Abstract
The expansion of brain size is accompanied by a relative enlargement of the subventricular zone during development. Epithelial-like neural stem cells divide in the ventricular zone at the ventricles of the embryonic brain, self-renew and generate basal progenitors1 that delaminate and settle in the subventricular zone in enlarged brain regions2. The length of time that cells stay in the subventricular zone is essential for controlling further amplification and fate determination. Here we show that the interphase centrosome protein AKNA has a key role in this process. AKNA localizes at the subdistal appendages of the mother centriole in specific subtypes of neural stem cells, and in almost all basal progenitors. This protein is necessary and sufficient to organize centrosomal microtubules, and promote their nucleation and growth. These features of AKNA are important for mediating the delamination process in the formation of the subventricular zone. Moreover, AKNA regulates the exit from the subventricular zone, which reveals the pivotal role of centrosomal microtubule organization in enabling cells to both enter and remain in the subventricular zone. The epithelial-to-mesenchymal transition is also regulated by AKNA in other epithelial cells, demonstrating its general importance for the control of cell delamination.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data generated and analysed in this study are included in the article and its Supplementary Information. Full gel blots can be found in the Extended Data Figures or Supplementary Fig. 1. Any other data in this study are available from the corresponding author upon reasonable request.
References
Taverna, E., Götz, M. & Huttner, W. B. The cell biology of neurogenesis: toward an understanding of the development and evolution of the neocortex. Annu. Rev. Cell Dev. Biol. 30, 465–502 (2014).
Fernández, V., Llinares-Benadero, C. & Borrell, V. Cerebral cortex expansion and folding: what have we learned? EMBO J. 35, 1021–1044 (2016).
Pinto, L. et al. Prospective isolation of functionally distinct radial glial subtypes—lineage and transcriptome analysis. Mol. Cell. Neurosci. 38, 15–42 (2008).
Pinto, L. et al. AP2γ regulates basal progenitor fate in a region- and layer-specific manner in the developing cortex. Nat. Neurosci. 12, 1229–1237 (2009).
Stahl, R. et al. Trnp1 regulates expansion and folding of the mammalian cerebral cortex by control of radial glial fate. Cell 153, 535–549 (2013).
Siddiqa, A. et al. Regulation of CD40 and CD40 ligand by the AT-hook transcription factor AKNA. Nature 410, 383–387 (2001).
Insolera, R., Bazzi, H., Shao, W., Anderson, K. V. & Shi, S. H. Cortical neurogenesis in the absence of centrioles. Nat. Neurosci. 17, 1528–1535 (2014).
Nigg, E. A. & Stearns, T. The centrosome cycle: centriole biogenesis, duplication and inherent asymmetries. Nat. Cell Biol. 13, 1154–1160 (2011).
Aprea, J. et al. Transcriptome sequencing during mouse brain development identifies long non-coding RNAs functionally involved in neurogenic commitment. EMBO J. 32, 3145–3160 (2013).
Uzbekov, R. & Alieva, I. Who are you, subdistal appendages of centriole? Open Biol. 8, 180062 (2018).
Thawani, A., Kadzik, R. S. & Petry, S. XMAP215 is a microtubule nucleation factor that functions synergistically with the γ-tubulin ring complex. Nat. Cell Biol. 20, 575–585 (2018).
Askham, J. M., Vaughan, K. T., Goodson, H. V. & Morrison, E. E. Evidence that an interaction between EB1 and p150Glued is required for the formation and maintenance of a radial microtubule array anchored at the centrosome. Mol. Biol. Cell 13, 3627–3645 (2002).
Ibi, M. et al. Trichoplein controls microtubule anchoring at the centrosome by binding to Odf2 and ninein. J. Cell Sci. 124, 857–864 (2011).
Tanaka, N., Meng, W., Nagae, S. & Takeichi, M. Nezha/CAMSAP3 and CAMSAP2 cooperate in epithelial-specific organization of noncentrosomal microtubules. Proc. Natl Acad. Sci. USA 109, 20029–20034 (2012).
Kasioulis, I., Das, R. M. & Storey, K. G. Inter-dependent apical microtubule and actin dynamics orchestrate centrosome retention and neuronal delamination. eLife 6, e26215 (2017).
Tsai, J. W., Lian, W. N., Kemal, S., Kriegstein, A. R. & Vallee, R. B. Kinesin 3 and cytoplasmic dynein mediate interkinetic nuclear migration in neural stem cells. Nat. Neurosci. 13, 1463–1471 (2010).
Itoh, Y. et al. Scratch regulates neuronal migration onset via an epithelial–mesenchymal transition-like mechanism. Nat. Neurosci. 16, 416–425 (2013).
Zander, M. A., Burns, S. E., Yang, G., Kaplan, D. R. & Miller, F. D. Snail coordinately regulates downstream pathways to control multiple aspects of mammalian neural precursor development. J. Neurosci. 34, 5164–5175 (2014).
Singh, S. et al. Zeb1 controls neuron differentiation and germinal zone exit by a mesenchymal–epithelial-like transition. eLife 5, e12717 (2016).
Sahu, S. K. et al. JNK-dependent gene regulatory circuitry governs mesenchymal fate. EMBO J. 34, 2162–2181 (2015).
