Abstract
HIV-1 envelope glycoprotein (Env), which consists of trimeric (gp160)3 cleaved to (gp120 and gp41)3, interacts with the primary receptor CD4 and a coreceptor (such as chemokine receptor CCR5) to fuse viral and target-cell membranes. The gp120–coreceptor interaction has previously been proposed as the most crucial trigger for unleashing the fusogenic potential of gp41. Here we report a cryo-electron microscopy structure of a full-length gp120 in complex with soluble CD4 and unmodified human CCR5, at 3.9 Å resolution. The V3 loop of gp120 inserts into the chemokine-binding pocket formed by seven transmembrane helices of CCR5, and the N terminus of CCR5 contacts the CD4-induced bridging sheet of gp120. CCR5 induces no obvious allosteric changes in gp120 that can propagate to gp41; it does bring the Env trimer close to the target membrane. The N terminus of gp120, which is gripped by gp41 in the pre-fusion or CD4-bound Env, flips back in the CCR5-bound conformation and may irreversibly destabilize gp41 to initiate fusion. The coreceptor probably functions by stabilizing and anchoring the CD4-induced conformation of Env near the cell membrane. These results advance our understanding of HIV-1 entry into host cells and may guide the development of vaccines and therapeutic agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic structure coordinates are deposited in the RCSB Protein Data Bank (PDB) under the accession numbers 6MEO and 6MET; and the electron microscopy maps have been deposited in the Electron Microscopy Data Bank (EMDB) under the accession numbers EMD-9108 and EMD-9109. All other related data generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Harrison, S. C. Viral membrane fusion. Nat. Struct. Mol. Biol. 15, 690–698 (2008).
Kwong, P. D. et al. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393, 648–659 (1998).
Julien, J. P. et al. Crystal structure of a soluble cleaved HIV-1 envelope trimer. Science 342, 1477–1483 (2013).
Lee, J. H., Ozorowski, G. & Ward, A. B. Cryo-EM structure of a native, fully glycosylated, cleaved HIV-1 envelope trimer. Science 351, 1043–1048 (2016).
Pancera, M. et al. Structure and immune recognition of trimeric pre-fusion HIV-1 Env. Nature 514, 455–461 (2014).
Wang, H. et al. Cryo-EM structure of a CD4-bound open HIV-1 envelope trimer reveals structural rearrangements of the gp120 V1V2 loop. Proc. Natl Acad. Sci. USA 113, E7151–E7158 (2016).
Tan, Q. et al. Structure of the CCR5 chemokine receptor-HIV entry inhibitor maraviroc complex. Science 341, 1387–1390 (2013
Tamamis, P. & Floudas, C. A. Molecular recognition of CXCR4 by a dual tropic HIV-1 gp120 V3 loop. Biophys. J. 105, 1502–1514 (2013).
Tamamis, P. & Floudas, C. A. Molecular recognition of CCR5 by an HIV-1 gp120 V3 loop. PLoS ONE 9, e95767 (2014).
Zheng, Y. et al. Structure of CC chemokine receptor 5 with a potent chemokine antagonist reveals mechanisms of chemokine recognition and molecular mimicry by HIV. Immunity 46, 1005–1017.e1005 (2017).
Berger, E. A., Murphy, P. M. & Farber, J. M. Chemokine receptors as HIV-1 coreceptors: roles in viral entry, tropism, and disease. Annu. Rev. Immunol. 17, 657–700 (1999).
Connor, R. I., Sheridan, K. E., Ceradini, D., Choe, S. & Landau, N. R. Change in coreceptor use correlates with disease progression in HIV-1–infected individuals. J. Exp. Med. 185, 621–628 (1997).
Verhofstede, C., Nijhuis, M. & Vandekerckhove, L. Correlation of coreceptor usage and disease progression. Curr. Opin. HIV AIDS 7, 432–439 (2012).
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010).
Qin, L. et al. Crystal structure of the chemokine receptor CXCR4 in complex with a viral chemokine. Science 347, 1117–1122 (2015).
Scholten, D. J. et al. Pharmacological modulation of chemokine receptor function. Br. J. Pharmacol. 165, 1617–1643 (2012).
Lin, G., Baribaud, F., Romano, J., Doms, R. W. & Hoxie, J. A. Identification of gp120 binding sites on CXCR4 by using CD4-independent human immunodeficiency virus type 2 Env proteins. J. Virol. 77, 931–942 (2003).
