Abstract
There is growing evidence that tumour neoantigens have important roles in generating spontaneous antitumour immune responses and predicting clinical responses to immunotherapies1,2. Despite the presence of numerous neoantigens in patients, complete tumour elimination is rare, owing to failures in mounting a sufficient and lasting antitumour immune response3,4. Here we show that durable neoantigen-specific immunity is regulated by mRNA N6-methyadenosine (m6A) methylation through the m6A-binding protein YTHDF15. In contrast to wild-type mice, Ythdf1-deficient mice show an elevated antigen-specific CD8+ T cell antitumour response. Loss of YTHDF1 in classical dendritic cells enhanced the cross-presentation of tumour antigens and the cross-priming of CD8+ T cells in vivo. Mechanistically, transcripts encoding lysosomal proteases are marked by m6A and recognized by YTHDF1. Binding of YTHDF1 to these transcripts increases the translation of lysosomal cathepsins in dendritic cells, and inhibition of cathepsins markedly enhances cross-presentation of wild-type dendritic cells. Furthermore, the therapeutic efficacy of PD-L1 checkpoint blockade is enhanced in Ythdf1−/− mice, implicating YTHDF1 as a potential therapeutic target in anticancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. RIP–seq, Ribo-seq and m6A-seq datasets have been deposited in the Gene Expression Omnibus (GEO) under the accession number GSE115106. A summary of sequencing experiments is provided in Supplementary Table 3. The differential translational efficiency results provided in Supplementary Table 4. Source Data for bar graphs and box-plots in the Figures and Extended Data Figures are provided in separate Excel files.
Change history
25 March 2019
In this Letter, a citation to ‘Fig. 1e’ has been corrected to ‘Fig. 1d’ in the sentence starting “By contrast, the anti-tumour response…”. This has been corrected online
References
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Yarchoan, M., Johnson, B. A. III, Lutz, E. R., Laheru, D. A. & Jaffee, E. M. Targeting neoantigens to augment antitumour immunity. Nat. Rev. Cancer 17, 209–222 (2017).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Wang, X. et al. N 6-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Desrosiers, R., Friderici, K. & Rottman, F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc. Natl Acad. Sci. USA 71, 3971–3975 (1974).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Jia, G. et al. N 6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Wang, X. et al. N 6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m6A-dependent translation control. Nature 552, 126–131 (2017).
Li, Z. et al. FTO plays an oncogenic role in acute myeloid leukemia as a N 6-Methyladenosine RNA demethylase. Cancer Cell 31, 127–141 (2017).
Vu, L. P. et al. The N 6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 23, 1369–1376 (2017).
Liu, J. et al. m6A mRNA methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer. Nat. Cell Biol. 20, 1074–1083 (2018).
Shi, H. et al. m6A facilitates hippocampus-dependent learning and memory through YTHDF1. Nature 563, 249–253 (2018).
Yadav, M. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014).
Mellman, I., Coukos, G. & Dranoff, G. Cancer immunotherapy comes of age. Nature 480, 480–489 (2011).
Jongbloed, S. L. et al. Human CD141+ (BDCA-3)+ dendritic cells (DCs) represent a unique myeloid DC subset that cross-presents necrotic cell antigens. J. Exp. Med. 207, 1247–1260 (2010).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Naik, S. H. et al. Cutting edge: generation of splenic CD8+ and CD8– dendritic cell equivalents in Fms-like tyrosine kinase 3 ligand bone marrow cultures. J. Immunol. 174, 6592–6597 (2005).
Mayer, C. T. et al. Selective and efficient generation of functional Batf3-dependent CD103+ dendritic cells from mouse bone marrow. Blood 124, 3081–3091 (2014).
Kretzer, N. M. et al. RAB43 facilitates cross-presentation of cell-associated antigens by CD8α+ dendritic cells. J. Exp. Med. 213, 2871–2883 (2016).
Driessens, G., Kline, J. & Gajewski, T. F. Costimulatory and coinhibitory receptors in anti-tumor immunity. Immunol. Rev. 229, 126–144 (2009).
Fuertes, M. B. et al. Host type I IFN signals are required for antitumor CD8+ T cell responses through CD8α+ dendritic cells. J. Exp. Med. 208, 2005–2016 (2011).
Woo, S. R. et al. STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors. Immunity 41, 830–842 (2014).
Cebrian, I. et al. Sec22b regulates phagosomal maturation and antigen crosspresentation by dendritic cells. Cell 147, 1355–1368 (2011).
