Abstract
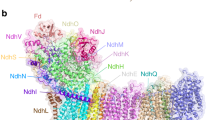
Cyclic electron flow around photosystem I (PSI) is a mechanism by which photosynthetic organisms balance the levels of ATP and NADPH necessary for efficient photosynthesis1,2. NAD(P)H dehydrogenase-like complex (NDH) is a key component of this pathway in most oxygenic photosynthetic organisms3,4 and is the last large photosynthetic membrane-protein complex for which the structure remains unknown. Related to the respiratory NADH dehydrogenase complex (complex I), NDH transfers electrons originating from PSI to the plastoquinone pool while pumping protons across the thylakoid membrane, thereby increasing the amount of ATP produced per NADP+ molecule reduced4,5. NDH possesses 11 of the 14 core complex I subunits, as well as several oxygenic-photosynthesis-specific (OPS) subunits that are conserved from cyanobacteria to plants3,6. However, the three core complex I subunits that are involved in accepting electrons from NAD(P)H are notably absent in NDH3,5,6, and it is therefore not clear how NDH acquires and transfers electrons to plastoquinone. It is proposed that the OPS subunits—specifically NdhS—enable NDH to accept electrons from its electron donor, ferredoxin3,4,5,7. Here we report a 3.1 Å structure of the 0.42-MDa NDH complex from the thermophilic cyanobacterium Thermosynechococcus elongatus BP-1, obtained by single-particle cryo-electron microscopy. Our maps reveal the structure and arrangement of the principal OPS subunits in the NDH complex, as well as an unexpected cofactor close to the plastoquinone-binding site in the peripheral arm. The location of the OPS subunits supports a role in electron transfer and defines two potential ferredoxin-binding sites at the apex of the peripheral arm. These results suggest that NDH could possess several electron transfer routes, which would serve to maximize plastoquinone reduction and avoid deleterious off-target chemistry of the semi-plastoquinone radical.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Cryo-EM maps for the dataset 1 overall, peripheral arm, with NdhS, without NdhS, with X-cofactor and without X-cofactor and dataset 2 overall have been deposited with the Electron Microscopy Data Bank under accession numbers EMD-0415, EMD-0416, EMD-0417, EMD-0418, EMD-0419, EMD-0420 and EMD-0425, respectively. Atomic coordinates for the overall complex for dataset 1, dataset 2 and composite have been deposited with the PDB under accession codes 6NBQ, 6NBX and 6NBY, respectively. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE62 partner repository with the dataset identifier PXD012206. All other data are available from the corresponding author upon reasonable request.
References
Kramer, D. M. & Evans, J. R. The importance of energy balance in improving photosynthetic productivity. Plant Physiol. 155, 70–78 (2011).
Arnon, D. I. The light reactions of photosynthesis. Proc. Natl Acad. Sci. USA 68, 2883–2892 (1971).
Shikanai, T. Chloroplast NDH: A different enzyme with a structure similar to that of respiratory NADH dehydrogenase. Biochim. Biophys. Acta 1857, 1015–1022 (2016).
Strand, D. D., Fisher, N. & Kramer, D. M. The higher plant plastid NAD(P)H dehydrogenase-like complex (NDH) is a high efficiency proton pump that increases ATP production by cyclic electron flow. J. Biol. Chem. 292, 11850–11860 (2017).
Yamamoto, H., Peng, L., Fukao, Y. & Shikanai, T. An Src homology 3 domain-like fold protein forms a ferredoxin binding site for the chloroplast NADH dehydrogenase-like complex in Arabidopsis. Plant Cell 23, 1480–1493 (2011).
Peltier, G., Aro, E.-M. & Shikanai, T. NDH-1 and NDH-2 plastoquinone reductases in oxygenic photosynthesis. Annu. Rev. Plant Biol. 67, 55–80 (2016).
He, Z. et al. NDH-1L interacts with ferredoxin via the subunit NdhS in Thermosynechococcus elongatus. Photosynth. Res. 126, 341–349 (2015).
Baradaran, R., Berrisford, J. M., Minhas, G. S. & Sazanov, L. A. Crystal structure of the entire respiratory complex I. Nature 494, 443–448 (2013).
Zickermann, V. et al. Mechanistic insight from the crystal structure of mitochondrial complex I. Science 347, 44–49 (2015).
Zhu, J., Vinothkumar, K. R. & Hirst, J. Structure of mammalian respiratory complex I. Nature 536, 354–358 (2016).