Tiwari, N. et al. Sox4 is a master regulator of epithelial-mesenchymal transition by controlling Ezh2 expression and epigenetic reprogramming. Cancer Cell 23, 768–783 (2013).
Meng, W., Mushika, Y., Ichii, T. & Takeichi, M. Anchorage of microtubule minus ends to adherens junctions regulates epithelial cell–cell contacts. Cell 135, 948–959 (2008).
Cooper, J. A. Molecules and mechanisms that regulate multipolar migration in the intermediate zone. Front. Cell. Neurosci. 8, 386 (2014).
Petry, S. & Vale, R. D. Microtubule nucleation at the centrosome and beyond. Nat. Cell Biol. 17, 1089–1093 (2015).
Stiess, M. et al. Axon extension occurs independently of centrosomal microtubule nucleation. Science 327, 704–707 (2010).
Sakakibara, A. et al. Dynamics of centrosome translocation and microtubule organization in neocortical neurons during distinct modes of polarization. Cereb. Cortex 24, 1301–1310 (2014).
Aillaud, C. et al. Vasohibins/SVBP are tubulin carboxypeptidases (TCPs) that regulate neuron differentiation. Science 358, 1448–1453 (2017).
Song, Y. & Brady, S. T. Post-translational modifications of tubulin: pathways to functional diversity of microtubules. Trends Cell Biol. 25, 125–136 (2015).
Pongrakhananon, V., Wattanathamsan, O., Takeichi, M., Chetprayoon, P. & Chanvorachote, P. Loss of CAMSAP3 promotes EMT via the modification of microtubule–Akt machinery. J. Cell Sci. 131, jcs216168 (2018).
Chen, C., Lee, G. A., Pourmorady, A., Sock, E. & Donoghue, M. J. Orchestration of neuronal differentiation and progenitor pool expansion in the developing cortex by SoxC genes. J. Neurosci. 35, 10629–10642 (2015).
Sims-Mourtada, J. C. et al. The human AKNA gene expresses multiple transcripts and protein isoforms as a result of alternative promoter usage, splicing, and polyadenylation. DNA Cell Biol. 24, 325–338 (2005).
Martínez-Martínez, M. A. et al. A restricted period for formation of outer subventricular zone defined by Cdh1 and Trnp1 levels. Nat. Commun. 7, 11812 (2016).
de Juan Romero, C., Bruder, C., Tomasello, U., Sanz-Anquela, J. M. & Borrell, V. Discrete domains of gene expression in germinal layers distinguish the development of gyrencephaly. EMBO J. 34, 1859–1874 (2015)
Acknowledgements
We thank M. Takeichi (RIKEN CDB) for the Camsap3 antibody; C. Dehay and V. Cortay (SBRI) for immunostaining of macaque cortex; E. Llorens and T. Mata-Balaguer (IN) for ferret western blots; M. Drukker and E. Rusha (HMGU) for the iPSC line; I. Poser (MPI-CBG), C. Lie (University of Erlangen), U. Müller (Scripps Research Institute), A. Akhmanova (Utrecht University, EB3-GFP constructs) and G. Masserdotti for plasmids; A. Steiner for technical assistance; and S. Cappello (MPI-P) and A. O’Neill (HMGU) for comments on the manuscript. Funding was provided by the DFG (GO 640/12-1, SFB 870 A06 to M.G.; JU 2957/1-1, SFB 1032 A11 to R.J.; INST86/1581-1FUGG, IRTG 2290 to S.R.; SFB 1089 to F.B.), MINECO (SAF2015-69168-R to V.B.), Fundación Francisco Cobos (fellowship to C.D.J.R.), the ERC (Chroneurorepair to M.G.; Cortexfolding – 306933 to V.B.; MolMap – 680241 to R.J.), ERANET (AXON REPAIR and RATER SCI to F.B.; STEM-MCD and NEUROTALK to L.N.), the F.R.S.-FNRS (EOS O019118F-RG36 to L.N.), and NIH (R01DA024681 and R01NS085004 to S.-H.S.).
Reviewer information
Nature thanks Yukiko Gotoh, Xavier Morin and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.G. conceived and designed the project, and M.G. and P.A.J. started the project. P.A.J. and S.F. performed the in vivo experiments, which were also analysed by K.C. and A. Gavranovic. S.F., E.P. and L.B. performed and analysed all live imaging experiments. G.C.O. discovered the centrosome localization and cell-specific heterogeneity of Akna, and performed all biochemical and in vitro experiments. T. Steininger and T.K. contributed to the data analysis. K.D. and G.C.O. performed the human organoid experiments. M.W.-B. and W.B.H. contributed the immuno-electron microscopy experiments and analysis. A. Geerlof and R.F. generated the Akna antibodies, and G.C.O. validated them. J.M.-P. and S.M.H. contributed proteome expertise and analysis. S.V. and F.B. provided high-resolution light microscopy images, analysis of microtubule nucleation in NMuMG cells and expertise in centrosome function. C.D.J.R. and V.B. contributed the ferret experiments. W.S. and S.-H.S. provided samples of Sas4fl/flp53fl/flEmx1-cre mice. S.K.S. and G.C.O. performed experiments in NMuMG cells and analysed, with S.F., the role of Akna during EMT. S.R. and W.H. performed microtubule co-sedimentation experiments, and TIRF microscopy and analysis. R.J., T. Schlichthaerle and G.C.O. performed DNA-PAINT microscopy in E13 primary cortical cells. L.N. provided cell migration expertise, S.R. provided microtubule expertise, and V.K.T. provided EMT expertise. M.G., G.C.O. and S.F. wrote the manuscript, and all authors contributed corrections and comments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 AKNA expression, centrosome localization and antibody verification.
