Abstract
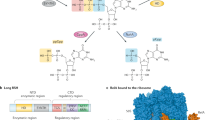
Cyclic dinucleotides (CDNs) have central roles in bacterial homeostasis and virulence by acting as nucleotide second messengers. Bacterial CDNs also elicit immune responses during infection when they are detected by pattern-recognition receptors in animal cells. Here we perform a systematic biochemical screen for bacterial signalling nucleotides and discover a large family of cGAS/DncV-like nucleotidyltransferases (CD-NTases) that use both purine and pyrimidine nucleotides to synthesize a diverse range of CDNs. A series of crystal structures establish CD-NTases as a structurally conserved family and reveal key contacts in the enzyme active-site lid that direct purine or pyrimidine selection. CD-NTase products are not restricted to CDNs and also include an unexpected class of cyclic trinucleotide compounds. Biochemical and cellular analyses of CD-NTase signalling nucleotides demonstrate that these cyclic di- and trinucleotides activate distinct host receptors and thus may modulate the interaction of both pathogens and commensal microbiota with their animal and plant hosts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the Article and associated Supplementary Information. X-ray crystallographic coordinates and structure factor files are available from the PDB: RmCdnE apo (6E0K); RmCdnE–Apcpp–Upnpp (6E0L); EmCdnE apo (6E0M); EmCdnE–GTP–Apcpp (6E0N); EmCdnE–pppA[3′–5′]pA (6E0O); RECON–cAAG (6M7K). CD-NTase sequences and information on CD-NTase-encoding bacteria are available in Supplementary Table 2 and CD-NTase effector genes sequences are available in Supplementary Table 3. CD-NTase alignments for tree construction are provided as Source Data for Fig. 4a and Source Data are also available for Extended Data Fig. 8b in the online version of the paper. Source gel images are available in Supplementary Fig. 1.
References
Wu, J. & Chen, Z. J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Corrales, L. et al. Direct activation of STING in the tumor microenvironment leads to potent and systemic tumor regression and immunity. Cell Rep. 11, 1018–1030 (2015).
Fu, J. et al. STING agonist formulated cancer vaccines can cure established tumors resistant to PD-1 blockade. Sci. Transl. Med. 7, 283ra52 (2015).
Ross, P. et al. Regulation of cellulose synthesis in Acetobacter xylinum by cyclic diguanylic acid. Nature 325, 279–281 (1987).
Danilchanka, O. & Mekalanos, J. J. Cyclic dinucleotides and the innate immune response. Cell 154, 962–970 (2013).
Nelson, J. W. & Breaker, R. R. The lost language of the RNA world. Sci. Signal. 10, eaam8812 (2017).
Krasteva, P. V. & Sondermann, H. Versatile modes of cellular regulation via cyclic dinucleotides. Nat. Chem. Biol. 13, 350–359 (2017).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP–AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Kranzusch, P. J. et al. Structure-guided reprogramming of human cGAS dinucleotide linkage specificity. Cell 158, 1011–1021 (2014).
Davies, B. W., Bogard, R. W., Young, T. S. & Mekalanos, J. J. Coordinated regulation of accessory genetic elements produces cyclic di-nucleotides for V. cholerae virulence. Cell 149, 358–370 (2012).
Hu, D. et al. Origins of the current seventh cholera pandemic. Proc. Natl Acad. Sci. USA 113, E7730–E7739 (2016).
Dziejman, M. et al. Comparative genomic analysis of Vibrio cholerae: genes that correlate with cholera endemic and pandemic disease. Proc. Natl Acad. Sci. USA 99, 1556–1561 (2002).
Severin, G. B. et al. Direct activation of a phospholipase by cyclic GMP–AMP in El Tor Vibrio cholerae. Proc. Natl Acad. Sci. USA 115, E6048–E6055 (2018).
