Abstract
Aberrant cleavage of Notch by γ-secretase leads to several types of cancer, but how γ-secretase recognizes its substrate remains unknown. Here we report the cryo-electron microscopy structure of human γ-secretase in complex with a Notch fragment at a resolution of 2.7 Å. The transmembrane helix of Notch is surrounded by three transmembrane domains of PS1, and the carboxyl-terminal β-strand of the Notch fragment forms a β-sheet with two substrate-induced β-strands of PS1 on the intracellular side. Formation of the hybrid β-sheet is essential for substrate cleavage, which occurs at the carboxyl-terminal end of the Notch transmembrane helix. PS1 undergoes pronounced conformational rearrangement upon substrate binding. These features reveal the structural basis of Notch recognition and have implications for the recruitment of the amyloid precursor protein by γ-secretase.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM maps of the structure of human γ-secretase crosslinked to Notch-100 have been deposited in the Electron Microscopy Data Bank (EMDB) with accession code EMD-9648. The atomic coordinates for the corresponding model has been deposited in the Protein Data Bank (PDB) under accession code 6IDF.
References
Brown, M. S., Ye, J., Rawson, R. B. & Goldstein, J. L. Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100, 391–398 (2000).
Rawson, R. B. et al. Complementation cloning of S2P, a gene encoding a putative metalloprotease required for intramembrane cleavage of SREBPs. Mol. Cell 1, 47–57 (1997).
Urban, S., Lee, J. R. & Freeman, M. Drosophila rhomboid-1 defines a family of putative intramembrane serine proteases. Cell 107, 173–182 (2001).
Sherrington, R. et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer’s disease. Nature 375, 754–760 (1995).
Weihofen, A., Binns, K., Lemberg, M. K., Ashman, K. & Martoglio, B. Identification of signal peptide peptidase, a presenilin-type aspartic protease. Science 296, 2215–2218 (2002).
Manolaridis, I. et al. Mechanism of farnesylated CAAX protein processing by the intramembrane protease Rce1. Nature 504, 301–305 (2013).
Li, Y. M. et al. Photoactivated γ-secretase inhibitors directed to the active site covalently label presenilin 1. Nature 405, 689–694 (2000).
Li, Y. M. et al. Presenilin 1 is linked with γ-secretase activity in the detergent solubilized state. Proc. Natl Acad. Sci. USA 97, 6138–6143 (2000).
Kimberly, W. T. et al. γ-Secretase is a membrane protein complex comprised of presenilin, nicastrin, Aph-1, and Pen-2. Proc. Natl Acad. Sci. USA 100, 6382–6387 (2003).
De Strooper, B. et al. A presenilin-1-dependent γ-secretase-like protease mediates release of Notch intracellular domain. Nature 398, 518–522 (1999).
De Strooper, B. et al. Deficiency of presenilin-1 inhibits the normal cleavage of amyloid precursor protein. Nature 391, 387–390 (1998).
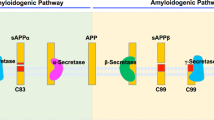
Golde, T. E., Estus, S., Younkin, L. H., Selkoe, D. J. & Younkin, S. G. Processing of the amyloid protein precursor to potentially amyloidogenic derivatives. Science 255, 728–730 (1992).
Takami, M. et al. γ-Secretase: successive tripeptide and tetrapeptide release from the transmembrane domain of β-carboxyl terminal fragment. J. Neurosci. 29, 13042–13052 (2009).
Suzuki, N. et al. An increased percentage of long amyloid β protein secreted by familial amyloid β protein precursor (βAPP717) mutants. Science 264, 1336–1340 (1994).
Hardy, J. A. & Higgins, G. A. Alzheimer’s disease: the amyloid cascade hypothesis. Science 256, 184–185 (1992).
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
Struhl, G., Fitzgerald, K. & Greenwald, I. Intrinsic activity of the Lin-12 and Notch intracellular domains in vivo. Cell 74, 331–345 (1993).