Chen, X., He, Z., Xu, M., Peng, L. & Mi, H. NdhV subunit regulates the activity of type-1 NAD(P)H dehydrogenase under high light conditions in cyanobacterium Synechocystis sp. PCC 6803. Sci. Rep. 6, 28361 (2016).
Fan, X., Zhang, J., Li, W. & Peng, L. The NdhV subunit is required to stabilize the chloroplast NADH dehydrogenase-like complex in Arabidopsis. Plant J. 82, 221–231 (2015).
Arteni, A. A. et al. Structural characterization of NDH-1 complexes of Thermosynechococcus elongatus by single particle electron microscopy. Biochim. Biophys. Acta 1757, 1469–1475 (2006).
Wulfhorst, H., Franken, L. E., Wessinghage, T., Boekema, E. J. & Nowaczyk, M. M. The 5 kDa protein NdhP is essential for stable NDH-1L assembly in Thermosynechococcus elongatus. PLoS ONE 9, e103584 (2014).
Wirth, C., Brandt, U., Hunte, C. & Zickermann, V. Structure and function of mitochondrial complex I. Biochim. Biophys. Acta 1857, 902–914 (2016).
Zhang, J. et al. NdhP is an exclusive subunit of large complex of NADPH dehydrogenase essential to stabilize the complex in Synechocystis sp. strain PCC 6803. J. Biol. Chem. 289, 18770–18781 (2014).
Shimizu, H. et al. CRR23/NdhL is a subunit of the chloroplast NAD(P)H dehydrogenase complex in Arabidopsis. Plant Cell Physiol. 49, 835–842 (2008).
Yamamoto, H. & Shikanai, T. In planta mutagenesis of Src homology 3 domain-like fold of NdhS, a ferredoxin-binding subunit of the chloroplast NADH dehydrogenase-like complex in Arabidopsis: a conserved Arg-193 plays a critical role in ferredoxin binding. J. Biol. Chem. 288, 36328–36337 (2013).
Kubota-Kawai, H. et al. X-ray structure of an asymmetrical trimeric ferredoxin–photosystem I complex. Nat. Plants 4, 218–224 (2018).
Veit, S. et al. The cyanobacterial cytochrome b 6 f subunit PetP adopts an SH3 fold in solution. Biochim. Biophys. Acta 1857, 705–714 (2016).
Kurisu, G. et al. Structure of the electron transfer complex between ferredoxin and ferredoxin-NADP+ reductase. Nat. Struct. Biol. 8, 117–121 (2001).
Srivastava, A. P. et al. Identification of the ferredoxin-binding site of a ferredoxin-dependent cyanobacterial nitrate reductase. Biochemistry 56, 5582–5592 (2017).
Dai, S. et al. Structural snapshots along the reaction pathway of ferredoxin-thioredoxin reductase. Nature 448, 92–96 (2007).
Ishikita, H. & Knapp, E.-W. Function of redox-active tyrosine in photosystem II. Biophys. J. 90, 3886–3896 (2006).
Ekberg, M., Sahlin, M., Eriksson, M. & Sjöberg, B. M. Two conserved tyrosine residues in protein R1 participate in an intermolecular electron transfer in ribonucleotide reductase. J. Biol. Chem. 271, 20655–20659 (1996).
Zhang, P. et al. Isolation, subunit composition and interaction of the NDH-1 complexes from Thermosynechococcus elongatus BP-1. Biochem. J. 390, 513–520 (2005).