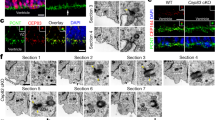
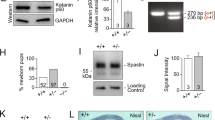
a, Quantitative PCR with reverse transcription (RT-qPCR) data show higher levels of Akna mRNA in E14 cerebral cortex than in E11 or E18 cerebral cortex (n = 3 biologically independent samples, mean ± s.e.m., one-way ANOVA with Tukey’s post hoc test. b, Microarray data depicting higher expression of AKNA in NSCs that generate basal progenitors (CD133+, hGFAP–GFPhigh (transgenic mice expressing GFP under control of the human GFAP promoter)3 (hGFAP–GFP, n = 4; hGFAP-GFPlow, n = 3 independent samples, mean ± s.e.m., two-sided Student’s t-test, note that all GFAP–GFP+ cells were double-stained for CD133 prior to FACS (see ref. 3). c, Western blot of AKNA in E14 cerebral cortex lysate running at a higher-than-predicted (153 kDa) molecular mass, owing to phosphorylation (data not shown, see also Extended Data Fig. 3d) (n = 8 independent experiments). d, e, Western blot of AKNA in Neuro2a cell lysates after transfection with Akna shRNA no. 1, Akna shRNA no. 2 or control shRNA (d, n = 2 independent experiments) or 3 different miRNAs and control (e, 1 experiment), using the antibody clone 14D7. f, AKNA immunofluorescence using clone 25F1 in primary E14 cortical cells, which show TBR2+ basal progenitors that lack AKNA immunofluorescence signal upon transfection with Akna siRNAs but not with control siRNA, demonstrating the specificity of the immunostaining (n = 3 independent experiments). g, Immunofluorescence of AKNA and TUBG in primary E14 cortical cells, showing AKNA signal surrounding a single TUBG+ centriole (n > 10 independent experiments). h, A20 BAC transgenic cell line showing GFP-tagged AKNA at centrosomes marked by TUBG in interphase (h′ and h′′′) but not during mitosis (h′′) (n = 3 independent experiments). i, Predicted domains of mouse AKNA protein. j, Amino acid sequence of AT-hook-containing transcription-factor domain of AKNA, and of the AT-hook-like domain. Note that AKNA lacks the GRP core sequence surrounded by several K and R amino acids, which is required for DNA- or RNA-binding AT-hook domains (such as the AT-hook domains of HMGA1). k–m, Immunostaining of dissociated cerebral organoid cells derived from human induced pluripotent stem cells (hiPSCs), showing AKNA localization at centrosomes with 3 different monoclonal antibodies (n = 2 independent experiments each). Clone 9G1 and 4F5 recognize epitopes in the N- and C-terminal parts of the protein, respectively, which suggests that different splice variants6,31 still localize at centrosomes. One (mother) centriole is enriched for AKNA. n, Western blot of AKNA in nuclear and cytoplasmic cell fractions of A20 cells, showing AKNA signal in the cytoplasm only (n = 2 independent experiments). o, Mass spectrometric analysis of sucrose-gradient-based isolated cellular sub-fractions of A20 cells. AKNA is enriched in fractions 3 and 4, which contain the centrosomal components listed on the right side (1 experiment). Scale bars, 5 μm (f, h, k–m), 2.5 μm (g).
Extended Data Fig. 2 Mechanisms of AKNA localization.
a, b, Electron microscopy micrographs showing AKNA immunogold-labelling at SDAs in the SVZ (a) and ventricular zone (b) of E13 cerebral cortex sections; b′, a magnification of the boxed area showing an SDA in b (n = 3 embryos). c, Stimulated emission depletion nanoscopy image showing AKNA immunofluorescence signal surrounding ODF2 immunofluorescence signal; this reveals the more-distal localization of AKNA at the SDA, as compared to ODF2, which localizes proximally with respect to the centriole in E14 brain NSCs (summarized in the schematic below; AKNA, green, ODF2, magenta) (n = 2 independent experiments). d, Micrographs showing DNA points accumulation for imaging in nanoscale topography super-resolution images of AKNA and TUBA, illustrating that AKNA is present along microtubules in E14 cortical cells (n = 3 independent experiments). e, Coomassie staining of a sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) gel showing samples of purified in vitro polymerized tubulin, with and without addition of purified recombinant AKNA as indicated (n = 2 independent experiments). Note that full-length AKNA co-precipitates with polymerized tubulin (lane 4), showing that AKNA is able to bind microtubules in vitro. f–h, E14 primary cortical cells treated with DMSO (f), nocodazole (g) or DCTN2 overexpression (h) show AKNA immunofluorescence remaining at centrosomes after microtubule depolymerization (n = 4 (f, g) and 3 (h) independent experiments). i, Micrographs showing AKNA immunofluorescence signal at centrosomes at the apical, ventricular surface in sections of E14 wild-type, but not Sas4−/−p53−/− (Sas4 is also known as Cenpji, and p53 is also known as Trp53) mice that lack centrioles7 (n = 2 independent experiments). Note that PCNT+ pericentriolar material is present in the absence of centrioles in Sas4−/−p53−/− mice. j, k, AKNA immunofluorescence of wild-type (j) or CRISPR–Cas9-generated Odf2 knock-out (k) mammary epithelial cells, showing that AKNA is lost from centrioles lacking SDAs (n = 3 independent experiments each). l, Schematic of different truncated AKNA variants used to analyse sub-cellular localization, as indicated to the right. m, n, Micrographs showing examples of the localization of truncated AKNA forms expressed in primary E14 cortical cells. Constructs containing the last 370 amino acids (n) localize to centrosomes, whereas AKNA(1–1080) is not localized at the centrosome (square) and is mainly cytoplasmic (m) (n = 3 independent experiments each). Notably, the clone used6 to first characterize AKNA lacked the C-terminal centrosome-targeting part. o, Representative images obtained from total internal reflection fluorescence (TIRF) time-lapse recordings of microtubules assembled in vitro from bovine brain tubulin, labelled with Cy5 in the presence of AKNA(1–1080)–GFP. BRB80 buffer with high salt concentration (100 mM KCl) was used to rule out unspecific binding to microtubules (n = 2 independent experiments). These data demonstrate that AKNA is an integral component of SDAs, is able to bind microtubules and is not recruited to centrosomes by microtubule or dynein–dynactin motors. Scale bars, 0.1 μm (a–c), 2 μm (d), 3 μm (o), 5 μm (f–h, m), 10 μm (i–k, n).
