Abstract
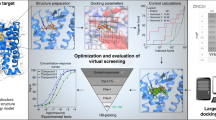
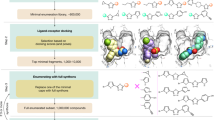
Despite intense interest in expanding chemical space, libraries containing hundreds-of-millions to billions of diverse molecules have remained inaccessible. Here we investigate structure-based docking of 170 million make-on-demand compounds from 130 well-characterized reactions. The resulting library is diverse, representing over 10.7 million scaffolds that are otherwise unavailable. For each compound in the library, docking against AmpC β-lactamase (AmpC) and the D4 dopamine receptor were simulated. From the top-ranking molecules, 44 and 549 compounds were synthesized and tested for interactions with AmpC and the D4 dopamine receptor, respectively. We found a phenolate inhibitor of AmpC, which revealed a group of inhibitors without known precedent. This molecule was optimized to 77 nM, which places it among the most potent non-covalent AmpC inhibitors known. Crystal structures of this and other AmpC inhibitors confirmed the docking predictions. Against the D4 dopamine receptor, hit rates fell almost monotonically with docking score, and a hit-rate versus score curve predicted that the library contained 453,000 ligands for the D4 dopamine receptor. Of 81 new chemotypes discovered, 30 showed submicromolar activity, including a 180-pM subtype-selective agonist of the D4 dopamine receptor.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Active molecules reported here are available from B.K.S. or directly from Enamine. The four structures of AmpC determined with the new docking hits are available from the PDB with accession numbers 6DPZ, 6DPY, 6DPX and 6DPT. The compounds docked in this study are freely available from our ZINC lead-like make-on-demand library (http://zinc15.docking.org). All active compounds are available either from the authors or may be purchased from Enamine. Figures with associated raw data include: Fig. 2, for which electron density and reflection files are deposited with the PDB; Figs. 3, 4 and Extended Data Fig. 5, for which Source Data are available in the online version of the paper; Extended Data Fig. 1, for which the data are included in Supplementary Table 1; Extended Data Fig. 6, for which raw clustering or no-clustering rank numbers are included in Supplementary Tables 8, 9. Further data are provided in Supplementary Tables 3, 5 (aggregation assays for AmpC inhibitors and D4 ligands); Extended Data Table 1 (crystallographic data collection and refinement); Supplementary Tables 9, 10 and Supplementary Data 12–15 (chemical purity of active ligands and their spectra); Supplementary Data 11 and 14 (synthetic routes to compounds). All other data are available from the authors on request.
References
Bohacek, R. S., McMartin, C. & Guida, W. C. The art and practice of structure-based drug design: a molecular modeling perspective. Med. Res. Rev. 16, 3–50 (1996).
Ertl, P. Cheminformatics analysis of organic substituents: identification of the most common substituents, calculation of substituent properties, and automatic identification of drug-like bioisosteric groups. J. Chem. Inf. Comput. Sci. 43, 374–380 (2003).
Fink, T., Bruggesser, H. & Reymond, J. L. Virtual exploration of the small-molecule chemical universe below 160 Daltons. Angew. Chem. Int. Ed. 44, 1504–1508 (2005).
Chevillard, F. & Kolb, P. SCUBIDOO: a large yet screenable and easily searchable database of computationally created chemical compounds optimized toward high likelihood of synthetic tractability. J. Chem. Inf. Model. 55, 1824–1835 (2015).
Keserü, G. M. & Makara, G. M. The influence of lead discovery strategies on the properties of drug candidates. Nat. Rev. Drug Discov. 8, 203–212 (2009).
McGovern, S. L., Caselli, E., Grigorieff, N. & Shoichet, B. K. A common mechanism underlying promiscuous inhibitors from virtual and high-throughput screening. J. Med. Chem. 45, 1712–1722 (2002).
Brenner, S. & Lerner, R. A. Encoded combinatorial chemistry. Proc. Natl Acad. Sci. USA 89, 5381–5383 (1992).
Ahn, S. et al. Allosteric “beta-blocker” isolated from a DNA-encoded small molecule library. Proc. Natl Acad. Sci. USA 114, 1708–1713 (2017).