Doranz, B. J. et al. Two distinct CCR5 domains can mediate coreceptor usage by human immunodeficiency virus type 1. J. Virol. 71, 6305–6314 (1997).
Rizzuto, C. D. et al. A conserved HIV gp120 glycoprotein structure involved in chemokine receptor binding. Science 280, 1949–1953 (1998).
Huang, C. C. et al. Structures of the CCR5 N terminus and of a tyrosine-sulfated antibody with HIV-1 gp120 and CD4. Science 317, 1930–1934 (2007).
Farzan, M. et al. Tyrosine sulfation of the amino terminus of CCR5 facilitates HIV-1 entry. Cell 96, 667–676 (1999).
Farzan, M. et al. The role of post-translational modifications of the CXCR4 amino terminus in stromal-derived factor 1α association and HIV-1 entry. J. Biol. Chem. 277, 29484–29489 (2002).
Oppermann, M. Chemokine receptor CCR5: insights into structure, function, and regulation. Cell. Signal. 16, 1201–1210 (2004).
Chen, J. et al. Effect of the cytoplasmic domain on antigenic characteristics of HIV-1 envelope glycoprotein. Science 349, 191–195 (2015).
Colin, P. et al. HIV-1 exploits CCR5 conformational heterogeneity to escape inhibition by chemokines. Proc. Natl Acad. Sci. USA 110, 9475–9480 (2013).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Wu, H., Kwong, P. D. & Hendrickson, W. A. Dimeric association and segmental variability in the structure of human CD4. Nature 387, 527–530 (1997).
Ozorowski, G. et al. Open and closed structures reveal allostery and pliability in the HIV-1 envelope spike. Nature 547, 360–363 (2017).
Huang, C. C. et al. Structure of a V3-containing HIV-1 gp120 core. Science 310, 1025–1028 (2005).
Gallivan, J. P. & Dougherty, D. A. Cation-π interactions in structural biology. Proc. Natl Acad. Sci. USA 96, 9459–9464 (1999).
Bannert, N. et al. Sialylated O-glycans and sulfated tyrosines in the NH2-terminal domain of CC chemokine receptor 5 contribute to high affinity binding of chemokines. J. Exp. Med. 194, 1661–1673 (2001).
Berro, R. et al. Multiple CCR5 conformations on the cell surface are used differentially by human immunodeficiency viruses resistant or sensitive to CCR5 inhibitors. J. Virol. 85, 8227–8240 (2011).
Moore, J. P. & Kuritzkes, D. R. A pièce de resistance: how HIV-1 escapes small molecule CCR5 inhibitors. Curr. Opin. HIV AIDS 4, 118–124 (2009).
Manglik, A. & Kruse, A. C. Structural basis for G protein-coupled receptor activation. Biochemistry 56, 5628–5634 (2017).
Kwon, Y. D. et al. Crystal structure, conformational fixation and entry-related interactions of mature ligand-free HIV-1 Env. Nat. Struct. Mol. Biol. 22, 522–531 (2015).
Burke, V. et al. Structural basis of the cross-reactivity of genetically related human anti-HIV-1 mAbs: implications for design of V3-based immunogens. Structure 17, 1538–1546 (2009).
Jiang, X. et al. Conserved structural elements in the V3 crown of HIV-1 gp120. Nat. Struct. Mol. Biol. 17, 955–961 (2010).
Pan, R. et al. Increased epitope complexity correlated with antibody affinity maturation and a novel binding mode revealed by structures of rabbit antibodies against the third variable loop (V3) of HIV-1 gp120. J. Virol. e01894-17 (2018).
Lyumkis, D. et al. Cryo-EM structure of a fully glycosylated soluble cleaved HIV-1 envelope trimer. Science 342, 1484–1490 (2013).
Moore, J. P., McKeating, J. A., Weiss, R. A. & Sattentau, Q. J. Dissociation of gp120 from HIV-1 virions induced by soluble CD4. Science 250, 1139–1142 (1990).
Thali, M., Furman, C., Helseth, E., Repke, H. & Sodroski, J. Lack of correlation between soluble CD4-induced shedding of the human immunodeficiency virus type 1 exterior envelope glycoprotein and subsequent membrane fusion events. J. Virol. 66, 5516–5524 (1992).
