Abstract
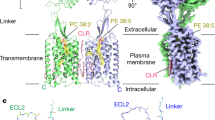
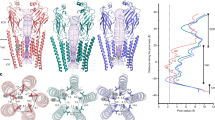
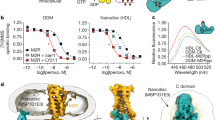
Type A γ-aminobutyric acid (GABAA) receptors are pentameric ligand-gated ion channels and the main drivers of fast inhibitory neurotransmission in the vertebrate nervous system1,2. Their dysfunction is implicated in a range of neurological disorders, including depression, epilepsy and schizophrenia3,4. Among the numerous assemblies that are theoretically possible, the most prevalent in the brain are the α1β2/3γ2 GABAA receptors5. The β3 subunit has an important role in maintaining inhibitory tone, and the expression of this subunit alone is sufficient to rescue inhibitory synaptic transmission in β1–β3 triple knockout neurons6. So far, efforts to generate accurate structural models for heteromeric GABAA receptors have been hampered by the use of engineered receptors and the presence of detergents7,8,9. Notably, some recent cryo-electron microscopy reconstructions have reported ‘collapsed’ conformations8,9; however, these disagree with the structure of the prototypical pentameric ligand-gated ion channel the Torpedo nicotinic acetylcholine receptor10,11, the large body of structural work on homologous homopentameric receptor variants12 and the logic of an ion-channel architecture. Here we present a high-resolution cryo-electron microscopy structure of the full-length human α1β3γ2L—a major synaptic GABAA receptor isoform—that is functionally reconstituted in lipid nanodiscs. The receptor is bound to a positive allosteric modulator ‘megabody’ and is in a desensitized conformation. Each GABAA receptor pentamer contains two phosphatidylinositol-4,5-bisphosphate molecules, the head groups of which occupy positively charged pockets in the intracellular juxtamembrane regions of α1 subunits. Beyond this level, the intracellular M3–M4 loops are largely disordered, possibly because interacting post-synaptic proteins are not present. This structure illustrates the molecular principles of heteromeric GABAA receptor organization and provides a reference framework for future mechanistic investigations of GABAergic signalling and pharmacology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sigel, E. & Steinmann, M. E. Structure, function, and modulation of GABAA receptors. J. Biol. Chem. 287, 40224–40231 (2012).
Olsen, R. W. & Sieghart, W. GABAA receptors: subtypes provide diversity of function and pharmacology. Neuropharmacology 56, 141–148 (2009).
Braat, S. & Kooy, R. F. The GABAA receptor as a therapeutic target for neurodevelopmental disorders. Neuron 86, 1119–1130 (2015).
Macdonald, R. L., Kang, J.-Q. & Gallagher, M. J. Mutations in GABAA receptor subunits associated with genetic epilepsies. J. Physiol. (Lond.) 588, 1861–1869 (2010).
Fritschy, J. M. & Mohler, H. GABAA receptor heterogeneity in the adult rat brain: differential regional and cellular distribution of seven major subunits. J. Comp. Neurol. 359, 154–194 (1995).
Nguyen, Q.-A. & Nicoll, R. A. The GABAA receptor β subunit is required for inhibitory transmission. Neuron 98, 718–725.e3 (2018).
Miller, P. et al. Heteromeric GABAA receptor structures in positively-modulated active states. Preprint at https://www.biorxiv.org/content/early/2018/06/05/338343 (2018).
Phulera, S. et al. Cryo-EM structure of the benzodiazepine-sensitive α1β1γ2S tri-heteromeric GABAA receptor in complex with GABA. eLife 7, 531 (2018).
Zhu, S. et al. Structure of a human synaptic GABAA receptor. Nature 559, 67–72 (2018).
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003).
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4 Å resolution. J. Mol. Biol. 346, 967–989 (2005).
Nemecz, Á., Prevost, M. S., Menny, A. & Corringer, P.-J. Emerging molecular mechanisms of signal transduction in pentameric ligand-gated ion channels. Neuron 90, 452–470 (2016).
Dostalova, Z. et al. Human α1β3γ2L gamma-aminobutyric acid type A receptors: high-level production and purification in a functional state. Protein Sci. 23, 157–166 (2014).
