Abstract
The development of neural circuits relies on axon projections establishing diverse, yet well-defined, connections between areas of the nervous system. Each projection is formed by growth cones—subcellular specializations at the tips of growing axons, encompassing sets of molecules that control projection-specific growth, guidance, and target selection1. To investigate the set of molecules within native growth cones that form specific connections, here we developed growth cone sorting and subcellular RNA–proteome mapping, an approach that identifies and quantifies local transcriptomes and proteomes from labelled growth cones of single projections in vivo. Using this approach on the developing callosal projection of the mouse cerebral cortex, we mapped molecular enrichments in trans-hemispheric growth cones relative to their parent cell bodies, producing paired subcellular proteomes and transcriptomes from single neuron subtypes directly from the brain. These data provide generalizable proof-of-principle for this approach, and reveal molecular specializations of the growth cone, including accumulations of the growth-regulating kinase mTOR2, together with mRNAs that contain mTOR-dependent motifs3,4. These findings illuminate the relationships between subcellular distributions of RNA and protein in developing projection neurons, and provide a systems-level approach for the discovery of subtype- and stage-specific molecular substrates of circuit wiring, miswiring, and the potential for regeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated and analysed during the current study are available in the Harvard Dataverse repository https://doi.org/10.7910/DVN/ISOEB6.
References
Lowery, L. A. & Van Vactor, D. The trip of the tip: understanding the growth cone machinery. Nat. Rev. Mol. Cell Biol. 10, 332–343 (2009).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Meyuhas, O., Avni, D. & Shama, S. Translational control of ribosomal protein mRNAs in eukaryotes. 30, 363–388 (1996).
Fonseca, B. D. et al. La-related protein 1 (LARP1) represses terminal oligopyrimidine (TOP) mRNA translation downstream of mTOR complex 1 (mTORC1). J. Biol. Chem. 290, 15996–16020 (2015).
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Pfenninger, K. H., Ellis, L., Johnson, M. P., Friedman, L. B. & Somlo, S. Nerve growth cones isolated from fetal rat brain: subcellular fractionation and characterization. Cell 35, 573–584 (1983).
Lohse, K. et al. Axonal origin and purity of growth cones isolated from fetal rat brain. Brain Res. Dev. Brain Res. 96, 83–96 (1996).
Fame, R. M., MacDonald, J. L. & Macklis, J. D. Development, specification, and diversity of callosal projection neurons. Trends Neurosci. 34, 41–50 (2011).
Greig, L. C., Woodworth, M. B., Greppi, C. & Macklis, J. D. Ctip1 controls acquisition of sensory area identity and establishment of sensory input fields in the developing neocortex. Neuron 90, 261–277 (2016).
Llorca, O. et al. Eukaryotic type II chaperonin CCT interacts with actin through specific subunits. Nature 402, 693–696 (1999).
Moccia, R. et al. An unbiased cDNA library prepared from isolated Aplysia sensory neuron processes is enriched for cytoskeletal and translational mRNAs. J. Neurosci. 23, 9409–9417 (2003).
Leung, K.-M. et al. Asymmetrical beta-actin mRNA translation in growth cones mediates attractive turning to netrin-1. Nat. Neurosci. 9, 1247–1256 (2006).
Crino, P. B. & Eberwine, J. Molecular characterization of the dendritic growth cone: regulated mRNA transport and local protein synthesis. Neuron 17, 1173–1187 (1996).
Taylor, A. M. et al. Axonal mRNA in uninjured and regenerating cortical mammalian axons. J. Neurosci. 29, 4697–4707 (2009).
Zivraj, K. H. et al. Subcellular profiling reveals distinct and developmentally regulated repertoire of growth cone mRNAs. J. Neurosci. 30, 15464–15478 (2010).
Jung, H., Yoon, B. C. & Holt, C. E. Axonal mRNA localization and local protein synthesis in nervous system assembly, maintenance and repair. Nat. Rev. Neurosci. 13, 308–324 (2012).
Catapano, L. A., Arnold, M. W., Perez, F. A. & Macklis, J. D. Specific neurotrophic factors support the survival of cortical projection neurons at distinct stages of development. J. Neurosci. 21, 8863–8872 (2001).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Özdinler, P. H. & Macklis, J. D. IGF-I specifically enhances axon outgrowth of corticospinal motor neurons. Nat. Neurosci. 9, 1371–1381 (2006).