Chang, M. I., Panorchan, P., Dobrowsky, T. M., Tseng, Y. & Wirtz, D. Single-molecule analysis of human immunodeficiency virus type 1 gp120-receptor interactions in living cells. J. Virol. 79, 14748–14755 (2005).
Dobrowsky, T. M., Zhou, Y., Sun, S. X., Siliciano, R. F. & Wirtz, D. Monitoring early fusion dynamics of human immunodeficiency virus type 1 at single-molecule resolution. J. Virol. 82, 7022–7033 (2008).
Brandenberg, O. F., Magnus, C., Regoes, R. R. & Trkola, A. The HIV-1 entry process: a stoichiometric view. Trends Microbiol. 23, 763–774 (2015).
Floyd, D. L., Ragains, J. R., Skehel, J. J., Harrison, S. C. & van Oijen, A. M. Single-particle kinetics of influenza virus membrane fusion. Proc. Natl Acad. Sci. USA 105, 15382–15387 (2008).
Zhu, P. et al. Distribution and three-dimensional structure of AIDS virus envelope spikes. Nature 441, 847–852 (2006).
Rosen, O., Sharon, M., Quadt-Akabayov, S. R. & Anglister, J. Molecular switch for alternative conformations of the HIV-1 V3 region: implications for phenotype conversion. Proc. Natl Acad. Sci. USA 103, 13950–13955 (2006).
Ho, S. H., Trunova, N., Gettie, A., Blanchard, J. & Cheng-Mayer, C. Different mutational pathways to CXCR4 coreceptor switch of CCR5-using simian-human immunodeficiency virus. J. Virol. 82, 5653–5656 (2008).
Edo-Matas, D., van Dort, K. A., Setiawan, L. C., Schuitemaker, H. & Kootstra, N. A. Comparison of in vivo and in vitro evolution of CCR5 to CXCR4 coreceptor use of primary human immunodeficiency virus type 1 variants. Virology 412, 269–277 (2011).
Westby, M. et al. Reduced maximal inhibition in phenotypic susceptibility assays indicates that viral strains resistant to the CCR5 antagonist maraviroc utilize inhibitor-bound receptor for entry. J. Virol. 81, 2359–2371 (2007).
Seclén, E. et al. Primary resistance to maraviroc in a large set of R5-V3 viral sequences from HIV-1-infected patients. J. Antimicrob. Chemother. 65, 2502–2504 (2010).
Jiang, X. et al. Characterizing the diverse mutational pathways associated with R5-tropic maraviroc resistance: HIV-1 that uses the drug-bound CCR5 coreceptor. J. Virol. 89, 11457–11472 (2015).
Pugach, P. et al. HIV-1 clones resistant to a small molecule CCR5 inhibitor use the inhibitor-bound form of CCR5 for entry. Virology 361, 212–228 (2007).
Lin, G. et al. Replication-competent variants of human immunodeficiency virus type 2 lacking the V3 loop exhibit resistance to chemokine receptor antagonists. J. Virol. 81, 9956–9966 (2007).
Zolla-Pazner, S. et al. The cross-clade neutralizing activity of a human monoclonal antibody is determined by the GPGR V3 motif of HIV type 1. AIDS Res. Hum. Retroviruses 20, 1254–1258 (2004).
Frey, G. et al. A fusion-intermediate state of HIV-1 gp41 targeted by broadly neutralizing antibodies. Proc. Natl Acad. Sci. USA 105, 3739–3744 (2008).
Kovacs, J. M. et al. HIV-1 envelope trimer elicits more potent neutralizing antibody responses than monomeric gp120. Proc. Natl Acad. Sci. USA 109, 12111–12116 (2012).
Freeman, M. M. et al. Crystal structure of HIV-1 primary receptor CD4 in complex with a potent antiviral antibody. Structure 18, 1632–1641 (2010).
Cai, Y. et al. Antigenicity-defined conformations of an extremely neutralization-resistant HIV-1 envelope spike. Proc. Natl Acad. Sci. USA 114, 4477–4482 (2017).
Brady, A. E. & Limbird, L. E. G protein-coupled receptor interacting proteins: emerging roles in localization and signal transduction. Cell. Signal. 14, 297–309 (2002).