Masiulis, S. et al. GABAA receptor signalling mechanisms revealed by structural pharmacology. Nature https://doi.org/10.1038/s41586-018-0832-5 (2018).
Gielen, M., Thomas, P. & Smart, T. G. The desensitization gate of inhibitory Cys-loop receptors. Nat. Commun. 6, 6829 (2015).
Papke, D. & Grosman, C. The role of intracellular linkers in gating and desensitization of human pentameric ligand-gated ion channels. J. Neurosci. 34, 7238–7252 (2014).
Hille, B., Dickson, E. J., Kruse, M., Vivas, O. & Suh, B.-C. Phosphoinositides regulate ion channels. Biochim. Biophys. Acta 1851, 844–856 (2015).
Hansen, S. B., Tao, X. & MacKinnon, R. Structural basis of PIP2 activation of the classical inward rectifier K+ channel Kir2.2. Nature 477, 495–498 (2011).
Mennerick, S., Taylor, A. A. & Zorumski, C. F. Phosphatidylinositol 4,5-bisphosphate depletion fails to affect neurosteroid modulation of GABAA receptor function. Psychopharmacology (Berl.) 231, 3493–3501 (2014).
Amin, J. & Weiss, D. S. GABAA receptor needs two homologous domains of the β subunit for activation by GABA but not by pentobarbital. Nature 366, 565–569 (1993).
Hibbs, R. E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011).
Miller, P. S. & Aricescu, A. R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Laverty, D. et al. Crystal structures of a GABAA-receptor chimera reveal new endogenous neurosteroid-binding sites. Nat. Struct. Mol. Biol. 24, 977–985 (2017).
Miller, P. S. et al. Structural basis for GABAA receptor potentiation by neurosteroids. Nat. Struct. Mol. Biol. 24, 986–992 (2017).
Auerbach, A. & Akk, G. Desensitization of mouse nicotinic acetylcholine receptor channels. A two-gate mechanism. J. Gen. Physiol. 112, 181–197 (1998).
Bali, M. & Akabas, M. H. The location of a closed channel gate in the GABAA receptor channel. J. Gen. Physiol. 129, 145–159 (2007).
Forman, S. A. & Miller, K. W. Mapping general anesthetic sites in heteromeric γ-aminobutyric acid type A receptors reveals a potential for targeting receptor subtypes. Anesth. Analg. 123, 1263–1273 (2016).
Li, G.-D. et al. Identification of a GABAA receptor anesthetic binding site at subunit interfaces by photolabeling with an etomidate analog. J. Neurosci. 26, 11599–11605 (2006).
Jurd, R. et al. General anesthetic actions in vivo strongly attenuated by a point mutation in the GABAA receptor β3 subunit. FASEB J. 17, 250–252 (2003).
Jayakar, S. S. et al. Positive and negative allosteric modulation of an α1β3γ2 γ-aminobutyric acid type A (GABAA) receptor by binding to a site in the transmembrane domain at the γ+/β− interface. J. Biol. Chem. 290, 23432–23446 (2015).
Tretter, V. et al. Gephyrin, the enigmatic organizer at GABAergic synapses. Front. Cell. Neurosci. 6, 23 (2012).
Unwin, N. Segregation of lipids near acetylcholine-receptor channels imaged by cryo-EM. IUCrJ 4, 393–399 (2017).
Ritchie, T. K. et al. Reconstitution of membrane proteins in phospholipid bilayer nanodiscs. Methods Enzymol. 464, 211–231 (2009).
Javaheri, A. et al. Helicobacter pylori adhesin HopQ engages in a virulence-enhancing interaction with human CEACAMs. Nat. Microbiol. 2, 16189 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. W. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012).
Vilas, J. L. et al. MonoRes: automatic and accurate estimation of local resolution for electron microscopy maps. Structure 26, 337–344.e4 (2018).
de la Rosa-Trevín, J. M. et al. Scipion: A software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 195, 93–99 (2016).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
McNicholas, S., Potterton, E., Wilson, K. S. & Noble, M. E. M. Presenting your structures: the CCP4mg molecular-graphics software. Acta Crystallogr. D 67, 386–394 (2011).