Goodnow, R. A. Jr, Dumelin, C. E. & Keefe, A. D. DNA-encoded chemistry: enabling the deeper sampling of chemical space. Nat. Rev. Drug Discov. 16, 131–147 (2017).
Jorgensen, W. L. The many roles of computation in drug discovery. Science 303, 1813–1818 (2004).
de Graaf, C. et al. Crystal structure-based virtual screening for fragment-like ligands of the human histamine H1 receptor. J. Med. Chem. 54, 8195–8206 (2011).
Katritch, V. et al. Structure-based discovery of novel chemotypes for adenosine A2A receptor antagonists. J. Med. Chem. 53, 1799–1809 (2010).
Manglik, A. et al. Structure-based discovery of opioid analgesics with reduced side effects. Nature 537, 185–190 (2016).
Wang, S. et al. D4 dopamine receptor high-resolution structures enable the discovery of selective agonists. Science 358, 381–386 (2017).
Negri, A. et al. Discovery of a novel selective kappa-opioid receptor agonist using crystal structure-based virtual screening. J. Chem. Inf. Model. 53, 521–526 (2013).
Jazayeri, A., Andrews, S. P. & Marshall, F. H. Structurally enabled discovery of adenosine A2A receptor antagonists. Chem. Rev. 117, 21–37 (2017).
Lane, J. R. et al. Structure-based ligand discovery targeting orthosteric and allosteric pockets of dopamine receptors. Mol. Pharmacol. 84, 794–807 (2013).
Langmead, C. J. et al. Identification of novel adenosine A2A receptor antagonists by virtual screening. J. Med. Chem. 55, 1904–1909 (2012).
Becker, O. M. et al. G protein-coupled receptors: in silico drug discovery in 3D. Proc. Natl Acad. Sci. USA 101, 11304–11309 (2004).
Kooistra, A. J. et al. Function-specific virtual screening for GPCR ligands using a combined scoring method. Sci. Rep. 6, 28288 (2016).
Congreve, M. et al. Discovery of 1,2,4-triazine derivatives as adenosine A2A antagonists using structure based drug design. J. Med. Chem. 55, 1898–1903 (2012).
Kiss, R. et al. Discovery of novel human histamine H4 receptor ligands by large-scale structure-based virtual screening. J. Med. Chem. 51, 3145–3153 (2008).
Oprea, T. I. & Gottfries, J. Chemography: the art of navigating in chemical space. J. Comb. Chem. 3, 157–166 (2001).
Mysinger, M. M., Carchia, M., Irwin, J. J. & Shoichet, B. K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594 (2012).
Bemis, G. W. & Murcko, M. A. The properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 39, 2887–2893 (1996).
Gaulton, A. et al. The ChEMBL database in 2017. Nucleic Acids Res. 45, D945–D954 (2017).
Katz, B. A. et al. A novel serine protease inhibition motif involving a multi-centered short hydrogen bonding network at the active site. J. Mol. Biol. 307, 1451–1486 (2001).
Congreve, M., Langmead, C. J., Mason, J. S. & Marshall, F. H. Progress in structure based drug design for G protein-coupled receptors. J. Med. Chem. 54, 4283–4311 (2011).
Vaidehi, N. Dynamics and flexibility of G-protein-coupled receptor conformations and their relevance to drug design. Drug Discov. Today 15, 951–957 (2010).
Irwin, J. J. & Shoichet, B. K. Docking screens for novel ligands conferring new biology. J. Med. Chem. 59, 4103–4120 (2016).
Vass, M. et al. in Computational Methods for GPCR Drug Discovery (Heifetz, A.) Ch. 4, 73–113 (Humana, Springer, New Jersey, 2018).
Isberg, V. et al. Generic GPCR residue numbers – aligning topology maps while minding the gaps. Trends Pharmacol. Sci. 36, 22–31 (2015).
McCorvy, J. D. et al. Structural determinants of 5-HT2B receptor activation and biased agonism. Nat. Struct. Mol. Biol. 25, 787–796 (2018).
Powers, R. A., Morandi, F. & Shoichet, B. K. Structure-based discovery of a novel, noncovalent inhibitor of AmpC β-lactamase. Structure 10, 1013–1023 (2002).