Visegrády, A., Boros, A., Némethy, Z., Kiss, B. & Keseru, G. M. Application of the BD ACTOne TM technology for the high-throughput screening of Gs-coupled receptor antagonists. J. Biomol. Screen. 12, 1068–1073 (2007).
Melar, M., Ott, D. E. & Hope, T. J. Physiological levels of virion-associated human immunodeficiency virus type 1 envelope induce coreceptor-dependent calcium flux. J. Virol. 81, 1773–1785 (2007).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Ru, H. et al. Molecular mechanism of V(D)J recombination from synaptic RAG1–RAG2 complex structures. Cell 163, 1138–1152 (2015).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protocols 5, 725–738 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Feng, Y., Broder, C. C., Kennedy, P. E. & Berger, E. A. HIV-1 entry cofactor: functional cDNA cloning of a seven-transmembrane, G protein-coupled receptor. Science 272, 872–877 (1996).
Wu, L. et al. CD4-induced interaction of primary HIV-1 gp120 glycoproteins with the chemokine receptor CCR-5. Nature 384, 179–183 (1996).
Alkhatib, G. et al. CC CKR5: a RANTES, MIP-1α, MIP-1β receptor as a fusion cofactor for macrophage-tropic HIV-1. Science 272, 1955–1958 (1996).
Choe, H. et al. The β-chemokine receptors CCR3 and CCR5 facilitate infection by primary HIV-1 isolates. Cell 85, 1135–1148 (1996).
Deng, H. et al. Identification of a major co-receptor for primary isolates of HIV-1. Nature 381, 661–666 (1996).
Doranz, B. J. et al. A dual-tropic primary HIV-1 isolate that uses fusin and the β-chemokine receptors CKR-5, CKR-3, and CKR-2b as fusion cofactors. Cell 85, 1149–1158 (1996).
Dragic, T. et al. HIV-1 entry into CD4+ cells is mediated by the chemokine receptor CC-CKR-5. Nature 381, 667–673 (1996).
Jin, J. et al. CCR5 adopts three homodimeric conformations that control cell surface delivery. Sci. Signal. 11, eaal2869 (2018).
Acknowledgements
We thank S. Harrison and A. Kruse for advice, K. Song, J. Chen, R. Martin and W. Chang for technical assistance, N. Grigorieff and T. Grant for discussion at the early stage of the project, and S. Harrison and A. Kruse for critical reading of the manuscript. This work was supported by NIH grants AI141002 (to B.C.), AI106488 (to B.C.), AI129721 (to B.C.), AI127193 (to B.C. and J.J.C.), the Center for HIV/AIDS Vaccine Immunology - Immunogen Design AI-100645 (to B. F. Haynes), and Collaboration for AIDS Vaccine Discovery (CAVD) grant OPP1169339 (to D. H. Barouch from the Bill and Melinda Gates Foundation).
Reviewer information
Nature thanks G. Melikyan, S. Subramaniam and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
B.C. and M.M.S. designed the experiments. H.P. and M.M.S. purified the CD4–gp120–CCR5 complex. J.L. performed CCR5 chemokine receptor assays. M.M.S. and S.R.-V. carried out CCR5 coreceptor functional assays. M.M.S. performed electron microscopy data collection with contributions from C.X. M.L. processed the initial negative-stain data. M.M.S. processed the cryo-EM data with contributions from M.L., and built the atomic model with help from B.C. All authors analysed the data. B.C. and M.M.S. wrote the manuscript with input from M.L. and J.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Previously known structures of CCR5 and CXCR4.
CCR5 and CXCR4 were identified as the coreceptors for HIV-1 entry in 199671,72,73,74,75,76,77. a, b, Crystal structures of a modified CCR5 (C224–N226 deleted and replaced with rubredoxin; ΔF320–L352; and the point mutations C58Y, G163N, A233D and K303E) in complex with the HIV entry-inhibitor maraviroc (PDB ID: 4MBS7) (a) and a modified chemokine [5P7]CCL5 (an antagonist; PDB ID: 5UIW10) (b). CCR5 is shown in ribbon diagram in blue, with the internally fused rubredoxin in magenta and the ligands in yellow. c–e, Crystal structures of an engineered CXCR4 in complex with a viral chemokine antagonist vMIP-II (PDB ID: 4RWS15) (c), a small molecule antagonist IT1t (PDB ID: 3ODU14) (d) and a cyclic peptide antagonist CVX15 (PDB ID: 3OE014) (e). CXCR4 is shown in green, the fused T4 lysozyme in magenta and the ligands in yellow.