Jurcik, A. et al. CAVER Analyst 2.0: analysis and visualization of channels and tunnels in protein structures and molecular dynamics trajectories. Bioinformatics 34, 3586–3588 (2018).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Dostalova, Z. et al. High-level expression and purification of Cys-loop ligand-gated ion channels in a tetracycline-inducible stable mammalian cell line: GABAA and serotonin receptors. Protein Sci. 19, 1728–1738 (2010).
Desai, R. et al. Contrasting actions of a convulsant barbiturate and its anticonvulsant enantiomer on the α1β3γ2L GABAA receptor account for their in vivo effects. J. Physiol. (Lond.) 593, 4943–4961 (2015).
Forman, S. A. A hydrophobic photolabel inhibits nicotinic acetylcholine receptors via open-channel block following a slow step. Biochemistry 38, 14559–14564 (1999).
Yuan, H., Low, C.-M., Moody, O. A., Jenkins, A. & Traynelis, S. F. Ionotropic GABA and glutamate receptor mutations and human neurologic diseases. Mol. Pharmacol. 88, 203–217 (2015).
Carvill, G. L. et al. GABRA1 and STXBP1: novel genetic causes of Dravet syndrome. Neurology 82, 1245–1253 (2014).
Lachance-Touchette, P. et al. Novel α1 and γ2 GABAA receptor subunit mutations in families with idiopathic generalized epilepsy. Eur. J. Neurosci. 34, 237–249 (2011).
Kash, T. L., Jenkins, A., Kelley, J. C., Trudell, J. R. & Harrison, N. L. Coupling of agonist binding to channel gating in the GABAA receptor. Nature 421, 272–275 (2003).
Epi4K Consortium & Epilepsy Phenome/Genome Project De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Hernandez, C. C. et al. GABAA receptor coupling junction and pore GABRB3 mutations are linked to early-onset epileptic encephalopathy. Sci. Rep. 7, 15903 (2017).
Hernandez, C. C. et al. Altered channel conductance states and gating of GABAA receptors by a pore mutation linked to Dravet syndrome. eNeuro 4, ENEURO.0251–16.2017 (2017).
Yip, G. M. S. et al. A propofol binding site on mammalian GABAA receptors identified by photolabeling. Nat. Chem. Biol. 9, 715–720 (2013).
Acknowledgements
We thank G. Cannone and S. Chen for electron microscopy support; J. Grimmett and T. Darling for computing support; J. García-Nafría, L. Dong, T. Nakane and S. Scheres for advice on electron microscopy data processing; and members of the Aricescu laboratory for assistance with electron microscopy data collection, discussions and comments on the manuscript. This work was supported by the UK Medical Research Council grants MR/L009609/1, MC_UP_1201/15 (A.R.A., D.L. and S.M.) and MC_UP_A025_1013 (J.Z.); UK Biotechnology and Biological Sciences Research Council grant BB/M024709/1 (A.R.A. and D.L.); Human Frontier Science Program grant RGP0065/2014 (A.R.A.); Cancer Research UK grant C20724/A14414 (T.M.); and Swiss National Science Foundation fellowship 168735 (J.Z.). R.D., W.J.S. and K.W.M. were supported by a grant from the National Institute for General Medical Sciences (GM 58448) and by the Department of Anesthesia, Critical Care and Pain Medicine, Massachusetts General Hospital. We acknowledge the support and the use of resources of Instruct-ERIC (PID1271), part of the European Strategy Forum on Research Infrastructures (ESFRI), and the Research Foundation-Flanders (FWO) for their support of nanobody discovery, and FWO for a doctoral fellowship to T.U.
Reviewer information
Nature thanks G. Akk, A. Evers, M. Jansen and H. Nury for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
D.L. and A.R.A. conceived the project. D.L. carried out protein purification, collected and processed the electron microscopy data with assistance from S.M., and built and refined the model with assistance from A.R.A. R.D. and K.W.M. designed and analysed the electrophysiological experiments, which were performed by R.D. T.U., E.P. and J.S. designed and generated Mb38. S.M. developed the nanodisc reconstitution protocols. T.M. performed small-molecule docking. J.Z. developed contrast transfer function refinement algorithms. W.J.S. and K.W.M. designed and analysed the binding experiments, which were performed by W.J.S. D.L. and A.R.A. wrote the manuscript, with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM image processing procedure.