Meyuhas, O. & Kahan, T. The race to decipher the top secrets of TOP mRNAs. Biochim. Biophys. Acta 1849, 801–811 (2015).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol. Cell. Proteomics 11, M111.014050 (2012).
Liebermeister, W. et al. Visual account of protein investment in cellular functions. Proc. Natl Acad. Sci. USA 111, 8488–8493 (2014).
Tang, H. et al. Amino acid-induced translation of TOP mRNAs is fully dependent on phosphatidylinositol 3-kinase-mediated signaling, is partially inhibited by rapamycin, and is independent of S6K1 and rpS6 phosphorylation. Mol. Cell. Biol. 21, 8671–8683 (2001).
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Kye, M. J. et al. SMN regulates axonal local translation via miR-183/mTOR pathway. Hum. Mol. Genet. 23, 6318–6331 (2014).
Tcherkezian, J. et al. Proteomic analysis of cap-dependent translation identifies LARP1 as a key regulator of 5'TOP mRNA translation. Genes Dev. 28, 357–371 (2014).
Dhand, R. et al. PI 3-kinase: structural and functional analysis of intersubunit interactions. EMBO J. 13, 511–521 (1994).
Itoh, Y. et al. PDK1-Akt pathway regulates radial neuronal migration and microtubules in the developing mouse neocortex. Proc. Natl Acad. Sci. USA 113, E2955–E2964 (2016).
Lu, Y., Belin, S. & He, Z. Signaling regulations of neuronal regenerative ability. Curr. Opin. Neurobiol. 27, 135–142 (2014).
Okabe, M., Ikawa, M., Kominami, K., Nakanishi, T. & Nishimune, Y. ‘Green mice’ as a source of ubiquitous green cells. FEBS Lett. 407, 313–319 (1997).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Gallardo, T., Shirley, L., John, G. B. & Castrillon, D. H. Generation of a germ cell-specific mouse transgenic Cre line, Vasa-Cre. Genesis 45, 413–417 (2007).
Risson, V. et al. Muscle inactivation of mTOR causes metabolic and dystrophin defects leading to severe myopathy. J. Cell Biol. 187, 859–874 (2009).
Rhee, J. M. et al. In vivo imaging and differential localization of lipid-modified GFP-variant fusions in embryonic stem cells and mice. Genesis 44, 202–218 (2006).
Matsuda, T. & Cepko, C. L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl Acad. Sci. USA 101, 16–22 (2004).
Saito, T. & Nakatsuji, N. Efficient gene transfer into the embryonic mouse brain using in vivo electroporation. Dev. Biol. 240, 237–246 (2001).
Biesemann, C. et al. Proteomic screening of glutamatergic mouse brain synaptosomes isolated by fluorescence activated sorting. EMBO J. 33, 157–170 (2014).
Catapano, L. A., Arlotta, P., Cage, T. A. & Macklis, J. D. Stage-specific and opposing roles of BDNF, NT-3 and bFGF in differentiation of purified callosal projection neurons toward cellular repair of complex circuitry. Eur. J. Neurosci. 19, 2421–2434 (2004).
Molyneaux, B. J. et al. Novel subtype-specific genes identify distinct subpopulations of callosal projection neurons. J. Neurosci. 29, 12343–12354 (2009).
Galazo, M. J., Emsley, J. G. & Macklis, J. D. Corticothalamic projection neuron development beyond subtype specification: Fog2 and intersectional controls regulate intraclass neuronal diversity. Neuron 91, 90–106 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Patro, R., Mount, S. M. & Kingsford, C. Sailfish enables alignment-free isoform quantification from RNA-seq reads using lightweight algorithms. Nat. Biotechnol. 32, 462–464 (2014).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Berriz, G. F. & Roth, F. P. The Synergizer service for translating gene, protein and other biological identifiers. Bioinformatics 24, 2272–2273 (2008).
Binder, J. X. et al. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database (Oxford) 2014, bau012 (2014).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Cox, J. & Mann, M. 1D and 2D annotation enrichment: a statistical method integrating quantitative proteomics with complementary high-throughput data. BMC Bioinformatics 13 (Suppl. 16), S12 (2012).