Extended Data Fig. 2 Characterization of stable cell lines (HEK293T and Expi293F) expressing wild-type human CCR5.
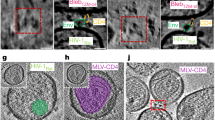
a, Chemokine receptor assay. HEK293T and HEK293T-CCR5 (stable) cells were treated with different concentrations of CCL5. Ft/F0 is a fluorescence-signal ratio proportional to that of intracellular cAMP concentration at 40 min after CCL5 activation and at time 0. The dose–response curves were plotted for both HEK293T (black) and HEK293T-CCR5 (red) cells. The experiment was carried out in quadruplicate, and repeated at least three times with similar results. Error bars indicate the standard deviation calculated by the STDEV function in Excel. b, Flow cytometry histograms of HIV-1 gp120 binding to CCR5 expressed on the cell surfaces in the absence (orange) or presence (red) of soluble CD4. HEK293T cells (black), CCR5-expressing cells only (grey) and CCR5-expressing cells with soluble CD4 only (blue) were negative controls. The experiment was repeated independently at least twice with similar results. c, HIV-1 Env-mediated cell–cell fusion. HEK293T cells stably transfected with CCR5 were mixed with HIV-1 Env (gp160)-expressing cells in the absence or presence of soluble CD4. The CCR5 cells fuse with CD4-triggered Env cells very efficiently, and form large syncytia that cover almost the entire well. The experiment was repeated independently twice with similar results. d, Chemokine receptor assay by various ligands. As in a, Expi293F and Expi293F-CCR5 (stable) cells were treated with CCL5, gp120, CD4 or the complex of gp120 and CD4. The dose–response curves were plotted for both Expi293F as a control (left) and Expi293F-CCR5 (right) cells, with different ligands as indicated. The experiment was carried out in quadruplicate and repeated at least three times with similar results. Error bars indicate the standard deviation calculated by the STDEV function in Excel. e, Left, kinetic curves of 5 representative wells of HEK293T-CCR5 cells treated with 5 different ligands as indicated. ATP activates the endogenous Gq-coupled G-protein-coupled receptor (P2Y receptor), as a positive control. The ratio represents fluorescence intensity divided by baseline intensity. Right, dose–response curve of each ligand. The y axis is a background-subtracted ratio (peak fluorescent intensity ratio − 1). We conclude that our gp120 and gp120–CD4 do not activate G-protein-mediated calcium flux at the concentrations tested here. The experiment was carried out in quadruplicate and repeated twice with similar results. Error bars indicate the standard deviation calculated by the STDEV function in Excel.
Extended Data Fig. 3 Purification of the CD4-gp120-CCR5 complex.
a, Schematic of expression constructs for HIV-1 gp120, human CCR5 and CD4. Segments of gp120 are designated as follows: C1–C5, conserved regions 1–5; V1–V5, variable regions 1–5; and His-tag, a six-histidine tag. Tree-like symbols represent glycans. Abbreviations used for segments of CCR5 are: N, N terminus; TM1–TM7, transmembrane helices 1–7; ECL1–ECL3, extracellular loops 1–3; ICL1–ICL3, intracellular loops 1–3; and CT, cytoplasmic tail. For CD4, the following abbreviations are used: D1–D4, immunoglobulin (Ig) domains 1–4; and strep tag, a purification tag. The transmembrane segment (TM) and cytoplasmic tail (CT) in grey are truncated in the expression construct. b, Unmodified human CCR5 in complex with HIV-1 gp120 and four-domain CD4 was purified by the following steps. (1) Complex formation: HIV-1 gp120 (light blue) and strep-tagged, four-domain CD4 (green) were incubated with CCR5 (magenta)-expressing cells to allow formation of the CD4–gp120–CCR5 complex on cell surfaces. (2) Strep-tag purification: the CCR5 complex and some of the CD4–gp120 complex were captured to strep-tactin resin via the strep-tagged CD4 (strep tag in purple). They were eluted by d-desthiobiotin under mild conditions. (3) Negative selection by an anti-V3 antibody to remove the CD4–gp120 complex. The CCR5 complex was further purified by size-exclusion chromatography. c, The purified CD4–gp120–CCR5 complex was resolved by gel-filtration chromatography on a Superose 6 column in the presence of the detergent LMNG. The molecular-mass standards include thyoglobulin (670 kDa), ferritin (440 kDa), γ-globulin (158 kDa) and ovalbumin (44 kDa). The expected size of the CCR5 complex is ~310 kDa (120 kDa for gp120, 50 kDa for four-domain CD4, 40 kDa for CCR5 and ~100 kDa for LMNG micelle). Peak fractions were analysed by Coomassie-stained SDS–PAGE (lanes 1–3). Labelled bands were confirmed by western blot and protein sequencing. The experiment was repeated independently at least 15 times with similar results.