Feng, B. Y. et al. A high-throughput screen for aggregation-based inhibition in a large compound library. J. Med. Chem. 50, 2385–2390 (2007).
Babaoglu, K. et al. Comprehensive mechanistic analysis of hits from high-throughput and docking screens against β-lactamase. J. Med. Chem. 51, 2502–2511 (2008).
Rowley, M. et al. 5-(4-chlorophenyl)-4-methyl-3-(1-(2-phenylethyl)piperidin-4-yl)isoxazole: a potent, selective antagonist at human cloned dopamine D4 receptors. J. Med. Chem. 39, 1943–1945 (1996).
Enguehard-Gueiffier, C. et al. 2-[(4-phenylpiperazin-1-yl)methyl]imidazo(di)azines as selective D4-ligands. Induction of penile erection by 2-[4-(2-methoxyphenyl)piperazin-1-ylmethyl]imidazo[1,2-a]pyridine (PIP3EA), a potent and selective D4 partial agonist. J. Med. Chem. 49, 3938–3947 (2006).
Löber, S., Hübner, H. & Gmeiner, P. Synthesis and biological investigations of dopaminergic partial agonists preferentially recognizing the D4 receptor subtype. Bioorg. Med. Chem. Lett. 16, 2955–2959 (2006).
Lindsley, C. W. & Hopkins, C. R. Return of D4 dopamine receptor antagonists in drug discovery. J. Med. Chem. 60, 7233–7243 (2017).
Tirado-Rives, J. & Jorgensen, W. L. Contribution of conformer focusing to the uncertainty in predicting free energies for protein-ligand binding. J. Med. Chem. 49, 5880–5884 (2006).
Abagyan, R., Totrov, M. & Kuznetsov, D. ICM—a new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J. Comput. Chem. 15, 488–506 (1994).
Halgren, T. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 47, 1750–1759 (2004).
Goodsell, D. S. & Olson, A. J. Automated docking of substrates to proteins by simulated annealing. Proteins 8, 195–202 (1990).
Kufareva, I., Katritch, V., Stevens, R. C. & Abagyan, R. Advances in GPCR modeling evaluated by the GPCR Dock 2013 assessment: meeting new challenges. Structure 22, 1120–1139 (2014).
Kramer, B., Rarey, M. & Lengauer, T. Evaluation of the FLEXX incremental construction algorithm for protein-ligand docking. Proteins 37, 228–241 (1999).
McGann, M. FRED pose prediction and virtual screening accuracy. J. Chem. Inf. Model. 51, 578–596 (2011).
Jones, G., Willett, P., Glen, R. C., Leach, A. R. & Taylor, R. Development and validation of a genetic algorithm for flexible docking. J. Mol. Biol. 267, 727–748 (1997).
Corbeil, C. R., Williams, C. I. & Labute, P. Variability in docking success rates due to dataset preparation. J. Comput. Aided Mol. Des. 26, 775–786 (2012).
Hawkins, P. C., Skillman, A. G., Warren, G. L., Ellingson, B. A. & Stahl, M. T. Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. J. Chem. Inf. Model. 50, 572–584 (2010).
Hawkins, G. D. et al. AMSOL version 7.1 https://comp.chem.umn.edu/amsol/ (2004).
Wei, B. Q., Baase, W. A., Weaver, L. H., Matthews, B. W. & Shoichet, B. K. A model binding site for testing scoring functions in molecular docking. J. Mol. Biol. 322, 339–355 (2002).
Mysinger, M. M. & Shoichet, B. K. Rapid context-dependent ligand desolvation in molecular docking. J. Chem. Inf. Model. 50, 1561–1573 (2010).
Sterling, T. & Irwin, J. J. ZINC 15—ligand discovery for everyone. J. Chem. Inf. Model. 55, 2324–2337 (2015).
Barelier, S. et al. Increasing chemical space coverage by combining empirical and computational fragment screens. ACS Chem. Biol. 9, 1528–1535 (2014).
Gray, D. L. et al. Impaired β-arrestin recruitment and reduced desensitization by non-catechol agonists of the D1 dopamine receptor. Nat. Commun. 9, 674 (2018).
