (loading control) in the livers of mice treated with either control adenovirus (pAdV) or PSMD9 adenovirus. Data are mean ± s.e.m. c, Liver and plasma lipidomics of adenovirus-treated mice. Top panel (above first dotted line) shows relative fold change of total lipid classes. Middle and bottom panels show relative fold changes of individual diacylglycerol and triacylglycerol lipid species, respectively. P values determined by t-test with permutation-based FDR correction. Filled bubbles are significant (q < 0.05) changes, larger bubbles indicate greater significance.
Extended Data Fig. 7 ASO knockdown of PSMD9 in C57BL/6J and DBA/2J mice.
a–c, Assessment of hepatotoxicity as measured by plasma levels (U/L) of aspartate transaminase (AST) and alanine transaminase (ALT) (a), percentage of liver weight to body weight (b) and total body weight (c) of mice on a normal chow diet and treated with PBS, control ASO or PSMD9 ASO for 7 days (n = 4 per group, twice-weekly injection at 25 mg kg−1). d, Lipidomic analysis of total diacylglycerols and triacylglycerols in the plasma of mice on a chow diet (n = 4 C57BL/6J, n = 3 DBA/2J mice per group) or a Western diet (n = 6 mice per group, except n = 5 DBA/2J control ASO mice per group) and treated with either control or PSMD9 ASOs (twice-weekly ASO injection at 25 mg kg−1). e–g, Assessment of hepatotoxicity as measured by plasma AST and ALT levels (e), percentage change in body weight from baseline (f), and food consumption normalized to body weight (g) from in vivo de novo lipogenesis experimental animals (n = 6 control ASO, n = 8 PSMD9 ASO, 28 days on diet and weekly injection of ASO injection at 25 mg kg−1). *P < 0.05, **P < 0.01 control ASO versus PSMD9 ASO, t-test. Data are mean ± s.e.m.
Supplementary information
Supplementary Information
This file contains a guide for Supplementary Tables 1-16 and the uncropped blots for Figs. 3, 5 and Extended Data Fig. 6.
Supplementary Tables
This zipped file contains Supplementary Tables 1-16 – see Supplementary Information document for a full guide.
Rights and permissions
About this article
Cite this article
Parker, B.L., Calkin, A.C., Seldin, M.M. et al. An integrative systems genetic analysis of mammalian lipid metabolism. Nature 567, 187–193 (2019). https://doi.org/10.1038/s41586-019-0984-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0984-y
This article is cited by
-
Harnessing the power of proteomics in precision diabetes medicine
Diabetologia (2024)
-
Small molecule metabolites: discovery of biomarkers and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Mouse strain-dependent variation in metabolic associated fatty liver disease (MAFLD): a comprehensive resource tool for pre-clinical studies
Scientific Reports (2023)
-
Mechanisms Behind NAFLD: a System Genetics Perspective
Current Atherosclerosis Reports (2023)
-
The Possible Mechanism of Physiological Adaptation to the Low-Se Diet and Its Health Risk in the Traditional Endemic Areas of Keshan Diseases
Biological Trace Element Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.