Extended Data Fig. 3 AKNA dissociates from the centrosome during mitosis and upon increased phosphorylation.
a, Immunofluorescence of AKNA in primary E14 cortical cells at different phases of the cell cycle, showing the lack of AKNA immunofluorescence at the centrosome during mitosis (n = 3 independent experiments). b, Western blot of AKNA in synchronized A20 cells, showing that AKNA protein is not degraded during mitosis (as indicated by the presence of phospho-histone H3) (n = 2 independent experiments). c, Representative micrographs of AKNA and PCNT immunofluorescence in E14 primary cortical cells treated with 500 nM okadaic acid (OA). Note that AKNA immunofluorescence is observed at centrosomes at 0 h but is undetectable in most cells 3 h after treatment, which shows that the centrosomal localization of AKNA is phosphorylation-dependent (n = 3 independent experiments). d, Western blot of lysates of OA-treated cells shows that phosphorylation—here caused by protein phosphatase inhibition—shifts that AKNA band on SDS–PAGE, and subsequently leads to protein degradation as observed in lysates of cells 5 h after OA washout (n = 3 independent experiments). Scale bars, 5 μm (a), 10 μm (c).
Extended Data Fig. 4 Temporal and sub-type-specific regulation of AKNA in the developing telencephalon.
a–c, Micrographs showing that PCNT+ centrosomes lack AKNA in the cerebral cortex at E9 (a, n = 2 experiments) and E18 (b, n = 2 experiments), whereas AKNA is enriched in the ventricular zone and—specifically—the SVZ at E14 in the ganglionic eminence (c, n > 10 experiments) and the cerebral cortex (Fig. 1). d, Distribution of AKNA+ centrosomes in the E14 cortex (n = 3 independent experiments). e–g, Plots showing the percentage of AKNA+ centrosomes in E14 and E13 cerebral cortex regions as indicated (e, f, n = 3 independent experiments), and in dissociated primary E14 cortical cells (g), which reveals that cells with AKNA+ centrosomes are mostly differentiating NSCs (PAX6+TBR2+) and PAX6−TBR2+ basal progenitors (n = 3 independent experiments). EM, electron microscopy; IF, immunofluorescence. h, Micrographs of cells isolated from E14 cerebral cortex by FACS using prominin 1, and stained for PAX6 (red arrows) and TBR2 (blue arrows). This shows that double-positive (differentiating) NSCs have AKNA+ centrosomes, whereas PAX6+TBR2− (proliferating) NSCs do not (n = 2 experiments). Data in d–g are presented as mean ± s.e.m. Scale bars, 10 μm (a, c, h), 20 μm (b).
Extended Data Fig. 5 AKNA knock down elicits cell death, and delamination defects persist upon cell-death rescue by p53 reduction.