Carlsson, J. et al. Ligand discovery from a dopamine D3 receptor homology model and crystal structure. Nat. Chem. Biol. 7, 769–778 (2011).
Meng, E. C., Shoichet, B. K. & Kuntz, I. D. Automated docking with gridb-based energy evaluation. J. Comput. Chem. 13, 505–524 (1992).
Sharp, K. A., Friedman, R. A., Misra, V., Hecht, J. & Honig, B. Salt effects on polyelectrolyte-ligand binding: comparison of Poisson–Boltzmann, and limiting law/counterion binding models. Biopolymers 36, 245–262 (1995).
Gallagher, K. & Sharp, K. Electrostatic contributions to heat capacity changes of DNA-ligand binding. Biophys. J. 75, 769–776 (1998).
Coleman, R. G., Carchia, M., Sterling, T., Irwin, J. J. & Shoichet, B. K. Ligand pose and orientational sampling in molecular docking. PLoS ONE 8, e75992 (2013).
Tolmachev, A. et al. Expanding synthesizable space of disubstituted 1,2,4-oxadiazoles. ACS Comb. Sci. 18, 616–624 (2016).
Eidam, O. et al. Design, synthesis, crystal structures, and antimicrobial activity of sulfonamide boronic acids as β-lactamase inhibitors. J. Med. Chem. 53, 7852–7863 (2010).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Eidam, O. et al. Fragment-guided design of subnanomolar β-lactamase inhibitors active in vivo. Proc. Natl Acad. Sci. USA 109, 17448–17453 (2012).
Feng, B. Y. & Shoichet, B. K. A detergent-based assay for the detection of promiscuous inhibitors. Nat. Protoc. 1, 550–553 (2006).
Allen, J. A. et al. Discovery of β-arrestin-biased dopamine D2 ligands for probing signal transduction pathways essential for antipsychotic efficacy. Proc. Natl Acad. Sci. USA 108, 18488–18493 (2011).
Carpenter, B. et al. Stan: a probabilistic programming language. J. Stat. Softw. 1, 1–32 (2017).
Ryan, E. G., Drovandi, C. C., McGree, J. M. & Pettitt, A. N. A review of modern computational algorithms for Bayesian optimal design. Int. Stat. Rev. 84, 128–154 (2016).
Rainforth, T., Cornish, R., Yang, H., Warrington, A. & Wood, F. On Nesting Monte Carlo Estimators. In Proc. 35th International Conference on Machine Learning PMLR 80 (eds Dy, J. & Krause, A.) 4267–4276 (2018).
Acknowledgements
This research was supported by GM71896 (to J.J.I.); R35 GM122481 and a UCSF PBBR New Frontier Award (to B.K.S.); R01 MH112205, U24DK1169195 and the NIMH Psychoactive Drug Screening Contract (to B.L.R.); Strategic Priority Research Program of the Chinese Academy of Sciences, grant number XDB19000000 (to S.W.). We thank R. Stein and I. Fish for help with AmpC preparation, H. Torosyan for aggregation assays, R. H. J. Olsen for developing the D4 receptor BRET assay, B. Wong and C. Dandarchuluun for computer support, and M. Korczynska and J. Pottel for reading this manuscript; ChemAxon for a license to JChem, OpenEye Scientific software for a license to OEChem and Omega2, Molecular Networks for a license to Corina, and Molinspriation for a license to Mitools.
Reviewer information
Nature thanks M. M. Babu, D. E. Gloriam and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
J.J.I. and B.K.S. conceived the study. J.J.I. created the enlarged docking libraries. J.L., T.E.B. and A.L. performed the docking. S.W., T.C. and B.L.R. performed D4 receptor assays and analysis. I.S. performed all crystallography and assays for AmpC. Y.S.M., K.T. and A.A.T. directed the compound synthesis, purification and characterization. M.J.O. performed Bayesian modelling. E.A., T.E.B. and J.L. contributed enabling computer code. B.K.S., J.L., J.J.I., B.L.R., T.E.B., I.S., A.L., S.W., T.C. and M.J.O. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
B.K.S. and J.J.I. are founders of a company, BlueDolphin LLC, that works in the area of molecular docking. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Simulating the effect of library size on ligand enrichment among the top 1,000 docked molecules.