a, Overview of cryo-EM data collection and image-processing procedure (see Methods). b, Representative micrograph of the α1β3γ2L–Mb38–nanodisc complex obtained using the Falcon3 detector and Volta phase plate. c, Representative 2D class averages for downscaled data (box size of 240 Å). d, FSC curves for the reconstruction before and after applying a soft mask. e, The unsharpened map from refinement coloured by local resolution estimate (calculated using MonoRes38) and shown at a low isosurface level to enable visualization of the nanodisc (top) and at a high isosurface level (bottom). f, Angular-distribution histogram of particles used in calculating the final 3D reconstruction for the receptor assembly. g, FSC curves for cross-validation between maps and models: model versus summed map (black), model versus half map 1 (used in test refinement, green), model versus half map 2 (not used in test refinement, blue).
Extended Data Fig. 2 Sequence alignment of GABAA receptor α1, β3 and γ2 subunits, biochemical characterization and binding assays.
a, Alignment of wild-type GABAA receptor subunit sequences, in which 1 represents the first residue of the mature protein. α-helices (grey cylinders), β-strands (black arrows) and associated loops are indicated. Glycosylation sites are indicated by a blue pentagon and the associated subunit residue is highlighted in blue. Residues identified as coordinating PIP2 binding are highlighted in yellow and are indicated by yellow hexagons. The alignment graphic was prepared on the ESPript 3.0 server (http://espript.ibcp.fr/ESPript/ESPript/). b, Structure of a single α1 subunit. c, Western blot analysis of cell lysates from LMNG-solubilized control HEK293 cells and α1β3γ2L GABAA receptor cells, and purified α1β3γ2L GABAA receptors in nanodiscs. The arrowhead denotes the band corresponding to the full-length GABAA receptor subunits, which migrates as a species of about 51–55 kDa. With the exception of the α1 subunit (which displays a small degree of proteolysis following reconstitution, denoted by an asterisk), GABAA receptor subunits do not display apparent proteolysis during solubilization, purification and reconstitution. Western blots were repeated twice independently with similar results. d, GABA enhanced displaceable [3H]flunitrazepam binding to purified receptors in a concentration-dependent manner in the presence or absence of Mb38. Points represent individual samples from two separate experiments.
Extended Data Fig. 3 α1β3γ2 model-map validation and electron microscopy density.
a–m, Electron microscopy density segments for representative regions of each subunit and Mb38. Sharpened map contoured as detailed in Methods.
Extended Data Fig. 4 Comparison of TMD architecture with α1β2γ2 and α1β1γ2 GABAA receptor structures in detergent.
a–f, Superposition of subunit chains of α1β3γ2L GABAA receptor (grey) onto equivalent chains of α1β2γ2 GABAA receptor in DDM (conformation B; PDB ID: 6D6T) and the γ2 subunit of α1β2γ2 GABAA receptor (conformation A; PDB ID: 6D6U). R.m.s.d. values are for Cα atoms over the entire subunit. g, Overview of the TMD of α1β3γ2 in nanodisc. h–j, Superposition of TMD for α1β2γ2 GABAA receptor conformation A (h), conformation B (i), α1β1γ2 GABAA receptor (PDB ID: 6DW0; j) and the α1β3γ2–nanodisc complex (grey ribbon). Transmembrane helices of the γ2 subunit are labelled. The transmembrane helices of the γ2 show considerable distortion in detergent-bound complexes. M4 helices in α1 and γ2 subunits were not modelled in the α1β1γ2 GABAA receptor. k–n, Superposition of β− subunits reveals conformational differences of the γ2+ subunit (α1β3γ2–nanodisc complex in grey). Differences in distance (∆) between selected residue Cα atoms (spheres) are indicated by black lines. Disruption of the γ2 TMD induces substantial displacement of loop 7, loop 2 and the M2–M3 loop at the ECD–TMD interface in the detergent-bound α1β2γ2 structures (l, m), and to a lesser extent in α1β1γ2 GABAA receptor (n). o–r, Close-up view of M2 helices at the level of −2ʹ proline or alanine residues (Cα atoms shown as spheres) in nanodisc (o) and detergent-bound structures (p–r).