Hornung, V., Hartmann, R., Ablasser, A. & Hopfner, K.-P. OAS proteins and cGAS: unifying concepts in sensing and responding to cytosolic nucleic acids. Nat. Rev. Immunol. 14, 521–528 (2014).
Jean, S. S., Lee, W. S., Chen, F. L., Ou, T. Y. & Hsueh, P. R. Elizabethkingia meningoseptica: an important emerging pathogen causing healthcare-associated infections. J. Hosp. Infect. 86, 244–249 (2014).
Jenal, U., Reinders, A. & Lori, C. Cyclic di-GMP: second messenger extraordinaire. Nat. Rev. Microbiol. 15, 271–284 (2017).
Corrigan, R. M. & Gründling, A. Cyclic di-AMP: another second messenger enters the fray. Nat. Rev. Microbiol. 11, 513–524 (2013).
Woodward, J. J., Iavarone, A. T. & Portnoy, D. A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705 (2010).
Wang, C. et al. Synthesis of all possible canonical (3′–5′-linked) cyclic dinucleotides and evaluation of riboswitch interactions and immune-stimulatory effects. J. Am. Chem. Soc. 139, 16154–16160 (2017).
McFarland, A. P. et al. Sensing of bacterial cyclic dinucleotides by the oxidoreductase RECON promotes NF-κB activation and shapes a proinflammatory antibacterial state. Immunity 46, 433–445 (2017).
Burroughs, A. M., Zhang, D., Schäffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Kazlauskiene, M., Kostiuk, G., Venclovas, Č., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR–Cas systems. Science 357, 605–609 (2017).
Niewoehner, O. et al. Type III CRISPR–Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017).
Hallberg, Z. F. et al. Hybrid promiscuous (Hypr) GGDEF enzymes produce cyclic AMP–GMP (3′, 3′-cGAMP). Proc. Natl Acad. Sci. USA 113, 1790–1795 (2016).
Nelson, J. W. et al. Control of bacterial exoelectrogenesis by c-AMP–GMP. Proc. Natl Acad. Sci. USA 112, 5389–5394 (2015).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Whiteley, A. T. et al. c-di-AMP modulates Listeria monocytogenes central metabolism to regulate growth, antibiotic resistance and osmoregulation. Mol. Microbiol. 104, 212–233 (2017).
Kranzusch, P. J. & Whelan, S. P. J. Arenavirus Z protein controls viral RNA synthesis by locking a polymerase–promoter complex. Proc. Natl Acad. Sci. USA 108, 19743–19748 (2011).
Zhou, W. et al. Structure of the human cGAS–DNA complex reveals enhanced control of immune surveillance. Cell 174, 300–311 (2018).
Kulasakara, H. et al. Analysis of Pseudomonas aeruginosa diguanylate cyclases and phosphodiesterases reveals a role for bis-(3′-5′)-cyclic-GMP in virulence. Proc. Natl Acad. Sci. USA 103, 2839–2844 (2006).
Schubert, S., Dufke, S., Sorsa, J. & Heesemann, J. A novel integrative and conjugative element (ICE) of Escherichia coli: the putative progenitor of the Yersinia high-pathogenicity island. Mol. Microbiol. 51, 837–848 (2004).
Kranzusch, P. J. et al. Ancient origin of cGAS–STING reveals mechanism of universal 2′,3′ cGAMP signaling. Mol. Cell 59, 891–903 (2015).
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
Reverter, D. & Lima, C. D. Structural basis for SENP2 protease interactions with SUMO precursors and conjugated substrates. Nat. Struct. Mol. Biol. 13, 1060–1068 (2006).
Stetson, D. B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Sureka, K. et al. The cyclic dinucleotide c-di-AMP is an allosteric regulator of metabolic enzyme function. Cell 158, 1389–1401 (2014).