Kern, J. et al. Purification, characterisation and crystallisation of photosystem II from Thermosynechococcus elongatus cultivated in a new type of photobioreactor. Biochim. Biophys. Acta 1706, 147–157 (2005).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Wittig, I., Braun, H.-P. & Schägger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Wessel, D. & Flügge, U. I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 138, 141–143 (1984).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Beck, S. et al. The impact II, a very high-resolution quadrupole time-of-flight instrument for deep shotgun proteomics. Mol. Cell Proteomics 14, 2014–2029 (2015).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Biyani, N. et al. Focus: The interface between data collection and data processing in cryo-EM. J. Struct. Biol. 198, 124–133 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Kimanius, D., Forsberg, B. O., Scheres, S. H. W. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bai, X.-C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. W. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zivanov, J., Nakane, T. & Scheres, S. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Scheres, S. H. W. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
McGuffin, L. J., Bryson, K. & Jones, D. T. The PSIPRED protein structure prediction server. Bioinformatics 16, 404–405 (2000).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
The PyMOL Molecular Graphics System v.1.8.2 (Schrödinger, 2016).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
We thank the Yano/Yanchandra Laboratory for T. elongatus membranes; E. A. Montabana for cryo-EM training of T.G.L. and advice on sample preparation; D. Toso for assistance during data collection; P. Tobias for technical support; T. H. D. Nyugen and B. J. Greber for advice on model-building; and L. M. Oltrogge, A. Flamholz, J. J. Demarais, D. Serwas and N. Fisher for comments on the manuscript. We thank Donner Laboratories at Lawrence Berkeley National Laboratory and the Bay Area Cryo-EM facility at the University of California, Berkeley for microscope screening and data collection access, respectively. We thank the McGill Pharmacology SPR/MS facility and the Canadian Fund for Innovation for infrastructure support. T.G.L. was supported by a National Science Foundation Graduate Research Fellowship and Molecular Basis of Cell Function National Institutes of Health pre-doctoral training grant (NIGMS project 5T32GM007232-38). A.N.B. was supported by a Healthy Brain for Healthy Lives studentship and J.-F.T. was supported by a Tier 2 Canada Research Chair. D.F.S. and K.M.D. were supported by the US Department of Energy grants DE-SC00016240 and DE-AC02-O5CH11231, respectively.
Reviewer information
Nature thanks R.Cogdell, A. Leitner and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
T.G.L. purified NDH, prepared cryo-EM grids, acquired and processed electron microscopy data, built the atomic models and performed coordinate refinement. J.-F.T. and A.N.B. performed mass spectrometry experiments and analysed data. T.G.L. and K.M.D. designed the project. K.M.D. and D.F.S. supervised the project. T.G.L., K.M.D. and D.F.S. interpreted the structure and wrote the manuscript with contributions from J.-F.T. and A.N.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Purification and cryo-EM of NDH from T. elongatus.
a, Blue native PAGE gel of NDH solubilized by β-DDM and purified by Ni2+-affinity chromatography with fractions, positions of molecular-weight marker and NDH indicated. b, Size-exclusion chromatography trace of concentrated Ni2+-affinity eluates. The gold bar indicates the fraction that shows a single band corresponding to NDH on the blue native PAGE gel (right). See Supplementary Fig. 1 for source gels for a and b. c, Micrograph of frozen hydrated NDH used in this study, with exemplar particles boxed and 2D-class averages generated from dataset 1 showing clear internal features.
Extended Data Fig. 2 Cryo-EM data-processing workflow.
a, b, Schematic of pre-processing, classification and refinement procedures used to generate the maps obtained in this study (see Methods and Supplementary Methods for details) for dataset 1 (a) and dataset 2 (b). Boxed maps indicate those used in model building and solid boxes indicate those used for coordinate refinement. The masks used for focused classification of the X-cofactor are indicated in cyan. The apo- and holo X-cofactor focused maps are enlarged to emphasize the X-site (coloured purple). Resolutions are for an FSC of 0.143, as described in Methods.
Extended Data Fig. 3 Resolution assessment of cryo-EM maps and models.
a–c, Euler angle distributions (a), 3DFSC plots (b) and local resolution (RELION, unsharpened) maps (c) for the overall cryo-EM maps of dataset 1 (top) and dataset 2 (bottom). d, Sharpened maps used for coordinate refinement shown from both sides for dataset 1 (top left, yellow), dataset 2 (top right, pink), and the overlay (bottom left) and composite (bottom right, blue) maps. e, Model compared with map FSC for each dataset and composite map with their respective models calculated with PHENIX. f, Representation of the refined composite coordinate model, coloured according to B factor.
Extended Data Fig. 4 Comparison of the homologous cores of NDH and complex I.
a, Core subunits from T. elongatus (cyan) and T. thermophilus (orange, PDB: 4HEA) superimposed on NdhA/ND1 (the heel subunit) viewed from side (top) and top-down of the membrane arm (bottom). The antiporter domains are related by a rotation of approximately 10° about their central axis. The iron–sulfur clusters are similarly positioned. b, Core subunits and transverse helix aligned individually with observed differences highlighted in colour and labelled (white, conserved structure; blue, distinct to NDH; orange; distinct to complex I). c, Table of core subunit sequence homology (ID) and structural similarity (r.m.s.d.). d, e, Depiction of putative proton-translocation pathways (blue arrows) based on conserved charged residues for the distal (d) and PQ/Q-site adjacent (e) sites. f, Close-up view of the PQ/Q-site based on the location of the coordinating Tyr72 of NdhH (Tyr87 of Nqo4) reveals a difference in the of β1β2 loop of NdhH, which is displaced by approximately 9 Å relative to the complex I homologue Nqo4.