Spandidos, A., Wang, X., Wang, H. & Seed, B. PrimerBank: a resource of human and mouse PCR primer pairs for gene expression detection and quantification. Nucleic Acids Res. 38, D792–D799 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank A.-K. Hadjantonakis and S. Srinivas for reagents; B. Noble and I. Florea for technical support; J. LaVecchio, G. Buruzula and S. Ionescu of HSCRB-HSCI Flow Cytometry Core; W. Lane, J. Neveu and B. Budnik of the Harvard FAS CSB Mass Spectrometry and Proteomics Resource Laboratory; the Harvard Center for Biological Imaging for infrastructure and support; L. Liang for statistical advice; N. Brose, C. Biesemann, E. Herzog, L. Reijmers and J. Rinn for discussions. This work was supported by grants to J.D.M. from the Paul G. Allen Frontiers Group, Brain Research Foundation Scientific Innovations Award program; NIH Pioneer Award DP1 NS106665, Emily and Robert Pearlstein Fund, and the Max and Anne Wien Professorship; with additional infrastructure support from NIH grants NS045523, NS075672, NS049553, and NS041590. A.P. was partially supported by a European Molecular Biology Organization Long Term Fellowship and a Human Frontier Science Program Long Term Fellowship. A.J.M. was partially supported by NIH Training Grant T32 AG000222. Work by R.K. and the Harvard Chan Bioinformatics Core was supported by funds from the Harvard NeuroDiscovery Center and Harvard Stem Cell Institute. J.D.M. is an Allen Distinguished Investigator of the Paul G. Allen Frontiers Group.
Reviewer information
Nature thanks D. Sabatini, J. Henley and the anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.P. and J.D.M. conceived of the project. A.P., A.J.M. and J.D.M. designed experiments. A.P., A.J.M., A.O., P.D. and J.H. performed experiments. A.P., A.J.M., A.O., J.H., R.K. and J.D.M. analysed data. A.P., A.J.M., P.D. and J.D.M. interpreted results. A.P. and J.D.M. wrote the manuscript. All authors contributed to discussions and manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
A US patent application (15/144,660) is pending on growth cone sorting technology and implications. The application lists J.D.M., A.P. and A.J.M. as inventors. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Isolated GC verification.
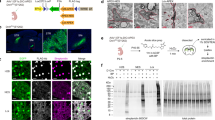
a, Enrichment analysis of GC fraction versus starting homogenate (input). Left, western blot detects protein enrichment of GAP43 (GC marker) and depletion of MAP2 (somato-dendritic protein marker) in GC fraction. Middle, native gel electrophoresis shows de-enriched presence of large (28S) and small (18S) ribosomal subunit rRNA in GC fraction. Right, RT–PCR detects mRNA for Actb (ubiquitous) but not Gfap (progenitor and glial marker) from the GC fraction. n = 6 biological replicates for protein and rRNA; n = 3 for mRNA. b, GC protection assay schema: bulk GC fraction isolated after subcellular fractionation is a suspension of GC particles enclosing GC-specific molecules (blue) within a medium that contains dilute soluble cytosolic molecules (red). Treatment with RNase or protease leads to hydrolysis of RNA and protein in the suspension medium not protected within GC particles, leaving only the GC-protected molecules (blue) after treatment. Addition of detergent before treatment results in hydrolysis of both cytosolic as well as GC-encapsulated molecules due to ruptures in the encapsulating GC plasma membrane, providing a control for the efficiency of enzymes. The difference in RNA or protein signal between hydrolysis control and GC-protected samples corresponds to the GC-encapsulated signal. c, d, GC protection assays with non-membrane-permeable degrading enzymes (protease in c and RNase in d) to test GC integrity and GC-specific membrane encapsulation of RNA and proteins in isolated GCs. Treatment with enzyme plus detergent, but neither alone, completely abolishes RNA and protein signal from GC fractions. Signals persisting in treatments with enzyme alone (lanes 3) correspond to RNA and protein encapsulated (protected) by the GC membrane, and correspond to the specific molecular content of isolated GCs. Treatment with protease alone has no effect on the signal from GAP43, confirming GC-specificity. Conversely, the signal from GM130, a Golgi matrix protein known to be excluded from GCs, is abolished with protease treatment alone, indicating there is no non-specific encapsulation in GCs. Reduced presence of both Actb and rRNA in samples treated with enzyme alone is consistent with their ubiquitous presence in both GCs and elsewhere in the homogenate. n = 5 independent biological replicates. e, Bioanalyzer profiles show GC-protected RNA compared to detergent-treated control, with characteristic peaks corresponding to 28S and 18S rRNA, and a spectrum of low intensity signal characteristic of mRNA. Experiments performed in n = 5 biological replicates with consistent results.