Gaspar, A. H. & Machner, M. P. VipD is a Rab5-activated phospholipase A1 that protects Legionella pneumophila from endosomal fusion. Proc. Natl Acad. Sci. USA 111, 4560–4565 (2014).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Terwilliger, T. C. Reciprocal-space solvent flattening. Acta Crystallogr. D 55, 1863–1871 (1999).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Holm, L. & Laakso, L. M. Dali server update. Nucleic Acids Res. 44, W351–W355 (2016).
Lohöfener, J. et al. The activation mechanism of 2′-5′-oligoadenylate synthetase gives new insights into OAS/cGAS triggers of innate immunity. Structure 23, 851–862 (2015).
Yang, Q., Nausch, L. W. M., Martin, G., Keller, W. & Doublié, S. Crystal structure of human poly(A) polymerase gamma reveals a conserved catalytic core for canonical poly(A) polymerases. J. Mol. Biol. 426, 43–50 (2014).
Kuhn, C.-D., Wilusz, J. E., Zheng, Y., Beal, P. A. & Joshua-Tor, L. On-enzyme refolding permits small RNA and tRNA surveillance by the CCA-adding enzyme. Cell 160, 644–658 (2015).
Freudenthal, B. D., Beard, W. A., Shock, D. D. & Wilson, S. H. Observing a DNA polymerase choose right from wrong. Cell 154, 157–168 (2013).
Moon, A. F., Gosavi, R. A., Kunkel, T. A., Pedersen, L. C. & Bebenek, K. Creative template-dependent synthesis by human polymerase mu. Proc. Natl Acad. Sci. USA 112, E4530–E4536 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Kato, K., Ishii, R., Hirano, S., Ishitani, R. & Nureki, O. Structural basis for the catalytic mechanism of DncV, bacterial homolog of cyclic GMP–AMP synthase. Structure 23, 843–850 (2015).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP–AMP synthase. Cell 153, 1094–1107 (2013).
Acknowledgements
We thank K. McCarty and V. Cabrera for technical assistance; S. Wilson for advice; M. Reniere for reading the manuscript; T. Levin for input on CD-NTase tree construction; T. Wyche and M. Henke from the HMS ICCB Longwood Screening Facility and the staff at the HMS East Quad NMR Facility for their advice and technical support; A. Govande and W. Zhou for providing purified proteins; C. Miller for assistance with X-ray crystallography data collection; and the members of the Mekalanos and Kranzusch laboratories for advice and discussions. This work was funded by the Claudia Adams Barr Program for Innovative Cancer Research (P.J.K.), Richard and Susan Smith Family Foundation (P.J.K.), Charles H. Hood Foundation (P.J.K.), a Cancer Research Institute CLIP Grant (P.J.K.), NIH/NIAID R01AI018045 and R01AI026289 (J.J.M.), the Searle Scholars Program (A.S.Y.L.) and a Sloan Research Fellowship (A.S.Y.L.). A.T.W. is supported as a fellow of The Jane Coffin Childs Memorial Fund for Medical Research, C.C.d.O.M. is supported as a Cancer Research Institute/Eugene V. Weissman Fellow and B.R.M. is supported by the NIH T32 Cancer Immunology training grant (5T32CA207021-02). X-ray data were collected at the Lawrence Berkeley National Laboratory Advanced Light Source beamlines 8.2.1 and 8.2.2, this work used Northeastern Collaborative Access Team beamlines 24-ID-C and 24-ID-E (P30 GM124165), a Pilatus detector (S10RR029205), an Eiger detector (S10OD021527) and Argonne National Laboratory Advanced Photon Source (DE-AC02-06CH11357).
Reviewer information
Nature thanks Regine Hengge, Joshua Woodward and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Experiments were designed and conceived by A.T.W., J.J.M. and P.J.K. Gene identification, activity screening and bacterial assays were performed by A.T.W. Biochemical experiments were performed by A.T.W., J.B.E. and P.J.K. Structural experiments and analysis were performed by J.B.E. and P.J.K. Nucleotide synthesis and purification was performed by A.T.W., B.L. and P.J.K. Mass spectrometry was performed by C.C.d.O.M. and D.S.K. NMR was performed by B.R.M. Cloning assistance was provided by E.A.N. Observations and strains were contributed by O.D. Bioinformatics and cell assays were performed by A.S.Y.L. Figures were prepared by A.T.W., J.B.E. and P.J.K. The manuscript was written by A.T.W., J.B.E., J.J.M. and P.J.K. All authors contributed to editing the manuscript and support the conclusions.