Extended Data Fig. 4 Characterization of the CD4–gp120–CCR5 complex by electron microscopy.
a, Representative image of the CD4–gp120–CCR5 complex in negative stain. The experiment was repeated independently at least 4 times with similar results. b, 2D averages of the negatively stained CD4–gp120–CCR5 complex. The box size of 2D averages is ~330 Å. c, 3D reconstruction of the negatively stained CD4–gp120–CCR5 complex, fitted with a gp120 structure containing an extended V3 loop (PDB ID: 2QAD20), four-domain CD4 (PDB ID: 1WIO) and CCR5 (PDB ID: 4MBS). d, A representative cryo-EM image of the four-domain-CD4–gp120–CCR5 complex. Scale bar, 25 nm. Five independent large datasets were collected with similar results. e, 2D averages of the cryo-EM particle images show secondary structural features for both gp120 and CCR5.
Extended Data Fig. 5 Single-particle cryo-EM analysis of the CD4–gp120–CCR5 complex.
a, Data-processing workflow for the CD4–gp120–CCR5 complex. b, 3D reconstructions of the CD4–gp120–CCR5 complex refined with no mask at an overall resolution of 4.5 Å (left), and with a mask to exclude the last two domains of CD4 at a resolution of 3.9 Å (right), are coloured according to local resolution estimated by RELION. c, The angular distribution of the cryo-EM particles used in the reconstruction is also shown in respect to both the side and top views of the electron microscopy map. d, Gold standard Fourier shell correlation curves of the unmasked and masked electron microscopy reconstructions shown in b.
Extended Data Fig. 6 Gallery of representative density for the CD4–gp120–CCR5 complex.
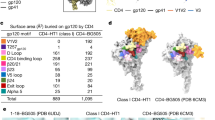
Representative density in grey mesh from the 3.9 Å resolution electron microscopy map is shown for transmembrane helices TM1–TM7, the N terminus of CCR5, extracellular loop 3 (ELC3) near TM6, Tys10, Tys14 and Tyr15 (red model); two V3 regions; and for helix α1, N terminus, V3 loop, the bridging sheet and N-linked glycan at N262 of gp120 (cyan model).
Extended Data Fig. 7 Comparison of the conformations of the V3 loop and [5P7]CCL5 in complex with CCR5, as well as of gp120-bound CCR5 and G-protein-bound β2 adrenergic receptor.
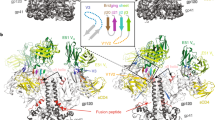
a, The structures of the CD4–gp120–CCR5 and [5P7]CCL5–CCR5 complexes are superposed on CCR5 (red). The V3 loop of gp120 with its Pro311 in stick model is in cyan and [5P7]CCL5 with its Pro3 in stick model in yellow. Residues 309–316 of the V3 loop and residues 1–8 of [5P7]CCL5 adopt a very similar structure, and are highlighted in a rectangular box. b, Superposition of the structures of the N terminus of the gp120-bound CCR5 (red) and the complementarity-determining region H3 loop of antibody 412d in complex with gp120 core (green). The electron microscopy density of the CD4–gp120–CCR5 complex is shown in grey. The positions of the sulfated tyrosine (‘Tys’) residues, including Tys10 and Tys14 (from CCR5) and Tys100 and Tys100c (from 412d), are indicated. c, A model for interactions of three CD4 receptors and three CCR5 coreceptors with the SOSIP Env trimer. The side and bottom views of a composite structure of the CD4–CCR5–SOSIP Env trimer complex are shown. The model was generated using the CD4-bound SOSIP trimer (PDB ID: 5VN3) and the structure of the CD4–gp120–CCR5 complex from this study. All the structures were aligned on the basis of the core region of gp120. CCR5 is shown in red, CD4 in green, gp120 in blue, the gp120 of SOSIP in dark blue and the gp41 of SOSIP in grey. The crystallographic dimer of CCR5 (PDB ID: 4MBS) is also shown, on the left only, in a rectangular box. The observed crystallographic dimer of CCR5 or the transmembrane helix 5-mediated dimer by modelling does not seem to be relevant to binding to either monomeric or trimeric gp1207,78. d, Superposition of the structures of the gp120-bound CCR5 (red) and the Gs-protein-bound β2 adrenergic receptor (blue). The position of TM6, which is critical for the activation of G-protein-coupled receptors, is indicated.