Extended Data Fig. 2 Quality control filtering of mass spectrometry measurements from sorted somata and GCs.
a, b, Multi-scatter plot of mass spectrometry signal intensity (log2-transformed label-free quantification (LFQ) of proteins) for each detected soma (a) and GC (b) protein in pairwise comparisons across six biological replicates. Quality control minimum stringency criteria were set based on average Pierson’s correlation coefficients across biological replicates. Soma samples displayed higher complexity than GC samples, which was reflected in the minimum acceptable correlation coefficients of 0.5 for somata (a) and 0.8 for GCs (b). All six soma replicates, and four out of six GC replicates met quality control criteria. Outlier GC samples (GC 3 and GC 6) are shaded grey (b).
Extended Data Fig. 3 Sorted GC–soma protein mass spectrometry intensities.
Scatter plot of paired protein intensities from trans-hemispheric sorted GC and sorted parent somata. Units represent log2-transformed peak-normalized intensities as measured by MaxLFQ45. Gene groups are coloured as indicated in the key. The GC marker GAP43 is indicated by a circled asterisk.
Extended Data Fig. 4 Sorted GC–soma proteome mapping.
Volcano plot of GC–soma proteome mapping, with log2-transformed λP values for each gene product plotted against significance (−log-transformed P values determined by two-tailed t-test). Significance thresholds were set to a 0.05 permutation-based FDR to indicate soma- and GC-specific mapping. n = 6 biological replicates, 3–6 litters each. Coloured gene groups are indicated. The proteome of sensorimotor cortex inter-hemispheric projection neurons distributes between cellular compartments with varying polarization that clusters with gene group, including GC-rich clusters (for example, proteasome), soma-rich clusters (for example, histones), and groups with moderate levels present in both GCs and somata, with moderate enrichments for one or the other compartment (for example, ribosomes and actin/tubulin).
Extended Data Fig. 5 Sorted GC mRNA-to-protein distribution.
Scatter plot pairing mRNA and corresponding protein relative abundance in trans-hemispheric-sorted GCs. Units represent log2-transformed peak-normalized mean intensities as measured by Sailfish TPM (mRNA) and MaxLFQ (protein) from 6 biological replicates of 3–6 litters each. Gene groups are indicated. The GC marker GAP43 is indicated by an asterisk. Trans-hemispheric GCs contain mRNA of select high-abundance GC proteins, as well as of most ribosomal protein mRNAs, whereas ribosomal proteins remain at GCs at moderate levels.
Extended Data Fig. 6 Subcellular RNA–proteome mapping.
2D RNA–proteome mapping of statistically significant enrichments (two-tailed t-test P value, significance threshold of 0.02 Benjamini–Hochberg corrected FDR, from 6 biological replicates of 3–6 litters each) in Gene Ontology (GO) terms and gene groups defined in Supplementary Table 7. Labels are displayed for non-redundant GO cell compartment terms and gene groups. Histograms along the x axis show RNA distributions of log2-transformed mean λR values. Histograms along the y axis on the right side show protein distributions of log2-transformed mean λP values of genes within each group. Four clusters emerge: the soma cluster (blue) contains groups of genes with both mRNA and protein enriched in the soma; the anterograde cluster (red) contains groups of genes with mRNA mapping to soma and protein to GC; the mito cluster (grey) contains mitochondrial genes with intermediate distributions; and the TOP cluster (green) exclusively contains TOP transcripts (including ribosomal protein genes), with mRNAs mapping to the GC and proteins to both GC and soma.