Extended Data Fig. 5 Mb38 binding and function.
a, b, Side (a) and top-down (b) views of neighbouring α1+ and β3− subunits bound to Mb38. c, Representative normalized current traces obtained in cells expressing the α1β3γ2L receptor exposed to GABA (3 μM) alone or with varying concentrations of Mb38 (n = 3–6 cells), applied for 4 s. Currents were normalized to peak-current amplitude obtained with GABA (3 μM) alone during the first 1-s phase of the trace. The concentration of Mb38 is indicated above each trace. d–f, Close-up view of the binding site when viewed approximately parallel to the plane of the membrane. Complementarity-determining-region loops 1 (d), 3 (e) and 2 (f) of the Mb38 are coloured in turquoise, dark green and teal, respectively, and residues involved in interactions are shown in ball-and-stick representation. Polar interactions are shown as dotted lines. g, Representative current trace obtained in cells expressing the α1β3γ2L receptor exposed to 3 μM Mb38. Mb38 (3 μM) opened 16 ± 11% (mean ± s.d.; n = 5 cells) of the receptors gated by EC10 GABA alone (therefore around 2% of the total receptors expressed).
Extended Data Fig. 6 Vestibular glycans and interface classes in the α1β3γ2L GABAA receptor.
a, Side view of the receptor shows the position of vestibular α1 N-linked glycans. For clarity, the near α1 and β3 subunits have been removed. b, View across the extracellular vestibule reveals the stacking of α1 N-linked glycans. The receptor surface is coloured according to electrostatic surface potential and reveals an electropositive ring in the middle portion of the ECD vestibule. c–f, Paired views of the interface between principle (+) and complementary (−) subunits viewed from the pore axis outwards (left) and open-book view of each subunit when viewed from the receptor periphery (right). Residues involved in forming interactions (defined using PDBePISA45) are coloured according to the type of interaction and mapped onto the isosurface representation: polar, cyan; electrostatic/salt bridges, magenta; van der Waals, orange. Arrowheads reveal the openings of defined tunnels between adjacent subunits. g, Calculated interfacial buried surface areas and solvation energy gain at complex formation (both calculated using PDBePISA44). The asterisk denotes the second β3/α1 (chain E/chain A) interface in the pentameric assembly. Radii of tunnels, denoted by arrowheads in c–f, were also calculated (see Methods). Open arrowheads in c, e denote cavities forming the proposed anaesthetic-binding sites.
Extended Data Fig. 7 Disease mutations associated with α1, β3 and γ2, lateral tunnels and fenestrations at the subunit interfaces.
a, b, Disease mutations associated with GABAA α1, β3 and γ2 subunits are mapped onto the structure and shown in sphere representation. The receptor is viewed parallel to the membrane plane (a) and from the extracellular aspect (b). Outlined boxes highlight the position of mutations shown in c–e. c–e, Close-up view of disease mutations associated with the α1 and β3 subunits. Polar interactions between residues are shown as dotted lines. f, Table summarizing several disease mutations identified in genes for α1, β3 and γ2. Functional effects as determined from experimental studies52,53,54,55,56,57,58 of channel properties are summarized. g, Exposed surface of the γ2/β3 subunit interface coloured according to electrostatic surface potential. h, Close-up view of an electronegative fenestration formed at the γ2/β3 extracellular domain interface. The continuous tunnel leading from extracellular space to the receptor vestibule is outlined. i, Exposed surface of the α1/β3 subunit interface coloured according to electrostatic surface potential. j, Close-up view of the α1/β3 extracellular interface reveals an upper tunnel leading to the mid-portion of the ECD vestibule. A lower tunnel (denoted by the arrow) opens into the upper aspect of the ion channel at the level of β3His267, a residue that is implicated in mediating the effects of propofol59.