Samie, M. & Cresswell, P. The transcription factor TFEB acts as a molecular switch that regulates exogenous antigen-presentation pathways. Nat. Immunol. 16, 729–736 (2015).
Benci, J.L. et al. Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade. Cell 167, 1540–1554 (2016).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “OMICs” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Uren, P. J. et al. Site identification in high-throughput RNA-protein interaction data. Bioinformatics 28, 3013–3020 (2012).
Cui, X. et al. Guitar: An R/Bioconductor package for gene annotation guided transcriptomic analysis of RNA-related genomic features. BioMed Res. Int. 2016, 8367534 (2016).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
This study was supported by the National Key Research and Development Program of China, Stem Cell and Translational Research (2018YFA0109700 to D.H.), Strategic Priority Research Program of the Chinese Academy of Science (XDA16010404 to D.H.), National Institute of Health (HG008935 and GM113194 to C.H.), Ludwig Center at the University of Chicago (to C.H. and R.R.W.), CAS Hundred Talent Program (to D.H.), National Natural Science Foundation of China (31870890 to M.M.X., 31741074 to D.H.), National Science Fund for Excellent Young Scholars (31622039 to B.S.), Science Foundation for Distinguished Young Scholars of Jiangsu Province (BK20160045 to B.S.) and Open Project of Key Laboratory of Genomic and Precision Medicine of the CAS. The Mass Spectrometry Facility of the University of Chicago is funded by National Science Foundation (CHE-1048528). C.H. is an investigator of the Howard Hughes Medical Institute. We thank J. Tauler for editing.
Reviewer information
Nature thanks J. Hanna, J. Neefjes and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
D.H. and M.M.X. conceived the project. D.H., M.M.X., J.L., L.D., X.H., Y.L. and R.C. performed experimental work. D.H. and C.C. performed bioinformatics analysis. Y.L., J.W. and B.S. generated Ythdf1 knockout mice. M.B.B. and U.D. provided human colon biopsy samples. D.H., M.M.X. and C.H. designed the study. D.H., M.M.X., C.H. and R.R.W. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
C.H. is a scientific founder and a member of the scientific advisory board of Accent Therapeutics, Inc. A patent application on YTHDF1 has been filed by the University of Chicago.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Deletion efficacy of Ythdf1−/− mice.
a, b, Off-target analysis of the CRISPR–Cas9 system in Ythdf1−/− mice. a, Ythdf1 single guide RNA (sgRNA) targeting sites and four putative off-target sites were amplified. b, PCR products from Ythdf1−/− mice and wild-type mice were mixed and digested by T7EI. The PCR product from wild-type mice was used as negative control. c, Immunoblot assays are shown to validate changes in YTH protein expression in Ythdf1−/− DCs. Data are representative of one experiment (a, b) and two independent biological replications (c).
Extended Data Fig. 2 Characterization of immune phenotypes of Ythdf1−/− mice.
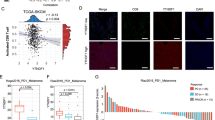
a, Data points for Fig. 1a. b, Wild-type or Ythdf1−/− mice were injected s.c. with 106 B16-OVA cells. Survival was monitored. Mice with tumour volumes less than 200 mm3 are considered to be surviving. One of three representative experiments is shown. c, Data points for Fig. 1b. d–h, Wild-type or Ythdf1−/− mice were injected s.c. with 106 B16-OVA cells. d, e, Frequency of tumour infiltrating MDSCs (Ly6C+CD11b+) was assessed 12 days after tumour inoculation. f, g, The percentages of Treg cells in spleen, DLN and tumour are shown. h, Degranulation of tumour NK cells in response to in vitro re-stimulation with PMA/ionomycin. i, Data points for Fig. 1d. Data are representative of two independent experiments (a, c). n, number of mice. Mean ± s.e.m., two-sided unpaired Student’s t-test (a, c, e, g–i); two-sided log-rank (Mantel–Cox) test (b).