a, Confocal micrographs showing staining of E14 cortex one day after co-IUE of membrane-tagged mKO2 (monomeric Kusabira-Orange2) and cytoplasmic GFP. Note that the vast majority of cells electroporated with one plasmid also express the marker of the second plasmid (n = 4 embryos). b, Line graph illustrating the distribution of GFP+ cells in the cerebral cortex after IUE for control shRNA (n = 6 embryos), Akna shRNA no. 1 (n = 5 embryos) and co-IUE of Akna shRNA and AKNA overexpression (0.2 μg μl−1) (n = 3 embryos). Note that the effect of AKNA knockdown is rescued with the appropriate amount of Akna expression, and is therefore specific. c–e, Micrographs showing TUNEL staining in E15 cerebral cortex, indicating cell death two days after IUE with Akna shRNA no. 1 (d, n = 3 embryos) or Akna shRNA no. 2 (d, n = 3 embryos) but not with control plasmids (c, n = 4 embryos). f, Dot and box plot showing the TUNEL+ area per electroporated (GFP+) area with control shRNA (n = 4 embryos) and Akna shRNA (n = 6 embryos, Akna shRNA no. 1 and Akna sRNA no. 2). g, h, Micrographs showing the distribution of GFP+ cells in E15 cerebral cortex two days after IUE with control shRNA (g) or Akna shRNA no. 2 plus p53 miRNA plasmids (h). Note that p53 downregulation rescues the apoptotic effect of AKNA knockdown. i, Line graph illustrating the distribution of GFP+ cells in the cerebral cortex after control shRNA (n = 6 embryos), p53 miRNA (n = 4 embryos), and Akna shRNA no. 2 and p53 miRNA (n = 4, embryos) IUE, showing the delamination defect that occurs upon AKNA knockdown is also present when apoptosis is blocked (Akna shRNA no. 2 and p53 miRNA). Note that p53 knockdown on its own does not alter the distribution of GFP+ cells. j, Dot and box plot showing the decrease of GFP+TBR2+ cells after IUE of Akna shRNA-positive, p53 miRNA, as compared to control; this shows that defects in delamination are accompanied by retaining an NSC fate (control shRNA, n = 6; Akna shRNA no. 2 and p53 miRNA, n = 8 embryos). k, Dot and box plot showing a decrease in proliferating (KI67+, n = 4 embryos each), and a concomitant increase in differentiated, NEUN+ cells analysed at E14 after IUE at E13 (GFP, n = 3; AKNA overexpression, n = 5 embryos). l, Box plot showing the fraction of PAX6+, KI67+ and NEUN+ in E14 primary cortical cells 48 h after transfection in vitro (n = 3 embryos each). b, i, Line graphs show mean ± s.e.m. as a transparent band in the same colour; control shRNA and Akna shRNA no. 1 data in b and control shRNA in i are the same data as shown in Fig. 1e. Box plots show median, quartiles (box) and range (whiskers); b, f, i–k, two-sided Mann–Whitney U test; l, two-sided Students t-test. Scale bars, 50 μm.
Extended Data Fig. 6 AKNA is sufficient to mediate microtubule polymerization and recruit γTuRC, CKAP5 and SDA components.
a–e, Micrographs of E14 primary cortex cells treated and immunostained as indicated. a, Time series depicting regrowth of the microtubule cytoskeleton after nocodazole-mediated depolymerization (n = 3 experiments). b, Regrowth of the microtubules 60 s after nocodazole-mediated depolymerization in AKNA overexpression (GFP+) cells (n > 10 experiments). Red arrows indicate centrosomes. Note that microtubules polymerize also from ectopic AKNA+ sites. c, Confocal images showing co-labelling of AKNA foci with the γTuRC components TUBG and GCP4, and microtubule nucleation factor CKAP5 (n = 2 experiments). d, Confocal micrographs illustrating that AKNA foci fail to recruit PCNT, NEDD1 and the microtubule minus-end capping-proteins CAMSAP2, CEP170 and CLASP2 (n = 2 experiments). e, Confocal micrographs depicting co-localization of ectopic AKNA foci with ODF2, DCTN1, EB1, dynein and CAMSAP3 (n = 2 experiments; 4 experiments for CAMSAP3). f, Co-immunoprecipitation experiments with lysates from E14 cerebral cortex, immunoprecipitation with AKNA antibody and western blot with AKNA, ODF2, DCTN1, EB1 AND GCP2, showing that these proteins are in the same complex, except GCP2 (that is, γTuRC) (n = 2 independent experiments for each). Scale bars, 5 μm (a–e).
Extended Data Fig. 7 AKNA effects on cilia formation or localization, the upstream regulation of Akna, and summary of AKNA effects.
a, b, Histograms depicting the percentages of ciliated cells (ARL13+) (a, n = 4 independent experiments) and short versus long cilia (b, n = 4 independent experiments), in E14 primary cortical cells transfected with control shRNA or Akna shRNA. Short cilia are defined as dot-like (short axoneme or cilium not fully exposed to cell surface), and long cilia are defined as rod-shaped (long axoneme exposed at the cell surface). c, d, Integrated correlative light and electron microscopy micrographs of in utero-electroporated cells (n = 2 embryos). The images compare two neighbouring NSCs; one electroporated (blue) and one non-electroporated (orange). Yellow and pink arrows show anti-GFP and anti-TUBA immunogold signal in the cytoplasm and cilium of the electroporated cell. Notice that AKNA electroporation does not notably affect cilia formation in vivo. e, Confocal micrographs of CAMSAP3, CDH2 (N-cadherin) and F-actin staining in E14 cortex. i, Model describing the expression and functional role of AKNA in delamination and seeding of the SVZ. j. Schematic indicating and summarizing the modes of AKNA function in NSC delamination. Data in a, b are presented as mean ± s.e.m. Scale bars, 30 μm (c, immunofluorescence), 1 μm (c, electron microscopy), 0.1 μm (d), 25 μm (e).