Kopan, R. & Ilagan, M. X. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137, 216–233 (2009).
Weng, A. P. et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306, 269–271 (2004).
Wang, B. et al. γ-Secretase gene mutations in familial acne inversa. Science 330, 1065 (2010).
Crump, C. J., Johnson, D. S. & Li, Y. M. Development and mechanism of γ-secretase modulators for Alzheimer’s disease. Biochemistry 52, 3197–3216 (2013).
Doody, R. S. et al. A phase 3 trial of semagacestat for treatment of Alzheimer’s disease. N. Engl. J. Med. 369, 341–350 (2013).
Bolduc, D. M., Montagna, D. R., Seghers, M. C., Wolfe, M. S. & Selkoe, D. J. The amyloid-beta forming tripeptide cleavage mechanism of γ-secretase. eLife 5, e17578 (2016).
De Strooper, B. & Chavez Gutierrez, L. Learning by failing: ideas and concepts to tackle γ-secretases in Alzheimer’s disease and beyond. Annu. Rev. Pharmacol. Toxicol. 55, 419–437 (2015).
Johnson, D. S., Li, Y. M., Pettersson, M. & St George-Hyslop, P. H. Structural and chemical biology of presenilin complexes. Cold Spring Harb. Perspect. Med. 7, a024067 (2017).
Lu, P. et al. Three-dimensional structure of human γ-secretase. Nature 512, 166–170 (2014).
Sun, L. et al. Structural basis of human γ-secretase assembly. Proc. Natl Acad. Sci. USA 112, 6003–6008 (2015).
Bai, X. C. et al. An atomic structure of human γ-secretase. Nature 525, 212–217 (2015).
Bolduc, D. M., Montagna, D. R., Gu, Y., Selkoe, D. J. & Wolfe, M. S. Nicastrin functions to sterically hinder γ-secretase-substrate interactions driven by substrate transmembrane domain. Proc. Natl Acad. Sci. USA 113, E509–E518 (2016).
Wolfe, M. S. et al. Two transmembrane aspartates in presenilin-1 required for presenilin endoproteolysis and γ-secretase activity. Nature 398, 513–517 (1999).
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015).
Takagi-Niidome, S. et al. Cooperative roles of hydrophilic loop 1 and the C-terminus of presenilin 1 in the substrate-gating mechanism of γ-secretase. J. Neurosci. 35, 2646–2656 (2015).
Crescenzi, O. et al. Solution structure of the Alzheimer amyloid β-peptide (1–42) in an apolar microenvironment. Similarity with a virus fusion domain. Eur. J. Biochem. 269, 5642–5648 (2002).
Deatherage, C. L. et al. Structural and biochemical differences between the Notch and the amyloid precursor protein transmembrane domains. Sci. Adv. 3, e1602794 (2017).
Nadezhdin, K. D., Bocharova, O. V., Bocharov, E. V. & Arseniev, A. S. Structural and dynamic study of the transmembrane domain of the amyloid precursor protein. Acta Naturae 3, 69–76 (2011).
Wahrle, S. et al. Cholesterol-dependent γ-secretase activity in buoyant cholesterol-rich membrane microdomains. Neurobiol. Dis. 9, 11–23 (2002).
Holmes, O., Paturi, S., Ye, W., Wolfe, M. S. & Selkoe, D. J. Effects of membrane lipids on the activity and processivity of purified γ-secretase. Biochemistry 51, 3565–3575 (2012).
Sun, L., Zhou, R., Yang, G. & Shi, Y. Analysis of 138 pathogenic mutations in presenilin-1 on the in vitro production of Aβ42 and Aβ40 peptides by γ-secretase. Proc. Natl Acad. Sci. USA 114, E476–E485 (2017).
Sato, C., Takagi, S., Tomita, T. & Iwatsubo, T. The C-terminal PAL motif and transmembrane domain 9 of presenilin 1 are involved in the formation of the catalytic pore of the γ-secretase. J. Neurosci. 28, 6264–6271 (2008).