Extended Data Fig. 3 Cross-priming of tumour neoantigens is increased in Ythdf1−/− mice.
a, Rag2−/− mice were inoculated with T cells isolated from wild-type or Ythdf1−/− mice on day 0. On the same day, mice were injected s.c. with 5 × 105 B16-OVA cells. Tumour growth was monitored over time. b, Wild-type or Ythdf1−/− mice were injected s.c. with 106 MC38-OTIp cells. Six days after tumour inoculation, CD8+ or CD11b+ DCs were sorted from DLNs. DCs were co-cultured with CD8+ T cells isolated from naive OT-I mice. Cross-priming capacity was determined by the production of IFNγ. c, Wild-type or Ythdf1−/− mice were injected s.c. with 106 MC38-SIY cells. Six days after tumour inoculation, DCs were sorted from DLNs and co-cultured with CD8+ T cells isolated from naive 2C mice. Cross-priming capacity was determined by the production of IFNγ. d, Wild-type or Mett14-deficient GMDCs were co-cultured with B16-OVA cells. Cross-priming capacity was determined by the production of IFNγ. e, Wild-type and Ythdf1−/− mice were injected s.c. with 106 B16-OVA cells. Data are shown as the expression of CD80 and CD86 on tumour-infiltrating DCs. f, Wild-type and Ythdf1−/− mice were injected s.c. with 106 B16-OVA cells. Six days after tumour inoculation, CD8+ or CD11b+ DCs were sorted from DLNs. DCs were pulsed with 1 μg/ml exogenous OT-I peptide and co-cultured with isolated CD8+ T cells from naive OT-I mice for 3 days, and then analysed by IFNγ CBA. Data are representative of two independent experiments with similar results (e). n, number of mice. Mean ± s.e.m., two-sided unpaired Student’s t-test (a–c, f) or one-sided unpaired Student’s t-test (d).
Extended Data Fig. 4 Development of DCs and T cells is similar in Ythdf1+/+ and Ythdf1−/− mice.
a, b, Percentages of CD11b+ and CD8α+ DCs in lymph node (LN) and spleen. c, d, Percentages of CD4+ and CD8+ T cells in lymph node and spleen. No significant difference was detected between wild-type and Ythdf1−/− mice. n, number of mice. Mean ± s.e.m., two-sided unpaired Student’s t-test.
Extended Data Fig. 5 In vitro functional analysis of GMDCs generated from Ythdf1−/− mice.
a, Production of IL-6, CCL2 and TNFα upon stimulation of Ythdf1−/− GMDCs with LPS. b, c, Wild-type and Ythdf1−/− mice were injected s.c. with 106 B16-OTI-zsGreen cells. The percentage of tumour-infiltrating zsGreen+ DCs six days after tumour inoculation is shown. Data are representative of two independent experiments (b). d, Splenic DCs from wild-type and Ythdf1−/− mice were stimulated with LPS overnight. The cross-presentation capacity of DCs in response to soluble OVA was assessed. n = 3 independent experiments (a); n = 6 independent experiments (d). n, number of mice. Mean ± s.e.m., two-sided unpaired Student’s t-test.
Extended Data Fig. 6 Transcriptome-wide analysis of YTHDF1-binding sites in FLT3L-DCs.
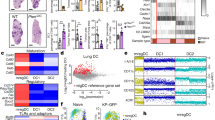
a, High reproducibility of YTHDF1 RIP–seq data. For each potential YTHDF1 binding peak, the fold-enrichment of the RIP/input signal was determined for both replicate 1 (Rep1) and replicate 2. Peaks identified in both replicates were considered as high-confidence peaks and are indicated in red. b, Overlap of YTHDF1-binding transcripts revealed from RIP–seq of two biological replicates. c, Meta-gene analysis to show the distribution of YTHDF1-binding sites along a normalized transcript. d, Distribution of YTHDF1-binding sites in transcripts. TTS, transcription termination site. e, Heatmap showing the translational efficiency of co-simulatory/inhibitory proteins (signal 2) and cytokines (signal 3) in wild-type and Ythdf1−/− FLT3L-DCs.
Extended Data Fig. 7 THDF1-deficient GMDCs exhibit lower translational rates.
a, High reproducibility of YTHDF1 RIP-seq data in GMDCs. For each potential YTHDF1 binding peak, the fold-enrichment of the RIP/input signal was determined for both Replicate 1 and Replicate 2. Peaks identified in both replicates were considered high-confidence peaks and are indicated in red. b, Volcano plots of genes with differential translational efficiencies in wild-type and Ythdf1−/− GMDCs. YTHDF1 targets are marked with yellow circles. P values calculated using two-sided likelihood ratio test with Benjamini–Hochberg adjustment; n = 4 (2 conditions × 2 biological replicates). c, Cumulative distribution of the fold change in translational efficiency between wild-type and Ythdf1−/− GMDCs. P values calculated using two-sided Kolmogorov–Smirnov test; n = 2 independent biological replicates. Box-plot elements: centre line, median; box limits, upper and lower quartiles; whiskers, 1–99%. d, Distribution of YTHDF1-binding sites in transcripts. e, Metagene plot depicting nearly unchanged distribution of m6A peaks and similar consensus motifs in wild-type and Ythdf1−/− GMDCs. P values of consensus motifs generated by HOMER29 using one-sided binomial test. f, KEGG and GO enrichment analysis of YTHDF1 target genes revealed enrichment of biological functions related to the innate immune system, lysosomes and phagosomes (n = 79). One-sided hypergeometric test was used to determine the statistical significance of enrichment. g, Heatmap showing translational efficiency of cathepsin genes in GMDCs and FLT3L-DCs. n, number of genes or m6A peaks.