Extended Data Fig. 8 CAMSAP3 in the developing cortex, AKNA binding to and effects on purified microtubules, and AKNA regulation and function during EMT in mammary epithelial cells.
a–c, AKNA overexpression in E14 cortex influences both the orientation (a, b, control, 117 EB3-comets; AKNA overexpression, 122 EB3-comets in 3 experiments; plot shows the distribution and solid line indicates the mean, ± s.e.m. as transparent band) and speed (c, control, 178 EB3-comets; AKNA overexpression, 113 EB3-comets) of microtubule polymerization monitored by live imaging of EB3–GFP in cortical slices 1 day after IUE. d, Representative kymographs obtained from TIRF time-lapse images of microtubules assembled in vitro from bovine brain tubulin labelled with Cy5 in the presence of AKNA–GFP. Cy5 is shown in red and GFP in green. e–g, Dynamic instability parameters of microtubules assembled from bovine brain tubulin with and without AKNA. Tubulin concentration was 15 μM and AKNA concentration was 75 ng ml−1. In reactions containing AKNA, the growth velocity was increased. Moreover, the mean depolymerization velocity of shrinking plus ends without an AKNA signal was 0.27 μm s−1, and the mean velocity was 0.10 μm s−1 when AKNA was associated with the plus end (g). e, f, Microtubule, n = 117; microtubule and AKNA, 174; g, microtubule, n = 84; microtubule and AKNA, n = 47 single measurements made from 2 (microtubule) and 3 (microtubule and AKNA) independent experiments. h, Box plot showing marker expression 24 h after IUE of GFP (control) (n = 4 embryos), AKNA overexpression (n = 4 embryos) and AKNA overexpression supplemented with Taxol (20 μM) (n = 6 embryos). GFP (control) and AKNA overexpression is the same data as shown in Fig. 1j. Note that the addition of Taxol during IUE rescues the phenotype of AKNA overexpression. i, Bar graph illustrating the mRNA expression of Twist1, Zeb2, Snai1, Snai2 and Fn1 by qRT–PCR (n = 3 individual experiments) after TGFβ1 treatment and siRNA transfection (data are shown as mean ± s.e.m.). j, AKNA immunofluorescence in untreated and TGFβ1-treated (two-day treatment) NMuMG cells. Red arrows indicate AKNA− centrosomes (PCNT+) in untreated cells, and blue arrows indicate AKNA+ centrosomes in TGFβ1-treated cells. k, Western blot showing AKNA protein increase during the first days after EMT induction, and subsequent decrease to levels similar to those in untreated cells. Coomassie-stained gel showing equal loading is shown below. l, Dot and violin plot depicting the EB3–GFP fluorescence intensity measured at the centrosome in control (Centrin–tdTomato) and AKNA–mKO2 overexpression NMuMG cells, indicating the increased centrosomal microtubule nucleation after AKNA overexpression (GFP, 57 cells; AKNA overexpression, 56 cells). m, Western blot of AKNA and de-tyrosinated tubulin in untreated cells, and in cells transfected with control and Akna siRNA after four days of TGFβ1 treatment. Note the efficient knock down of AKNA by siRNA treatment, and the concomitant increase in detyrosinated tubulin. Whole protein detection illustrating equal protein loading is shown below. n, Western blot of TUBA and detyrosinated tubulin in control (GFP) and AKNA overexpression-transfected NMuMG cells 12 h after transfection. Note the decrease in detyrosinated tubulin upon AKNA overexpression. o, Micrograph illustrating how the quantification for ZO-1 was performed in p. p, Line graph depicting the quantification of ZO-1 intensity profiles across cells along white lines (see also Fig. 3a) (untreated, n = 60 cells; TGFβ1-treated, control siRNA, n = 55 cells; TGFβ1-treatment, Akna siRNA, n = 76 cells; cell junctions = position 0). *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 (exact P values (from left to right) 5.2 × 10−6, 0.00004 and 0.04177). q, Line graph depicting the quantification of CAMSAP3 intensity profiles across cells, as indicated in Fig. 3e (untransfected, n = 26 cells; AKNA overexpression, n = 22 cells). NS, not significant, ***P ≤ 0.001 (exact P values (from left to right) 0.53142 and 0.00003). r, Line graph depicting the quantification of CAMSAP3 intensity profiles across cells, as in Fig. 3e (untreated, n = 42 cells, TGFβ1-treated and control siRNA, n = 51 cells; TGFβ1-treated and Akna siRNA, n = 50 cells). *P ≤ 0.05, ***P ≤ 0.001 (exact P values (from left to right) 0.02478, 0.00038, 0.00095 and 0.00086). f–h, Box plots show median, quartiles (box) and range (whiskers). c, e, l, Violin plots show distribution of individual measurements (dots) with mean ± s.e.m. in bold in the centre (two-sided Mann–Whitney U test). Scale bars, 5 μm (j), 25 μm (a).