Feng, L. et al. Structure of a site-2 protease family intramembrane metalloprotease. Science 318, 1608–1612 (2007).
Wu, Z. et al. Structural analysis of a rhomboid family intramembrane protease reveals a gating mechanism for substrate entry. Nat. Struct. Mol. Biol. 13, 1084–1091 (2006).
Baker, R. P., Young, K., Feng, L., Shi, Y. & Urban, S. Enzymatic analysis of a rhomboid intramembrane protease implicates transmembrane helix 5 as the lateral substrate gate. Proc. Natl Acad. Sci. USA 104, 8257–8262 (2007).
Vinothkumar, K. R. et al. The structural basis for catalysis and substrate specificity of a rhomboid protease. EMBO J. 29, 3797–3809 (2010).
Zoll, S. et al. Substrate binding and specificity of rhomboid intramembrane protease revealed by substrate–peptide complex structures. EMBO J. 33, 2408–2421 (2014).
Cho, S., Dickey, S. W. & Urban, S. Crystal structures and inhibition kinetics reveal a two-stage catalytic mechanism with drug design implications for rhomboid proteolysis. Mol. Cell 61, 329–340 (2016).
Vinothkumar, K. R., Pierrat, O. A., Large, J. M. & Freeman, M. Structure of rhomboid protease in complex with β-lactam inhibitors defines the S2′ cavity. Structure 21, 1051–1058 (2013).
Vosyka, O. et al. Activity-based probes for rhomboid proteases discovered in a mass spectrometry-based assay. Proc. Natl Acad. Sci. USA 110, 2472–2477 (2013).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S. H. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Bartesaghi, A. et al. 2.2 Å resolution cryo-EM structure of β-galactosidase in complex with a cell-permeant inhibitor. Science 348, 1147–1151 (2015).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr Protoc. Bioinformatics 5, 5–6 (2010).
Acknowledgements
We thank the Tsinghua University Branch of China National Center for Protein Sciences (Beijing) for the cryo-EM facility and the computational facility support, X. Li and X. Hu for technical support in EM data acquisition. This work was supported by funds from the National Natural Science Foundation of China (31621092). G.Y. and R.Z. are supported by postdoctoral fellowships of the Tsinghua–Peking Joint Center for Life Sciences and Beijing Advanced Innovation Center for Structural Biology.
Reviewer information
Nature thanks S. Blacklow, M. Freeman and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
G.Y., R.Z. and Y.S. conceived the project. G.Y., R.Z. and X.G. prepared the sample. G.Y., R.Z. and J.L. collected the EM data. G.Y. and Q.Z. processed the EM data. G.Y. and C.Y. built the atomic model. G.Y. and R.Z. designed and analysed the biochemical experiments. G.Y., R.Z. and X.G. performed the assays. M.K. helped with the APP-C99 modelling. G.Y., R.Z. and Y.S. wrote the manuscript. Y.S. supervised the project.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Discovery of an optimal crosslinking site between PS1 and Notch-100 for formation of a stable γ-secretase–Notch complex.