Extended Data Fig. 8 Antigen degradation is reduced in Ythdf1−/− mice and inhibition of protease cathepsins enhances cross-priming of wild-type DCs.
a, GMDCs were co-cultured with necrotic B16-OVA cells overnight. Immunoblot analysis of cathespins B, D and L (CTSB, CTSD and CTSL) in GMDCs. b, Wild-type and Ythdf1−/− DCs were treated with actinomycin D and RNAs were collected at different time points after treatment. mRNA levels were measured using RT–qPCR and represented as mRNA remaining after transcription inhibition (TI). c, GMDCs were co-cultured with necrotic B16-OVA cells overnight and OVA degradation in BMDCs was measured by immunoblot. d, Ex vivo purified wild-type cDCs were pre-treated with 0.04 μM E64 and pulsed with OVA protein for 4 h. The cross-priming capacity of DCs was compared by co-culturing DCs with CTV-labelled OT-I T cells. Proliferation was measured by the dilution of CTV. e, GMDCs were pre-treated with 0.2–2 μM E64 and co-cultured with B16-OVA cells. The cross-priming capacity of DCs was compared by co-culturing DCs with isolated CD8+ T cells from naive OT-I mice and analysed by IFNγ CBA. f, FLT3L-DCs were pre-treated with cathepsin inhibitor CA-074 or/and cathepsin L inhibitor III (CASIII), followed by co-culturing with necrotic B16-OVA cells. Synergistic inhibition was observed. The cross-priming capacity of DCs was determined. g, Data points for Fig. 4b. n = 3 independent experiments with similar results (a, c); n = 2 independent experiments (b). n, sample size. Mean ± s.e.m., two-sided unpaired Student’s t-test (e) or one-sided unpaired Student’s t-test (f).
Extended Data Fig. 9 IFNγ in tumour tissues is responsible for the upregulation of PD-L1 in Ythdf1−/− mice.
Tumour-bearing mice were treated with 50 μg anti-IFNγ monoclonal antibody intratumorally and PD-L1 expression on tumour cells is shown. n, number of mice. Mean ± s.e.m., two-sided unpaired Student’s t-test.
Supplementary information
Supplementary Figure 1
This file contains the uncropped scans of the western blots displayed in the Extended Data Figures.
Supplementary Table 1
A summary of antibody information used in this study.
Supplementary Table 2
A summary of patients’ sample information used in this study.
Supplementary Table 3
A summary of sequencing samples mapping information in this study.
Supplementary Table 4
A summary of ribosome profiling sequencing results for WT and Ythdf1-/- DCs. n = 4 (two conditions × two biological replicates), P values were calculated by a two-sided likelihood ratio test and adjusted by the Benjamini & Hochberg method. Sheet 1: List of translation efficiency for transcripts in WT and Ythdf1-/- Flt3L-DCs. Sheet 2: List of translation efficiency for transcripts in WT and Ythdf1-/- GMDCs.
Source data
Rights and permissions
About this article
Cite this article
Han, D., Liu, J., Chen, C. et al. Anti-tumour immunity controlled through mRNA m6A methylation and YTHDF1 in dendritic cells. Nature 566, 270–274 (2019). https://doi.org/10.1038/s41586-019-0916-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-0916-x
This article is cited by
-
Comprehensive analysis of m6A modification in immune infiltration, metabolism and drug resistance in hepatocellular carcinoma
Cancer Cell International (2024)
-
Regulation of inflammatory diseases via the control of mRNA decay
Inflammation and Regeneration (2024)
-
New horizons for the role of RNA N6-methyladenosine modification in hepatocellular carcinoma
Acta Pharmacologica Sinica (2024)
-
Identification of m6A/m5C-related lncRNA signature for prediction of prognosis and immunotherapy efficacy in esophageal squamous cell carcinoma
Scientific Reports (2024)
-
A lncRNA Dleu2-encoded peptide relieves autoimmunity by facilitating Smad3-mediated Treg induction
EMBO Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.