Extended Data Fig. 7 Subcellular transcriptome distribution follows mTOR dependence.
a, Sample transcripts from each class represented in Fig. 4 were verified by qPCR (y axis), and correlated with RNA-seq mapping values (x axis). Measurements from the two approaches displayed strong concurrence, with correlation coefficient R2 = 0.736. Centre points denote the mean, and error bars denote s.e.m. of three biological replicates from independent litters. Colour legend and schematics of mRNA classes based on known mTOR dependence: mRNAs containing a TOP motif are mTOR-dependent (green); schema indicates direct binding of the TOP motif by LARP1, which interacts with mTOR. mRNAs containing internal ribosome entry sites or lacking poly(A) tails are mTOR-independent (blue). Canonical mRNAs that undergo cap-dependent translation (grey) display moderate responses to mTOR. c, Expanded dataset presented in Fig. 4c. Single-molecule RNA chromogenic in situ hybridization (ISH, red) of two TOP transcripts: Rack1 (non-ribosomal TOP) and Rplp0 (ribosomal TOP), compared to a control transcript Ppib (soma-mapped canonical) in callosal projection neurons. Neurons were labelled with mem-GFP (green) via in utero electroporation at embryonic day (E) 15, cultured at P0, fixed and hybridized at in vitro day (DIV) 2–3. Nuclei were stained with DAPI (blue). Soma and GC close-ups shown in insets as overlays of transcript (red) with mem-GFP (green) in top rows, or with traced GC outlines in bottom rows. 92, 103 and 84 GCs imaged for Rack1, Rplp0 and Ppib probes, respectively, from n = 4 biological replicates from independent in utero electroporations. Five example GCs are shown per sample to capture the representative range. Scale bars, 10 μm.
Extended Data Fig. 8 Dense foci of mTOR in GCs.
a, Data relate to Fig. 5. Biochemical analysis of GC enrichment of mTOR pathway proteins and controls, shown in triplicate western blots of homogenate (input) and GC fraction pairs, derived from six independent preps. GAP43 is positive control for enrichment; GM130 is a negative control. Quantification as in Fig. 5. b, Close-ups of GCs from callosal projection neurons immunelabelled for endogenous mTOR pathway proteins (red in overlays, heat-mapped in underlying panels). Five example GCs are shown per sample to capture the representative range. Neurons were labelled via in utero electroporation at E15 with membrane-GFP (green in overlays, outlined in underlying panels), cultured at P0, fixed and stained at DIV2–DIV3. mTOR, LARP1, TSC1 and raptor (mTORC1 marker) appear in dense local foci within GCs. RICTOR (mTORC2 marker) and LAMP1 (lysosome marker) appear in fine granules distinct from GC foci. Bar (bottom right) indicates heat-map colour range, as well as 10-μm scale. 83 GCs imaged for mTOR, 47 for LARP1, 42 for LAMP1, 49 for TSC1, 26 for raptor, and 30 for RICTOR, from a minimum of n = 3 biological replicates from independent in utero electroporations.
Extended Data Fig. 9 GC-specific mTOR localization.
a, High-magnification views of representative GCs from callosal projection neurons immunolabelled for mTOR, equivalent to Fig. 5c and Extended Data Fig. 8b mTOR panels, but with a distinct mTOR antibody to independently confirm dense focal mTOR in GCs. Top, overlay images; bottom, heat maps of the same GCs. b, Example of mTOR labelling (red in two left panels; heat map in right panel) in 3-day-cultured neurons. A GFP-labelled axon from an electroporated inter-hemispheric neuron displays dense focal mTOR in its GC (arrow) compared to adjacent cell bodies (asterisks; DNA in blue indicates nuclei). Two other unlabelled GCs in the field can be recognized by virtue of their dense focal mTOR labelling alone (arrowheads). c, Example of dendritic GCs (arrows) lacking mTOR, juxtaposed to an unlabelled GC (arrowhead) with prominent mTOR focus. Bars in a–c (bottom right of each) indicate mTOR intensity heat-map colour range, as well as 10-μm scale. GCs imaged from n = 2 biological replicates from independent in utero electroporations.