Extended Data Fig. 8 Lipid-binding sites and functional modulation of GABAA receptor by PIP2.
a, Well-resolved density for the POPC lipid moiety (yellow, ball-and-stick representation) at the extracellular aspect of the lipid nanodisc. Electron microscopy density is shown in chicken-wire representation and contoured around the lipids. b, Sequence alignment of GABAA receptor and GlyR subunits for PIP2-binding regions: the M1–M2 loop, post-M3 and pre-M4 segments. α1 residues forming hydrogen bonds or salt-bridge interactions with PIP2 are identified by yellow hexagons, and those that are conserved among receptor subunits are highlighted in orange (identical) and yellow (similar). The alignment graphic was prepared on the ESPript 3.0 server (http://espript.ibcp.fr/ESPript/ESPript/). c, Representative normalized current traces from the same patch, obtained in a two-pulse protocol, in which inside-out patches were exposed to two 5-s etomidate (100 μM) pulses, 7.5 s apart. During the second pulse, etomidate was either applied alone or co-applied with poly-l-lysine (250 μg ml−1). Current traces were normalized to the peak-current amplitude obtained during the first etomidate pulse. d, Dot plot of peak-current amplitudes obtained during the second pulse (co-application of poly-l-lysine) normalized to the peak-current amplitudes obtained with the first pulse of etomidate (centre value represents mean ± s.d.; n = 9 patches). Unpaired and paired Student’s t-tests (two-tailed) were used; the P values obtained are indicated on the figure.
Extended Data Fig. 9 Comparisons of agonist sites and analogous pockets at other subunit interfaces.
a, View of the receptor from across the synaptic cleft with the agonist-binding sites highlighted. b, Cut-away view of a at the level of the loops-C reveals electron microscopy density (shown as magenta chicken-wire representation) at four inter-subunit pockets. c, Electron microscopy density in the orthosteric binding-pocket (grey surface representation). For comparison, the top binding conformation for GABA is displayed in grey ball-and-stick representation. d, e, Overlay of GABA binding conformations from molecular-docking calculations at the β3/α1 (d) and α1/β3 (e) binding pockets. The range of estimated free energies of binding (in kcal mol−1) is given. f, Comparison of the orthosteric binding-pocket at the β3/α1 interface (grey), with the three unique interfaces observed in the α1β3γ2–Mb38 receptor complex (coloured as in a). Superposition of the (−) subunit ECD reveals the relative movement of the (+) subunit ECD. g, h, Modelling of the intracellular end of M3 and M4 helices, contributing to the receptor intracellular domain, shown approximately parallel (g) and perpendicular (h) to the plane of the membrane.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References and the uncropped western blots. The Supplementary Discussion includes descriptions of the Mb38 megabody binding site and of inter-subunit interfaces. Determinants of heteromeric GABAAR assembly are also discussed
Video 1
: Cryo-EM map of the α1β3γ2 GABAAR in a lipid bilayer. This video first shows the sharpened EM density map for the GABAAR (coloured by subunit, α red; β blue; γ yellow) in a lipid nanodisc (pale blue) and in complex with Mb38 (green) parallel to the plane of the membrane. Bound lipids are coloured in pale brown. Slices through the map are then shown from the extracellular aspect through to the cytoplasmic side. EM densities in the orthosteric and analogous sites are coloured in magenta
Video 2
: Atomic model of the α1β3γ2 GABAAR and details of a PIP2 binding site. This video first shows the receptor model (coloured by subunit; ribbon representation) and bound lipid molecules (yellow ball-and-stick representation). The view then pans in to highlight the binding loci for PIP2 at the base of the α1 subunits. In rotating around the vertical axis, the video highlights how the C4 and C5 phosphate groups of PIP2 form an extensive network of interactions with α1-residues in the post-M3 and pre-M4 segments
Rights and permissions
About this article
Cite this article
Laverty, D., Desai, R., Uchański, T. et al. Cryo-EM structure of the human α1β3γ2 GABAA receptor in a lipid bilayer. Nature 565, 516–520 (2019). https://doi.org/10.1038/s41586-018-0833-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0833-4
This article is cited by
-
Lipid nanodisc scaffold and size alter the structure of a pentameric ligand-gated ion channel
Nature Communications (2024)
-
State-dependent binding of cholesterol and an anionic lipid to the muscle-type Torpedo nicotinic acetylcholine receptor
Communications Biology (2024)
-
EMC rectifies the topology of multipass membrane proteins
Nature Structural & Molecular Biology (2024)
-
Neurosteroids and their potential as a safer class of general anesthetics
Journal of Anesthesia (2024)
-
Applications and prospects of cryo-EM in drug discovery
Military Medical Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.