Corresponding authors
Ethics declarations
Competing interests
Harvard Medical School and the Dana-Farber Cancer Institute have patents pending for CD-NTase technologies on which A.T.W., J.B.E., J.J.M. and P.J.K are listed as inventors.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Detailed characterization of CdnE, a cUMP–AMP synthase.
a, b, Titration of reaction buffer pH in steps of 0.2 pH units. Recombinant DncV or CdnE was incubated with α32P-radiolabelled ATP, CTP, GTP and UTP at varying pH and the reactions were treated with alkaline phosphatase and visualized by PEI–cellulose TLC. CdnE activity is optimal at a pH of approximately 9.4, and this reaction condition was used in further experiments. Data are representative of two independent experiments. c, PEI–cellulose TLC of products after incubation of the indicated enzyme, wild-type CdnE or active site mutant CdnE with α32P-radiolabelled ATP, CTP, GTP and UTP as in Fig. 1b. Mutations that ablate the CdnE Mg2+-coordinating, active-site residues eliminate all detectable activity. Data are representative of three independent experiments. d, Nuclease P1 sensitivity of CDN products. Nuclease P1 specifically cleaves 3′–5′, canonical phosphodiester bonds. DncV and CdnE products are completely digested in the presence of P1 and alkaline phosphatase, whereas only one bond of 2′,3′-cGAMP is susceptible to digestion, producing the linear G[2′–5′]pA product. Data are representative of three independent experiments. e, Workflow of nucleotide production for mass spectrometry and NMR analysis. f, Anion-exchange chromatography of a CdnE reaction with ATP and UTP, eluted with a gradient of buffer B (2 M ammonium acetate) by FPLC. Individual fractions were concentrated before pooling for further analysis. g, Anion-exchange chromatography (IEX) fractions from f were separated by silica TLC, visualized by ultraviolet-light shadowing and compared to a radiolabelled reaction to confirm the appropriate absorbance at 254 nm (A254) peak. Fractions were pooled and concentrated before mass spectrometry and NMR analysis. h, k, l, 3′,3′-Cyclic uridine monophosphate–adenosine monophosphate proton-NMR spectrum (h) and associated magnified spectra (k, l) 1H NMR (400 MHz): δΗ 8.40 (s, 1H), 8.17 (s, 1H), 7.90 (d, J = 8.2 Hz, 1H), 6.15 (s, 1H), 5.75 (s, 1H), 5.55 (d, J = 8.2 Hz, 1H), 5.00–4.90 (m, 2H), 4.80 (d, J = 4.5 Hz, 1H) 4.70–4.61 (m, 1H), 4.55–4.38 (m, 4H), 4.17–4.02 (m, 2H). i, j, 3′,3′-Cyclic uridine monophosphate–adenosine monophosphate phosphate-NMR spectrum (i) and associated magnified spectrum (j). 31P{1H} NMR (162 MHz): δP −1.59 (s, 1P), −1.65 (s, 1P).
Extended Data Fig. 2 Order of the DncV, cGAS and CdnE reactions.
a–c, Incubation of CD-NTase enzymes with non-hydrolysable nucleotides traps reaction intermediates and identifies the order of the reaction. Left, PEI–cellulose TLC analysis of reactions as in Fig. 1b, in which individual NTPs have been replaced by their non-hydrolysable nucleotide analogues. Exposed γ-phosphates are labelled in parentheses to indicate that these are removed by phosphatase treatment before analysis. Data are representative of three independent experiments. Right, previously determined reaction mechanisms (DncV and cGAS)10,51 and proposed reaction mechanism for CdnE.