Extended Data Fig. 10 mTOR signalling is required for the extension of trans-hemispheric axons.
a, Data relate to Fig. 6. Electroporation of callosal projection neurons with GFP and genetic payloads at E15, fixation and analysis at P3. Control electroporations (left column, grey in quantifications) show soma migration into upper layers (middle row insets, examples from four brains), and callosal projections well into the contralateral cortex (bottom insets, examples from four brains). Electroporation with PI3K-DN (middle column, green in quantifications) hinders migration of somata, with failure of callosal axon growth. Electroporation of Cre in mice with homozygous floxed-mTOR alleles for conditional mTOR gene knockout (mTOR-KO, right column, blue in quantifications) results specifically in failure of callosal axon growth. Scale bars, 100 μm. b, Quantification of the location of the electroporation field shows comparable mediolateral electroporation positions across all samples. Plotted are histograms of binned GFP intensities along the tangential axis of the ipsilateral cortex ending at the midline, as schematized above. c, Quantification of extent of migration, with percentage of somata in layers II/III (dark colours in graph) versus somata still en route in deeper layers (light colours in graph), as schematized above. Inhibition of PI3K signalling (green) interferes with migration, whereas acute mTOR deletion (blue) does not significantly affect migration. d, Quantification of callosal axon extension, showing that PI3K inhibition (green) as well as knockout of mTOR (blue) disrupt the formation of axon projection across the corpus callosum. Plotted are binned GFP intensity histograms within the corpus callosum from ipsilateral, through the midline (indicated by dotted line), to the contralateral side, as schematized above. Data are mean ± s.e.m. from n = 4 mice from different litters for each condition.
Supplementary information
Supplementary Table 1: Trans-hemispheric GC sub-proteome, {P}GC
List of proteins that are detected by mass-spec from sorted trans-hemispheric GCs. Columns correspond to gene names, protein names, theoretical protein molecular weight, and ENSEMBL gene ID.
Supplementary Table 2: GC-soma proteome mapping
List of protein GC-soma ratios λP. Columns correspond to gene names, λP values (log2), specific enrichment based on significance threshold set to 0.05 permutation-based false discovery rate (FDR), two-tailed t-test P values (-log), protein names, and gene group. n=6 biological replicates from 3-6 litters each.
Supplementary Table 3: Protein GC-soma annotation enrichment
List of significantly enriched Gene Ontology (GO) terms based on λP values. Columns correspond to term type (GO-Biological Function, GO-Cellular Component, and GO-Molecular Function), term name, number of genes, rank distribution score51, two-tailed t-test P value, significance based on Benjamini-Hochberg corrected FDR, and mean λP value.
Supplementary Table 4: GC-soma RNA mapping
List of mRNA GC-soma ratios λR. Columns correspond to gene names, λR values (log2), specific enrichment based on significance threshold set to 0.05 permutation-based FDR, two-tailed t-test P values (-log), gene names, and gene group. n=6 biological replicates from 3-6 litters each.
Supplementary Table 5: mRNA GC-soma annotation enrichment
List of significantly enriched Gene Ontology (GO) terms based on λR values. Columns correspond to term type (GO-Biological Function, GO-Cellular Component, and GO-Molecular Function), term name, number of genes, rank distribution score51, two-tailed t-test P value, significance threshold of 0.02 Benjamini-Hochberg corrected FDR, and mean λR value.
Supplementary Table 6: 2D RNA-proteome GC-soma annotation enrichment
List of significantly enriched Gene Ontology (GO) and gene group terms based on paired λR and λP values. Columns correspond to term type (GO-Biological Function, GO-Cellular Component, GO-Molecular Function, and gene group), term name, number of genes, two-tailed t-test P value, significance threshold of 0.02 Benjamini-Hochberg corrected FDR, mean λR, and mean λP values.
Supplementary Table 7: Gene groups
List of gene names and ENSEMBL gene IDs that constitute the annotated gene groups in Fig. 2, Fig. 4, Extended Data Fig.3-5, and Supplementary Table 6.
Rights and permissions
About this article
Cite this article
Poulopoulos, A., Murphy, A.J., Ozkan, A. et al. Subcellular transcriptomes and proteomes of developing axon projections in the cerebral cortex. Nature 565, 356–360 (2019). https://doi.org/10.1038/s41586-018-0847-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0847-y
This article is cited by
-
The role of TSC1 and TSC2 proteins in neuronal axons
Molecular Psychiatry (2024)
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
Cytosolic Ptbp2 modulates axon growth in motoneurons through axonal localization and translation of Hnrnpr
Nature Communications (2023)
-
RNA-Binding Proteins: A Role in Neurotoxicity?
Neurotoxicity Research (2023)
-
Neuronal subtype-specific growth cone and soma purification from mammalian CNS via fractionation and fluorescent sorting for subcellular analyses and spatial mapping of local transcriptomes and proteomes
Nature Protocols (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.