Extended Data Fig. 9 Lower levels of AKNA in differentiating neurons mediate non-centrosomal microtubule polymerization and allow migration into the cortical plate.
a, b, Primary E14 cortical cells were sorted for PROMININ 1 (CD133) to isolate NSCs or for PSA-NCAM to isolate neurons as indicated in the panels. PROM1+ cells express the NSC marker nestin, whereas PSA-NCAM+ cells express the neuronal marker TUBB3, showing the specificity of the FACS (a) (n = 2 independent experiments). Microtubule regrowth assay in purified NSCs shows centrosomal microtubule polymerization (b, top), and purified neurons show largely non-centrosomal microtubule polymerization patterns (b, bottom) (n = 2 independent experiments). c, Micrographs showing co-electroporation of a CAG–dsRED and DCX–GFP at E13 and analysis at E15. Note the onset of Dcx–GFP expression (c′′) only in the SVZ and cortical plate of the developing cortex, whereas DsRED+ cells are also found in the ventricular zone (c, c′) (n = 3 embryos). d, e, Micrographs showing electroporated cells (GFP+) in control (Dcx-GFP) (d) and Akna overexpressing (Dcx-Akna) (e) conditions (n = 3 embryos each conditions). Note that many AKNA overexpression cells accumulate in the SVZ, and are unable to migrate into the cortical plate. f, Micrographs of AKNA overexpression cells retained in the SVZ co-stained for TBR1, CTIP2, AND CUX1, which label neurons of different layer identity (n = 3 embryos). Note that the neuronal populations accumulating below the cortical plate upon Dcx-driven Akna expression contain neurons that are positive for each of these neuronal identities. g, Micrographs showing that electroporated cells (GFP+) after control and Dcx-driven miRNA mediated downregulation of AKNA enter the cortical plate equally well (n = 3 embryos). h, Line graph illustrating the distribution of GFP+ cells after IUE for Dcx-miRNA(neg), and Dcx-Akna miRNA no. 4. Mean ± s.e.m. as a transparent band (Dcx-miRNA(neg), n = 4; Dcx-Akna miRNA no. 4, n = 3 embryos). Scale bars, 5 μm (a, b), 50 μm (c–g).
Extended Data Fig. 10 AKNA localization in ferret and macaque cerebral cortex and function in human cerebral cortex organoids.
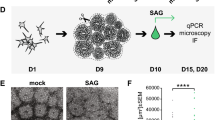
a, b, RT–qPCR of Akna and Sox4 in untreated or TGFβ1-treated NMuMG cells (treated for two days) showing upregulation during EMT (n = 4 independent experiments, mean ± s.e.m.). c, RT–qPCR of Akna after SOX4 overexpression in N2A cells for 36 h (n = 4 independent experiments, mean ± s.e.m.). d, qRT–PCR for Akna, showing the transient but strong upregulation of Akna mRNA in ferret ventricular zone tissue at E34—the time at which cells that form the outer SVZ (oSVZ) leave the ventricular zone32 (n = 4 (E30), 7 (E34), 4 (P1) biologically independent samples). e, f, Microarray33 (n = 3 biologically independent samples) (e) and western blot (f, 3 independent experiments) data showing higher levels of AKNA in the lateral sulcus (LS) compared to splenial gyrus (SG) in the oSVZ tissue of ferret brain at P1. g, Micrograph showing AKNA and TUBG co-localization at centrosomes in ferret brain cells (3 independent experiments). h, i, Comparison of AKNA immunofluorescence in ferret ventricular zone and oSVZ tissue in splenial gyrus versus lateral sulcus. Note the more-abundant immunofluorescence signal in the oSVZ of the lateral sulcus, corresponding with mRNA and protein levels as determined by western blot (n = 2 biologically independent samples). Given that the oSVZ of the gyrus contains more basal radial glia with bipolar morphology than does that of the sulcus, we propose that AKNA levels regulate the multipolar-to-bipolar transition in ferret SVZ, as AKNA levels do in mouse SVZ, with higher levels of AKNA leading to the retention of more cells in a multipolar state (see Fig. 4g–j). j, k, AKNA immunofluorescence in developing macaque germinal zone (GZ) and cortical plate (n = 2 sections from 2 animals). Arrows indicate AKNA+ centrosomes and denote abundant immunofluorescence signal. The square in j shows a representative example of a TUBG+AKNA+ centrosome in the germinal zone, and the square in (k) depicts an AKNA− centrosome in the cortical plate. l, Micrographs of human NPCs derived from hiPSCs that overexpress AKNA, showing multiple foci of microtubule polymerization (n = 2 independent experiments). m, n, In situ hybridization in hiPSC-derived cerebral organoids showing enrichment of Akna mRNA in non-apical SVZ-like areas (representative images of ventricular-zone-like structures from organoids of two independent culture batches). o, p, Micrographs showing sections of human brain organoids stained for GFP+ cells, electroporated with a control plasmid (o) or a plasmid overexpressing human AKNA cDNA (p). q, Line graph illustrating the distribution of GFP+ cells after IUE for GFP and AKNA overexpression (GFP, n = 36 ventricular zones; AKNA overexpression, n = 37 ventricular zones out of 2 independent organoid culture batches). r, Line graph illustrating the distribution of GFP+ electroporated cells after IUE for control miRNA and Akna miRNA knock down (control miRNA, n = 36 ventricular zones; Akna miRNA, n = 31 ventricular zones out of 2 independent organoid culture batches). q, r, Line graphs show mean ± s.e.m. as transparent band in the same colour. a–c, e, Two-sided Students t-test; d, one-way ANOVA followed by Tukey’s post hoc test; q, r, two-sided Mann–Whitney U test. Scale bars, 5 μm (g), 50 μm (h, i, m, n), 10 μm (l, j, k).
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion including detailed discussion of reasons for original mis-annotation of AKNA as transcription factor and species differences in the AT-hook sequence, Supplementary References, Supplementary Table 1, Supplementary Figures for FACS gating and the Supplementary Methods.
Supplementary Figure 1
The uncropped western blots with size-marker indications.