Extended Data Fig. 3 Detailed analysis of RmCdnE.
a, A thermophilic homologue of CdnE from R. marinus (RmCdnE) also synthesizes cUMP–AMP. Recombinant proteins were incubated with α32P-radiolabelled NTPs as indicated at either 37 °C (CdnE) or 70 °C (RmCdnE) and the reactions were visualized by PEI–cellulose TLC as in Fig. 1b. Data are representative of two independent experiments. b, Active site of RmCdnE superimposed with structures of cGAS (6CTA30) and DncV (4TY010). c, The analogous position to Asn166 was mutated in CdnE to a serine and that protein, CdnEN166S, was characterized further. Reactions, which were analysed as in Fig. 1b, demonstrate that CdnEN166S loses pyrimidine specificity. Data are representative of two independent experiments. d, Structure-based sequence alignment of CD-NTases, annotated with secondary structure features of RmCdnE and human cGAS (6CTA30). Mg2+-coordinating active-site residues are highlighted in red; blue arrows, cyan/pink annotations and purple highlights represent secondary structure; and the analogous residues to RmCdnE Asn166 are highlighted in orange.
Extended Data Fig. 4 Detailed structural analysis of EmCdnE.
a, Sequence alignment of CdnE homologues in Fig. 2c, annotated with RmCdnE secondary structure features. Mg2+-coordinating active-site residues are highlighted in red and the analogous residues to RmCdnE Asn166 are highlighted in orange. Yersinia enterocolitica (WP_050915017); P. aeruginosa (WP_096075289); Xanthomonas arboricola (WP_104644370); Xenorhabdus nematophila (WP_010848498); Bordetella parapertussis (WP_015040391); Burkholderia cepacia complex (WP_006482377); R. marinus (RmCdnE, WP_014072508); L. pneumophila (WP_042646516); Mycobacterium avium (WP_062886322); E. meningoseptica (EmCdnE, WP_016200549); Staphylococcus aureus (WP_031901603); Enterococcus faecalis (WP_050492554); Bacteroides thetaiotaomicron (WP_062695386). b, Biochemical deconvolution of EmCdnE, which has a natural serine substitution at the Asn166 analogous site. Recombinant protein was incubated with NTPs as indicated and reactions were visualized as in Fig. 1b. Data are representative of three independent experiments. c, Reactions of EmCdnE incubated with α32P-radiolabelled NTPs and non-hydrolysable nucleotide analogues as indicated and visualized as in Fig. 1b. Data are representative of three independent experiments. d, Anion exchange chromatography of an EmCdnE reaction with ATP and GTP, eluted with a gradient of buffer B (2 M ammonium acetate) by FPLC. Individual fractions were concentrated before pooling for further analysis. e, Anion exchange chromatography fractions from d were separated by silica TLC, visualized by ultraviolet-light shadowing and compared to a radiolabelled reaction to confirm the appropriate peak. Fractions were pooled and concentrated before mass spectrometry analysis. Mass spectrometry confirmed synthesis of products with masses corresponding to c-di-AMP, cGAMP and c-di-GMP. f, Crystal structure of EmCdnE in complex with GTP and non-hydrolysable ATP capturing the ‘first state’ structure before NTP hydrolysis. Mg2+ ions are omitted for clarity. g, Magnified cut-away of the active site of the complex shown in f, confirming the position of a serine at the analogous site to RmCdnE Asn166. Nucleotide and metal 2Fo − Fc electron densities are contoured at 1σ. h, Magnified cut-away of the active site of the EmCdnE–pppApA structure, capturing the ‘second state’ after the first reaction has occurred to form a linear intermediate, but before CDN formation. Nucleotide and metal 2Fo – Fc electron densities are contoured at 1σ. g, h, Mg2+ ions are shown in green. i, Biochemical deconvolution of mutant EmCdnE reverted to the ancestral asparagine at the Asn166 analogous site. This mutant loses preference for producing cyclic dipurine molecules and instead produces more pyrimidine-containing CDN products. Reactions were visualized as in Fig. 1b. Data are representative of two independent experiments.