Extended Data Fig. 8 Comparison of conformations of different structures of monomeric gp120 and various V3 loops.
a, Comparison of structures of an unliganded gp120 core (PDB ID:4OLV; purple), a CD4-bound monomeric gp120 core with the V3 loop (PDB ID: 2QAD; blue) and gp120 in complex with CD4 and CCR5 from this study (cyan). The gp120 core region is marked by a circle with a diameter of 50 Å. The N and C termini, V1V2 stem, V3 stem or loop and bridging sheet are indicated. b, Representative conformations that an HIV-1 V3 loop can adopt. From left to right, V3 loop in the unliganded SOSIP BG505 Env trimer (PDB ID: 4ZMJ); the first-V3-containing gp120 core in complex with CD4 and antibody X5 (PDB ID: 2B4C29); CD4- and 412d-bound monomeric gp120 core with V3 (PDB ID: 2QAD); CCR5-bound intact gp120 (this study); and V3 peptide in complex with antibody 447-52D (PDB ID: 3GHB36); antibody 268-D (PDB ID: 3GO137); antibody 2557 (PDB ID: 3MLV37); and antibody 10A37 (PDB ID: 5V6L38). The root-mean-square deviation of each structure (except for 5V6L), relative to the CCR5-bound gp120 monomer, is shown at the bottom in parentheses.
Extended Data Fig. 9 Model of HIV-1 Env activation to induce membrane fusion.
A hypothesis of how the cellular receptors CD4 and CCR5 trigger the HIV-1 Env trimer to induce membrane fusion and viral entry. Left, virus attaches to the target cell by gp120 (cyan) binding to CD4 (green). Helix collar (gp41), the four-helix collar gripping the N- and C termini of gp120. Right, immediate binding by CCR5 (red) prevents rapid dissociation between gp120 and CD4, stabilizes the CD4-induced conformational changes within the Env trimer and brings the trimer close to the cell membrane. Simultaneous binding of gp120 to both CD4 and CCR5 may require bending in the cell membrane. The fusion peptide (magenta) of gp41 (grey) flips out owing to intrinsic conformational dynamics, which enables the bending back of the N and C termini of gp120. This bending blocks the fusion peptide from resuming its original position in the trimer. The movements of the fusion peptide and gp120 termini effectively weaken the non-covalent association between the two subunits and may lead to partial or complete dissociation of gp120, as well as a series of refolding events in gp41 to adopt the pre-hairpin intermediate conformation (with the fusion peptides inserting into the target-cell membrane). Extended helix (gp41), three helices in the fusion-intermediate conformation of gp41.
Supplementary information
Rights and permissions
About this article
Cite this article
Shaik, M.M., Peng, H., Lu, J. et al. Structural basis of coreceptor recognition by HIV-1 envelope spike. Nature 565, 318–323 (2019). https://doi.org/10.1038/s41586-018-0804-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0804-9
This article is cited by
-
Structural basis of dimerization of chemokine receptors CCR5 and CXCR4
Nature Communications (2023)
-
Conserved HIV-1 V3 loop helix offers potential for broad neutralization
Nature Structural & Molecular Biology (2023)
-
HIV-1 Env trimers asymmetrically engage CD4 receptors in membranes
Nature (2023)
-
Trapping the HIV-1 V3 loop in a helical conformation enables broad neutralization
Nature Structural & Molecular Biology (2023)
-
Autoantibodies against chemokines post-SARS-CoV-2 infection correlate with disease course
Nature Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.