Supplementary Table 2
Mass-spec raw data of centrosome enriched fractions and protein clustering.
Video 1
High resolution light microscopy of AKNA, GT335 and ODF2 in an E14 primary cortical cell showing the 3+1 conformation of AKNA. The video exhibits AKNA (magenta) concentrated at the mother centriole (GT335 in green, adjacent to primary cilium). Observe two dots of AKNA at the same level of the DA/SDA marker ODF2 (white) and one at the PE of the MC. Moreover, an additional low intensity AKNA signal is also visible at the DC (n = 4 independent experiments).
Video 2
TIRF imaging of MTs with AKNA-mGFP protein. The recoding shows dynamic MTs in vitro (red) supplemented with AKNA-mGFP recombinant protein (green). Note (a) that AKNA decorates MTs after growth and (b) depolymerization stops when shrinking MTs meet an AKNA spot (n = 3 independent experiments).
Video 3
Time lapse imaging of AKNA-mKO2 in N2A cells. The video shows a transfected cell that undergoes mitosis. Notice how AKNA-mKO2 protein foci dissolve during mitosis, indicating that it is inactivated during this phase. At the end of M-phase it re-aggregates again (n = 4 independent experiments).
Video 4
Time lapse imaging of the cortical ventricular zone after Akna knockdown. The first part shows controls after in utero electroporation of a membrane-tagged EGFP (EGFP-f) at E13. The second part shows a movie where EGFP-f was co-electroporated with the Akna sh2 construct. Note that after Akna KD cells are retained in the VZ and SVZ and only few undergo interkinetic nuclear migration. Imaging started 23 hours after electroporation. Green arrows indicate cells in M-phase at the apical side (n = 3 independent experiments for each condition).
Video 5
Time lapse imaging of cells in the cortical ventricular zone after AKNA OE. The first part shows time lapse imaging of apically anchored control cells the ventricular zone after in utero electroporation of a membrane-tagged EGFP (EGFP-f) at E13. The cells undergo interkinetic nuclear migration and divide at the apical side. The second part shows a movie where EGFP-f was co-electroporated with the CAG driven AKNA overexpression construct and displays a cell which is initially apically anchored, rounding up at a non-apical position but then failing to undergo cytokinesis and turning into only one multipolar cell. The third part shows an apically anchored cell after co-electroporation with CAG-Akna that is delaminating from the apical side without rounding up or going through a division. In controls this is only very rarely observed. Imaging started 22 hours after electroporation (n = 3 independent experiments for each condition).
Video 6
Time lapse imaging of growing MTs in a NSC in the cortical ventricular zone in control conditions. EB3-GFP comets move from the apical site towards the basal process. As control RFP empty plasmid was used (n = 3 independent experiments).
Video 7
Time lapse imaging of growing MTs in a NSC in the cortical ventricular zone in AKNA OE conditions. EB3-GFP comets grow often from non-apical sites. Speed of MT nucleation is also faster than in controls. This indicates that elevating the levels of AKNA favor the re-orientation of MT nucleation and increases its speed. Overexpression was done with AKNA RFP (DsRED) plasmid (n = 3 independent experiments).
Video 8
Spinning disc imaging of MTs in NMUMG cells with CENTRIN OE as control. Untreated transgenic NMuMG EB3-NeonGreen epithelial cells transfected with CENTRIN-TdTOMATO (yellow arrow) showing that under control conditions MT organization is largely acentrosomal (n = 3 independent experiments). Scale bar: 5 µm.
Video 9
Spinning disc imaging of MTs in NMUMG cells with AKNA OE. Untreated transgenic NMuMG EB3-NeonGreen epithelial cells transfected with AKNA-mKO2 (yellow arrow) showing that MTs are mostly nucleated from the centrosome upon AKNA OE. Hence AKNA confers MTOC activity to the centrosome (n = 4 independent experiments). Scale bar: 5 µm.
Video 10
Time lapse imaging of newborn neurons migrating into the CP in control and AKNA OE conditions. In control conditions, two multipolar newborn neurons change to a bi-polar morphology. This is a critical step for migration into the CP. Upon AKNA OE, two multipolar newborn neurons are unable to change to a bi-polar morphology upon AKNA OE. These observations indicate that keeping the levels of AKNA high in young neurons impairs their repolarization and blocks their migration. OE was done with Dcx-driven plasmids. (n = 3 independent experiments).
Source data
Rights and permissions
About this article
Cite this article
Camargo Ortega, G., Falk, S., Johansson, P.A. et al. The centrosome protein AKNA regulates neurogenesis via microtubule organization. Nature 567, 113–117 (2019). https://doi.org/10.1038/s41586-019-0962-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0962-4
This article is cited by
-
Joint epigenome profiling reveals cell-type-specific gene regulatory programmes in human cortical organoids
Nature Cell Biology (2023)
-
Injury-specific factors in the cerebrospinal fluid regulate astrocyte plasticity in the human brain
Nature Medicine (2023)
-
Identification of Immune-Linked Hub Genes and Diagnostic Model Construction in Schizophrenia
Journal of Molecular Neuroscience (2023)
-
Upregulation of TUBG1 expression promotes hepatocellular carcinoma development
Medical Oncology (2023)
-
A complex epigenome-splicing crosstalk governs epithelial-to-mesenchymal transition in metastasis and brain development
Nature Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.