Extended Data Fig. 5 cUMP–AMP recognition helps to define innate immune receptor specificity.
a, e, Quantification of nucleotide interactions with the host receptors STING (a) or RECON (e), measured using a radiolabelled nucleotide bound to a concentration gradient of host protein, separated in a native PAGE gel shift (0, 4, 20, 100 μM protein). Quantification of the gels shown in b and f. Data are representative of n = 2 independent experiments. b, f, Native PAGE gel shift analysis of STING (b) or RECON (f) complex formation with indicated radiolabelled CDNs. Proteins are titrated at 0 (−), 4, 20 and 100 μM. STING readily binds all cyclic dipurine species, but does not form a high-affinity complex with cUMP–AMP. RECON readily binds all 3′,3′-CDN species that contain at least one adenine base, including cUMP–AMP. Data are representative of two independent experiments. c, In-cell STING reporter assay. Induction of an IFNβ reporter in HEK293T cells transfected with a concentration gradient of plasmid-overexpressing enzymes as indicated. DncV and CdnE were expressed with N-terminal MBP tags and IFNβ reporter induction was compared as fold over empty vector, shown as (−). Data are mean ± s.e.m. for n = 3 technical replicates and are representative of two independent experiments. d, Western blot of MBP-tagged DncV and CdnE expressed from plasmids analysed in c to validate in vivo expression. Data are representative of two independent experiments. Gel source data are available in Supplementary Fig. 1. g, Gel shift analysis as in f, with protein titration to measure the relative affinity of the RECON–cUMP–AMP interaction. Protein concentrations listed below. Data are representative of 2 independent experiments.
Extended Data Fig. 6 A biochemical screen of CD-NTases from diverse bacterial genera.
a, Chart of the number of bacterial genomes (n = 16,717) that have CD-NTases from clusters in Fig. 4a. See also Supplementary Table 2. b, Taxa of genome-sequenced bacteria isolated with unique CD-NTase genes, phyla are indicated in bold; Proteobacteria and Firmicutes are further divided by order and visualized by shades of colour. c–f, Type CD-NTases interrogated for product synthesis. Purified proteins were incubated with α32P-radiolabelled NTPs under different reaction conditions (indicated pH and divalent cation) and reaction products were visualized by either PEI–cellulose or silica TLC as in Fig. 1b and Fig. 4c, respectively. g, CD-NTase expression level and purity. Coomassie-stained SDS–PAGE analysis of purified CD-NTase enzymes used in each reaction. Data are representative of two independent experiments.
Extended Data Fig. 7 Detailed biochemical analysis of LpCdnE02.
a, Biochemical deconvolution of LpCdnE02 (CD-NTase057), analysed as in Fig. 1c, demonstrates specific synthesis of cyclic dipyrimidine products. Data are representative of three independent experiments. b, Nuclease sensitivity of the LpCdnE02 product, as described in Extended Data Fig. 1d. Data are representative of three independent experiments. c, Incubation of LpCdnE02 with non-hydrolysable nucleotides, as described in Extended Data Fig. 2. Non-hydrolysable UTP completely blocks the reaction, indicating the first step requires attack of the αP from UTP. However, the product formed when non-hydrolysable CTP is present cannot be distinguished from c-di-UMP in this assay, and it is unclear whether the reaction proceeds through a pppCpU reaction intermediate. Data are representative of three independent experiments. d, Anion exchange chromatography of a LpCdnE02 reaction with UTP and CTP, eluted with a gradient of buffer B (2 M ammonium acetate) by FPLC. Individual fractions were concentrated before pooling for further analysis. e, Anion exchange chromatography fractions from d were separated by silica TLC, visualized by ultraviolet-light shadowing and compared to a radiolabelled reaction to confirm the appropriate peak. Fractions were pooled and concentrated before mass spectrometry analysis. f, Mass spectrometry confirmed synthesis of c-di-UMP as the major product of LpCdnE02. g, Mass spectrometry confirmed synthesis of cCMP–UMP as a minor product of LpCdnE02.