a, b, The energy distribution of ligands (a) and decoys (b) from docking enrichment calculations against AmpC. The skewed normal fitting curves are plotted in red lines. The fitting parameters (shape (α), location (loc) and scale values) are shown. c, Heat maps of the number of active molecules in the top 1,000 docked molecules for 6 targets. The number of ligands in the top 1,000 docked molecules for a given library size and the ratio between ligands and decoys is coloured using a log10(number of ligands) scale ranging from 1 (blue) to 1,000 (red). Cells with zero ligands are shown in white. d, Large-library docking screens of AmpC (top, n = 99 million molecules) and D4 (bottom, n = 138 million molecules). Molecules that are known to bind to AmpC and D4, as well as close analogues, are treated as ligands and the rest of the molecules are treated as decoys. Left, the energy distributions of decoys (grey), ligands defined by ECFP4 Tc similarity ≥0.5 (blue), 0.6 (green) and 0.7 (orange) to ligands from ChEMBL. Middle, heat maps of the number of ligands in the top 1,000 docked molecules based on fit to full-library docking with the ligands (AmpC, Tc ≥ 0.5, green; D4, Tc ≥ 0.6, orange) and decoys (grey) distributions. Right, number of ligands in the top 1,000 docked molecules as the library grows based on actual distributions plotted in the left panel. Data are mean ± s.d. of 20 samples. See Supplementary Table 1 for retrospective performance on three more targets.
Extended Data Fig. 2 Initial hits and selected analogues against AmpC.
The five initial hits are shown in the left column. Under each compound, the first row includes the ZINC identifier; the second row is the cluster rank (position in cluster head list sorted by DOCK score) with global rank (position in unclustered hit list sorted by DOCK score) shown in brackets; the third row is the Tc value (Tanimoto coefficient to known AmpC inhibitors in ChEMBL); the fourth row is the Ki value. Five selected analogues for the corresponding hits are shown in the right column. Under each compound, the first row includes the ZINC identifier; the second row is the Tc value; and the third row is the Ki value.
Extended Data Fig. 3 Lineweaver–Burk plot and Ki analysis for analogues of each of the five series of AmpC inhibitors.
a–f, Lineweaver–Burk plots for ZINC776666294 (a), 275579920 (b), ZINC548592534 (c), ZINC1187516987 (d), 339204163 (e) and 549719643 (f), indicating competitive inhibition. IC50 values were determined by nonlinear regression fit in GraphPad Prism, and Ki values calculated by a replot of the slope of each Lineweaver–Burk plot versus the corresponding inhibitor concentration.
Extended Data Fig. 4 Electron density maps for AmpC–inhibitor complexes.
The initial Fo − Fc electron density map contoured at 2.5σ around the inhibitor (density in cyan) with refined 2Fo − Fc electron density contoured at 1σ for enzyme residues for the complexes with the following compounds. a, 547933290. b, 275579920. c, 339204163. d, 549719643. Inhibitor carbons are shown in cyan and enzyme carbons are shown in grey, oxygens in red, nitrogens in blue, sulfurs in yellow and chlorides in green.
Extended Data Fig. 5 Selected D4 hits from docking 138 million make-on-demand molecules.
Six ligands with docked poses (first column), cAMP Gαi/o activities (second column), Tango β-arrestin activities (third column) and 3H-N-methylspiperone displacement and chemical drawing (fourth column) are shown. The receptor structure is in grey and ligand carbons are in teal. Ballesteros–Weinstein residue numbers are included as superscripts. Functional assays represent normalized concentration–response curves of the ligands in cloned human D4-mediated activation of Gαi/o and β-arrestin translocation. Data are mean ± s.e.m. of three assays. The first row shows an example of an antagonist identified among the D4 hits. Both agonist (teal curve) and antagonist (purple curve) modes are shown for ZINC130532671 in the third panel; the concentration of quinpirole in the antagonist mode was 100 nM.