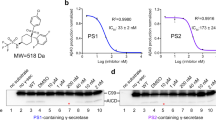
a, A schematic diagram of the screening procedure for an optimal crosslinking site between human γ-secretase and Notch-100. Sixteen γ-secretase variants were generated and tested for their crosslinking efficiencies. In each variant, a specific residue from Gly111-Gln112-Leu113-Ile114 of PS1 and a specific residue from Pro1728-Pro1729-Pro1730-Pro1731 of Notch were mutated to Cys. b, Examination of crosslinking efficiency by western blot analysis. In the absence of the reducing agent DTT, the intensity ratio of the crosslinked band (PS1-NTF + Notch-100) over PS1 was estimated to reflect the crosslinking efficiency. Q112C (in PS1) and P1728C (in Notch-100) were selected as the optimal crosslinking sites. c, PS1(Q112C) does not have a pronounced effect on the cleavage of Notch-100 by human γ-secretase. The amount of NICD generated by γ-secretase with PS1(Q112C) is similar to that produced by wild-type γ-secretase. d, The presence of an methionine residue at the amino terminus of Notch-100 does not have a pronounced effect on its cleavage efficiency by γ-secretase. e, The substrate Notch-100 is crosslinked to PS1 of human γ-secretase through an engineered disulfide bond. This disulfide bond involves two mutations on the extracellular side: PS1(Q112C) and Notch(P1728C). The enzyme–substrate complex was purified by affinity chromatography and gel filtration (see Methods). f, The purified γ-secretase–Notch-100 complex was visualized by SDS–PAGE with Coomassie staining. The crosslinked complex between PS1-NTF and Notch-100 can be reduced by DTT in vitro, generating free PS1-NTF. g, Analysis of the γ-secretase–Notch-100 complex by gel filtration. A representative chromatogram is shown (top), and the peak fractions were visualized by SDS–PAGE and Coomassie staining (bottom). PS1-NTF had been mostly crosslinked to Notch-100. Purification trials were repeated two times.
Extended Data Fig. 2 A flowchart for cryo-EM data processing.
The final average resolution for the entire human γ-secretase–Notch-100 complex is estimated to be 2.7 Å. For details, please refer to the Methods.
Extended Data Fig. 3 Cryo-EM analysis of the complex of human γ-secretase and Notch-100.
a, FSC curve for the 3D reconstruction of the cryo-EM map. The average resolution is estimated to be 2.7 Å on the basis of the FSC value of 0.143. b, Angular distribution of the particles used for reconstruction of the γ-secretase–Notch complex. Each cylinder represents one view and the height of the cylinder is proportional to the number of particles for that view. c, FSC curves of the refined model versus the overall 2.7 Å map that it was refined against (black); of the model refined in the first of the two independent maps used for the FSC calculation versus that same map (red); and of the model refined in the first of the two independent maps versus the second independent map (green). The small difference between the red and green curves indicates that the refinement of the atomic coordinates did not suffer from overfitting. d, Colour-coded local resolution distribution in Å of the final reconstruction as estimated by RELION-2.052.
Extended Data Fig. 4 Local cryo-EM density maps for representative regions of the γ-secretase–Notch-100 complex.
a, The local cryo-EM density maps for all nine transmembrane helices of PS1. The density for the side chains of TM2 is of sufficient quality for assignment of specific amino acids. TM6 is broken into two helices connected by a rigid coil. The sequences preceding TM7 form a β-strand in the presence of the substrate Notch-100. The contour level of the cryo-EM density for TM2 and TM6 of PS1 is 5σ. b, The local cryo-EM density maps for all seven transmembrane helices of APH-1. c, The local cryo-EM density map for the transmembrane helix of Notch-100. The cryo-EM density for the connecting loop of the transmembrane helix and the amino terminus is weak (left, 2.7 Å map, contour level 5σ), but can be improved by further classification and refinement (right, 3.3 Å map, contour level 8σ, map refers to Extended Data Fig. 8). d, The local cryo-EM density maps for the only transmembrane helix and three selected regions of NCT. e, The local cryo-EM density maps for the three transmembrane helices of PEN-2. The contour level of the cryo-EM density is 8σ, except where specifically stated in a and c.
Extended Data Fig. 5 Overall structure of the complex of human γ-secretase and Notch-100.
a, Overall structure of human γ-secretase bound to Notch-100. The structure is shown either in cartoon (cylindrical helices, top) or in surface (bottom). Glycans and lipids are displayed as sticks. Four perpendicular views are shown. b, The human γ-secretase–Notch-100 complex is represented by electrostatic surface potential. The four views are identical to those in a.