Extended Data Fig. 8 CD-NTases are encoded in conserved, poorly understood operons on mobile genetic elements.
a, Operon structure for CD-NTases selected for in-depth characterization showing the conserved protein domains in CD-NTase adjacent genes (see Fig. 4a–c). Conserved operons were identified previously and operons are vertically organized by similarity to one another22. Where found, linked genes demonstrating that CD-NTases are encoded on mobile genetic elements are indicated. b, CD-NTases and their adjacently encoded ‘effector’ proteins were coexpressed in E. coli and bacterial colony formation was quantified by spot-dilution analysis. CD-NTases were inducibly expressed from a chloramphenicol-resistance (CmR) vector and effectors were inducibly expressed from a carbenicillin-resistance (CarbR) vector. Bacteria that express cognate CD-NTase–effector plasmids or control plasmids were plated on medium containing both inducers and incubated for 24 h at 37 °C. Data were not determined (N.D.) for CD-NTase036, because the effector was toxic to E. coli under non-inducing conditions. Data are the mean ± s.e.m. of three independent experiments. c, Spot-dilution analysis of bacteria expressing the indicated cognate CD-NTase–effector pairs, quantified as in b. The CD-NTase036–effector pair was not analysed in this assay. Colony morphology indicates a potential interaction for some combinations; however, it is unclear how specific or meaningful this may be. Data are representative of three independent experiments.
Extended Data Fig. 9 Detailed biochemical analysis of EcCdnD02.
a, Titration of reaction buffer pH in steps of 0.2 pH units. Recombinant EcCdnD02 was incubated with α32P-radiolabelled NTPs at varying pH and the reactions were analysed and visualized by PEI–cellulose or silica TLC as in Fig. 1b and Fig. 4c, respectively. Silica TLC identified two products, denoted the major (blue triangle) and minor (red triangle) product. Quantification of TLC spots is shown at the bottom. Data are representative of two independent experiments. b, Biochemical deconvolution of EcCdnD02, recombinant protein was incubated with NTPs as indicated and analysed by TLC as in a. Data are representative of three independent experiments. c, Nuclease digestion of the EcCdnD02 product. Conventional nuclease digestion includes addition of a phosphatase, in this experiment reactions were first treated with Antarctic phosphatase to remove remaining NTPs and were then heat-inactivated. Next, reactions were untreated, treated with nuclease P1 (specific for 3′–5′-phosphodiester bonds) only or treated with nuclease P1 and phosphatase to remove exposed phosphate groups. 3′,3′-cGAMP (DncV) and EcCdnD02 product are digested into AMP and GMP constituents, which are phosphatase-sensitive. cAMP (CyaA) is insensitive to P1 digestion and cyclic monophosphates are phosphatase-resistant. These data demonstrate that the EcCdnD02 product does not contain a cyclic monophosphate. Data are representative of three independent experiments. d, Incubation of EcCdnD02 with non-hydrolysable nucleotides, as described in Extended Data Fig. 2. Non-hydrolysable ATP completely blocks the reaction, indicating the first step requires attack of the αP from ATP. However, when non-hydrolysable GTP is present the possible intermediates (pp(c)pGpA, pp(c)pGpApA, or pppApA) cannot be distinguished in this assay and it is unclear how the reaction proceeds. Silica TLC is not suited for analysing non-hydrolysable nucleotides, because they do not migrate beyond the origin. Data are representative of three independent experiments. e, Anion exchange chromatography of a EcCdnD02 reaction with ATP and GTP, eluted with a gradient of buffer B (2 M ammonium acetate) by FPLC. Individual fractions were concentrated before pooling for further analysis. f, g, 3′,3′,3′-Tricyclic adenosine monophosphate–adenosine monophosphate–guanosine monophosphate phosphate-NMR spectrum (f) and associated magnified spectrum (g). 31P{1H} NMR (162 MHz): δP −0.65 (s, 1P), −0.70 (s, 1P), −0.75 (s, 1P). h–j, 3′,3′,3′-Tricyclic adenosine monophosphate–adenosine monophosphate–guanosine monophosphate proton-NMR spectrum (h) and associated magnified spectra (i, j). 1H NMR (400 MHz): δΗ 8.43 (s, 1H), 8.39 (s, 1H), 8.19 (s, 1H), 8.12 (s, 1H), 8.01 (s, 1H), 6.15 (d, J = 7.0 Hz, 1H), 6.12 (d, J = 7.0 Hz, 1H), 5.92 (d, J = 7.5 Hz, 1H), 5.00–4.78 (m, 6H), 4.69–4.58 (m, 3H), 4.3–4.2 (m, 6H).