Extended Data Fig. 5 OPS subunits NdhL, NdhM, NdhN, NdhP and NdhQ.
a–d, Cartoon depictions of NdhL, NdhM, NdhN, NdhP and NdhQ, with surface representations indicating their location in the complex. a, NdhK (purple), NdhM (green) and NdhN (blue), with complementary β-strands formed between NdhK and NdhM (dashed red boxes). b, NdhL (red) connects the peripheral and membrane arms. c, NdhP (magenta) trapping a lipid (dark blue mesh) between NdhD (tan) and NdhF (light blue). d, NdhQ (orange) with density for dataset 1 (red mesh) and dataset 2 (blue mesh) for NdhQ and the NdhF transverse helix, which is of higher quality in the presence of NdhQ. All density is at σ = 3.
Extended Data Fig. 6 Electrostatics of the peripheral arm.
a–c, Left, surface model views of the peripheral arm with the NdhI β-hairpin, NdhO and NdhS coloured in cyan, purple and yellow, respectively. Right, corresponding electrostatic potential surfaces calculated at pH 8 (see Methods) with the colour key shown at the bottom. a, Peripheral arm viewed down the β-hairpin of NdhI. The electrostatic potential of this view shows two positive patches circled in white and labelled as NdhO-adjacent (O-site) and NdhS-adjacent (S-site). b, S-site viewed with the indicated rotational relationship to a with Arg69 of NdhS outlined in orange. c, O-site viewed with the indicated rotational relationship to b with the basic triplet (Lys89-Lys90-Lys91) of NdhI and Lys4-Lys5 of NdhO outlined in orange. d, Alignments of NdhO (top) and NdhS (bottom) homologues with β-strand regions indicated, numbered according to the T. elongatus sequence. Boxed regions indicate exposed regions facing the respective sites, and basic residues are coloured blue. Predicted chloroplast localization sequences are removed when applicable. The sequence of S. oleracea NdhS is not available. Residues 1–42 for T. elongatus NdhS are not conserved and are not observed in our structure, and have been removed for the purposes of alignment.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1–12, which show source gels for Extended Data Figure 1 and ESPript alignments of NDH and complex I core subunits used to generate Extended Data Figure 4. Full figure legends are available in a separate file.
Supplementary Information
This file contains the full legends for Supplementary Figures 1–12.
Supplementary Information
This file contains Supplementary Methods.
Supplementary Data
Mass-spectrometry proteomics analysis of purified T. elongatus NDH. Tables of proteins identified by MaxQuant using mass-spectrometry data derived from purified and digested T. elongatus NDH. The first sheet is for proteins digested in solution using both chymotrypsin and trypsin. The second sheet is for two BN-PAGE gel bands digested with chymotrypsin. In both sheets, proteins are clustered as “NDH subunits”, “Other T. elongatus proteins”, or “Contaminants”.
Rights and permissions
About this article
Cite this article
Laughlin, T.G., Bayne, A.N., Trempe, JF. et al. Structure of the complex I-like molecule NDH of oxygenic photosynthesis. Nature 566, 411–414 (2019). https://doi.org/10.1038/s41586-019-0921-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0921-0
This article is cited by
-
The cytochrome b6f complex: plastoquinol oxidation and regulation of electron transport in chloroplasts
Photosynthesis Research (2024)
-
Thermophilic cyanobacteria—exciting, yet challenging biotechnological chassis
Applied Microbiology and Biotechnology (2024)
-
Assessment of P700 Redox State of Tomato Plants Under the Combined Influence of Elevated Temperature and Fusarium oxysporum Infection by Differential Absorption Photometry Using Saturating Light Pulse Technology
Journal of Applied Spectroscopy (2024)
-
Cryo-EM and femtosecond spectroscopic studies provide mechanistic insight into the energy transfer in CpcL-phycobilisomes
Nature Communications (2023)
-
Metabolic landscape in cardiac aging: insights into molecular biology and therapeutic implications
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.