Extended Data Fig. 6 Pre-clustering the docking library yields much worse scores of scaffold representatives compared to full library docking.
a, b, Comparison of energy distributions of scaffold representatives between full library docking (orange) and pre-clustered library docking for D4 (a) and AmpC (b) using four strategies: the closest member to the centroid of molecular masses and clogP (blue), the closest member to the centroid of molecular masses (pink), the member with the largest molecular masses (magenta) and the member with the smallest molecular masses (green). The inset shows the ratio of the number of molecules at a given docking score for full library docking divided by the number at that score when only cluster representatives are docked (coloured by clustering method). For each target, two examples illustrate the effect on our experimentally active scaffold families. c, D4. d, AmpC. The scaffold for each molecule is highlighted in red. The ZINC identifier, post-cluster rank and pre-cluster rank are labelled for each pair. The arrow colour is as for the pre-clustering methods in a and b.
Extended Data Fig. 7 Comparison of hit rates achieved by combined docking score and human prioritization compared to the rates achieved by docking score alone.
a, The hit rates for selecting compounds at different scoring ranges by each strategy: human prioritization and docking score (orange), or docking score alone (blue). Hit rate is the ratio of active compounds/tested compounds; the raw numbers appear at the top of each bar. b, Distribution of the binding affinity level among the hits from a. There are 32 hits from human prioritization and docking score, and 26 hits from the docking score alone. These are divided into three affinity ranges: <100 nM (pale blue); 100 nM–1 μM (blue); 1–10 μM (dark blue). c, Functional activity distribution among the hits from b. There are 22 molecules from human prioritization and docking score, and 7 molecules from the docking score alone. These are divided into five activity ranges: <10 nM (pale green); 10 nM–1 μM (light green); 1–10 μM (olive); 10–50 μM (forest green); and not determined (dark green).
Extended Data Fig. 8 Bayesian prior modelling for balancing information gain and ligand discovery in molecule-selection design and error estimation.
a, Sigmoid functional form for the hit-rate model. b–d, Marginal Bayesian prior (teal) and posterior (red) distributions (n = 200,000) for each model parameter. b, Top. c, Dock50. d, Slope. e, Estimated hit rate based on evaluation by the authors of the docked poses before any molecules were tested. Brown, mean ± s.d.; n = 200, 220, 230, 230, 285, 235, 210, 230 and 200 compounds; n = 5, 4, 4, 4, 4, 4, 4, 4 and 4 experts. The prior mean (green) and samples (n = 200) from the prior (blue) are shown. f, Candidate (blue) and chosen (orange) experimental designs (inset, designs 1–6), with expected number of hits and information gain for each design. g, Expected number of active scaffolds (orange, mean; grey, posterior draws n = 200,000) superimposed on the total number of scaffold cluster heads (black). h, i, Marginal distribution of the number of active compounds (h) and scaffolds (i) over the posterior distributions (n = 200,000).
Supplementary information
Supplementary Information
This file contains Supplementary Tables and Data 1-15, except Supplementary Tables 2, 4, 7 and 8, which are provided as separate files.
Supplementary Table
This file contains Supplementary Table 2: Molecules tested against β-lactamase AmpC. We report the zinc id, smiles, and indicator of binding or non-binding. (supplied as a separate file. See Extended Data Fig. 2 for affinities and chemical drawings of potent binders).
Supplementary Table
This file contains Supplementary Table 4: All molecules tested against D4.
Supplementary Table
This file contains Supplementary Table 7: Full-library vs pre-clustering library docking for D4.
Supplementary Table
This file contains Supplementary Table 8: Full-library vs pre-clustering library docking for AmpC.
Rights and permissions
About this article
Cite this article
Lyu, J., Wang, S., Balius, T.E. et al. Ultra-large library docking for discovering new chemotypes. Nature 566, 224–229 (2019). https://doi.org/10.1038/s41586-019-0917-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0917-9
This article is cited by
-
An intriguing vision for transatlantic collaborative health data use and artificial intelligence development
npj Digital Medicine (2024)
-
Exploring the combinatorial explosion of amine–acid reaction space via graph editing
Communications Chemistry (2024)
-
A divergent intermediate strategy yields biologically diverse pseudo-natural products
Nature Chemistry (2024)
-
Computational drug development for membrane protein targets
Nature Biotechnology (2024)
-
Integrating QSAR modelling and deep learning in drug discovery: the emergence of deep QSAR
Nature Reviews Drug Discovery (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.