Extended Data Fig. 10 Structural analysis of cAAG inhibition of RECON.
a, cAAG interactions with STING or RECON, radiolabelled nucleotides incubated with a concentration gradient of each protein, separated in a native PAGE gel shift (0, 4, 20, 100 μM protein). Data are representative of two independent experiments. b, Co-crystal structure of the RECON–cAAG complex shown as cartoon (left) and surface (right). c, Overlay and orientation of RECON ligands cAAG, c-di-AMP (PDB 5UXF), co-substrate NAD (PDB 3LN3) demonstrate three individual binding pockets. d, Schematic representation of residues from RECON that interact with cAAG. Green dotted lines indicate hydrogen bonds, grey dotted lines indicate hydrophobic interactions. e, Magnified cutaways of individual RECON binding pockets as in d. f, Mammalian innate-immune sensors recognize CD-NTase products with overlapping specificities. 2′,3′-cGAMP and c-di-GMP are detected by STING; 3′,3′-cGAMP and c-di-AMP are detected by both STING and RECON; and cUMP–AMP and cAAG are detected by RECON.
Supplementary information
Supplementary Information
This file contains a detailed discussion of how to identify CD-NTases and bacteria that encode a given CD-NTase of interest. It also includes Supplementary Table 1, a summary of data collection, phasing and refinement statistics, and Supplementary Figure 1, which includes original gel source data.
Supplementary Table 2
CD-NTases and CD-NTase encoding bacteria. Sequences of CD-NTases selected for screening and subsequent in-depth analysis, CD-NTases identified in bacteria, and bacteria predicted to encode a CD-NTase.
Supplementary Table 3
CD-NTase effector genes. Sequences of CD-NTase effector genes encoded adjacently to CD-NTases chosen for in-depth analysis.
Supplementary Data
CD-NTase alignment and tree. Alignment and tree of CD-NTase amino acid sequences from Supplementary Table 2. These provide the source data for Fig. 4a.
Source data
Rights and permissions
About this article
Cite this article
Whiteley, A.T., Eaglesham, J.B., de Oliveira Mann, C.C. et al. Bacterial cGAS-like enzymes synthesize diverse nucleotide signals. Nature 567, 194–199 (2019). https://doi.org/10.1038/s41586-019-0953-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0953-5
This article is cited by
-
Activation of CBASS Cap5 endonuclease immune effector by cyclic nucleotides
Nature Structural & Molecular Biology (2024)
-
Reversible conjugation of a CBASS nucleotide cyclase regulates bacterial immune response to phage infection
Nature Microbiology (2024)
-
Inhibitors of bacterial immune systems: discovery, mechanisms and applications
Nature Reviews Genetics (2024)
-
Conservation and similarity of bacterial and eukaryotic innate immunity
Nature Reviews Microbiology (2024)
-
Phages overcome bacterial immunity via diverse anti-defence proteins
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.