Extended Data Fig. 6 Local cryo-EM density maps for key structural elements.
a, The local cryo-EM density map for TM6 of PS1. b, The local cryo-EM density map for the anti-parallel β-sheet. This β-sheet is formed by two β-strands from loop 2 of PS1 and one β-strand from Notch-100. c, The local cryo-EM density map for the amino terminus of Notch-100. At this interface, Gln1722 of Notch-100 may directly interact with Trp653 of NCT. The contour level of the cryo-EM density is 5σ for a and b and 6σ for c.
Extended Data Fig. 7 Phospholipids in the γ-secretase–Notch-100 complex.
a, The transmembrane region of the γ-secretase–Notch-100 complex is surrounded by co-purified lipid molecules (in gold). Shown here is the cryo-EM density map. b, The local cryo-EM density maps for two phosphatidylcholines (PC, top) and two cholesterols (bottom). The contour level of the cryo-EM density for the lipids is 4σ. c, A phosphatidylcholine molecule appears to stabilize the interface between PS1 and PEN-2. d, A phosphatidylcholine molecule is bound adjacent to TM2, TM6 and TM9 of PS1. e, Three cholesterol molecules are bound on the transmembrane region of APH-1 and appear to stabilize its structure. f, A close-up view of a cholesterol molecule bound between TM3 and TM6 of APH-1. g, A close-up view of a cholesterol molecule bound between TM6 and TM7 of APH-1. h, A close-up view of a cholesterol molecule at the interface between APH-1 and the transmembrane helix of NCT. Notably, all three cholesterols shown in f, g and h are in close contact with aromatic residues.
Extended Data Fig. 8 Cryo-EM analysis of the extracellular loop of Notch-100.
a, A schematic diagram of the procedure designed to improve the local cryo-EM density map of Notch-100. b, The cryo-EM density of the Notch-100 extracellular loop in the low-pass filtered map (low-pass filtered to 4 Å, coloured orange, left). In the cut-through section of the map (right), a continuous density can be clearly seen to connect the transmembrane helix and the density lobe near NCT. c, The local cryo-EM density map of the extracellular loop of Notch-100 after local mask application and refinement. d, Colour-coded resolution distribution of the local cryo-EM density map as estimated by RELION-2.0. The local resolution for the extracellular loop is around 3.5 Å. e, The FSC curve for the 3D reconstruction of the cryo-EM map with improved local resolution for the extracellular loop of Notch-100. The average resolution is estimated to be 3.3 Å on the basis of the FSC value of 0.143. f, The local cryo-EM density map for the extracellular loop of Notch-100 (the contour level is 8σ). g, The local cryo-EM density map for the crosslinking site between PS1 loop 1 and the extracellular loop of Notch-100 (the contour level is 6σ).
Extended Data Fig. 9 Substrate entry by lateral diffusion is most likely between TM2 and TM6 of PS1.
Shown here are three views of PS1 bound to Notch-100. The red arrows indicate potential routes for substrate entry; but these routes suffer from insurmountable physical restraints that do not allow substrate passage. The orange arrow indicates the only viable route of substrate entry by lateral diffusion.
Supplementary information
Supplementary Figure 1
The uncropped Western blot and gel scans with size marker indications. The cropped areas in each figure are indicated by rectangles.
Rights and permissions
About this article
Cite this article
Yang, G., Zhou, R., Zhou, Q. et al. Structural basis of Notch recognition by human γ-secretase. Nature 565, 192–197 (2019). https://doi.org/10.1038/s41586-018-0813-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0813-8
This article is cited by
-
Chromosome evolution screens recapitulate tissue-specific tumor aneuploidy patterns
Nature Genetics (2024)
-
The aldehyde dehydrogenase 2 rs671 variant enhances amyloid β pathology
Nature Communications (2024)
-
Microbial infection promotes amyloid pathology in a mouse model of Alzheimer’s disease via modulating γ-secretase
Molecular Psychiatry (2024)
-
New precision medicine avenues to the prevention of Alzheimer’s disease from insights into the structure and function of γ-secretases
The EMBO Journal (2024)
-
Applications and prospects of cryo-EM in drug discovery
Military Medical Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.