Abstract
Haematopoiesis is an essential process that evolved in multicellular animals. At the heart of this process are haematopoietic stem cells (HSCs), which are multipotent and self-renewing, and generate the entire repertoire of blood and immune cells throughout an animal’s life1. Although there have been comprehensive studies on self-renewal, differentiation, physiological regulation and niche occupation in vertebrate HSCs, relatively little is known about the evolutionary origin and niches of these cells. Here we describe the haematopoietic system of Botryllus schlosseri, a colonial tunicate that has a vasculature and circulating blood cells, and interesting stem-cell biology and immunity characteristics2,3,4,5,6,7,8. Self-recognition between genetically compatible B. schlosseri colonies leads to the formation of natural parabionts with shared circulation, whereas incompatible colonies reject each other3,4,7. Using flow cytometry, whole-transcriptome sequencing of defined cell populations and diverse functional assays, we identify HSCs, progenitors, immune effector cells and an HSC niche, and demonstrate that self-recognition inhibits allospecific cytotoxic reactions. Our results show that HSC and myeloid lineage immune cells emerged in a common ancestor of tunicates and vertebrates, and also suggest that haematopoietic bone marrow and the B. schlosseri endostyle niche evolved from a common origin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing data can be found on the NCBI Sequence Read Archive under accession PRJNA414486. RPKM values of gene expression and differential expression analysis results are shown in Supplementary Table 1. All other relevant data are available in the manuscript.
References
Weissman, I. L. Stem cells: units of development, units of regeneration, and units in evolution. Cell 100, 157–168 (2000).
Sabbadin, A. Le basi genetiche della capacita di fusione fra colonies in Botryllus schlosseri (Ascidiacea). Rend. Accad. Naz. Lincei. Ser. 8, 1031–1035 (1962).
Scofield, V. L., Schlumpberger, J. M., West, L. A. & Weissman, I. L. Protochordate allorecognition is controlled by a MHC-like gene system. Nature 295, 499–502 (1982).
Stoner, D. S., Rinkevich, B. & Weissman, I. L. Heritable germ and somatic cell lineage competitions in chimeric colonial protochordates. Proc. Natl Acad. Sci. USA 96, 9148–9153 (1999).
Laird, D. J., De Tomaso, A. W. & Weissman, I. L. Stem cells are units of natural selection in a colonial ascidian. Cell 123, 1351–1360 (2005).
Voskoboynik, A. et al. Identification of the endostyle as a stem cell niche in a colonial chordate. Cell Stem Cell 3, 456–464 (2008).
Voskoboynik, A. et al. Identification of a colonial chordate histocompatibility gene. Science 341, 384–387 (2013).
Corey, D. M. et al. Developmental cell death programs license cytotoxic cells to eliminate histocompatible partners. Proc. Natl Acad. Sci. USA 113, 6520–6525 (2016).
Darwin, C. On the Origin of the Species by Natural Selection (John Murray, London, 1859).
Delsuc, F., Brinkmann, H., Chourrout, D. & Philippe, H. Tunicates and not cephalochordates are the closest living relatives of vertebrates. Nature 439, 965–968 (2006).
Voskoboynik, A. et al. The genome sequence of the colonial chordate, Botryllus schlosseri. eLife 2, e00569 (2013).
Lauzon, R. J., Patton, C. W. & Weissman, I. L. A morphological and immunohistochemical study of programmed cell death in Botryllus schlosseri (Tunicata, Ascidiacea). Cell Tissue Res. 272, 115–127 (1993).
Hulett, H. R., Bonner, W. A., Barrett, J. & Herzenberg, L. A. Cell sorting: automated separation of mammalian cells as a function of intracellular fluorescence. Science 166, 747–749 (1969).
Ben-Ari Fuchs, S. et al. Geneanalytics: an integrative gene set analysis tool for next generation sequencing, rnaseq and microarray data. OMICS 20, 139–151 (2016).
Seita, J. et al. Gene Expression Commons: an open platform for absolute gene expression profiling. PLoS ONE 7, e40321 (2012).
Rinkevich, Y. et al. Repeated, long-term cycling of putative stem cells between niches in a basal chordate. Dev. Cell 24, 76–88 (2013).
Ermak, T. H. in Phylogeny of Thymus and Bone Marrow-Bursa Cells 45–56 (Elsevier, 1976).
Adams, G. B. et al. Stem cell engraftment at the endosteal niche is specified by the calcium-sensing receptor. Nature 439, 599–603 (2006).
Yusuf, R. Z. & Scadden, D. T. Homing of hematopoietic cells to the bone marrow. J. Vis. Exp. 25,1104 (2009).
Wright, D. E., Wagers, A. J., Gulati, A. P., Johnson, F. L. & Weissman, I. L. Physiological migration of hematopoietic stem and progenitor cells. Science 294, 1933–1936 (2001).
Ogasawara, M. et al. Gene expression profiles in young adult Ciona intestinalis. Dev. Genes Evol. 212, 173–185 (2002).
Cañestro, C., Bassham, S. & Postlethwait, J. H. Evolution of the thyroid: anterior-posterior regionalization of the Oikopleura endostyle revealed by Otx, Pax2/5/8, and Hox1 expression. Dev. Dyn. 237, 1490–1499 (2008).
Burighel, P. & Cloney, R. A. in Microscopic Anatomy of Invertebrates (eds Harrison, F. W. & Ruppert, E. E.) 221–347 (Alan R. Liss, 1997).
Ogasawara, M., Lauro, R. D. & Satoh, N. Ascidian homologs of mammalian thyroid transcription factor-1 gene are expressed in the endostyle. Zool. Sci. 16, 559–565 (1999).
Chen, J. Y. et al. Hoxb5 marks long-term haematopoietic stem cells and reveals a homogenous perivascular niche. Nature 530, 223–227 (2016).
Ballarin, L. & Cima, F. Cytochemical properties of Botryllus schlosseri haemocytes: indications for morpho-functional characterisation. Eur. J. Histochem. 49, 255–264 (2005).
Lauzon, R. J., Brown, C., Kerr, L. & Tiozzo, S. Phagocyte dynamics in a highly regenerative urochordate: insights into development and host defense. Dev. Biol. 374, 357–373 (2013).
Franchi, N. & Ballarin, L. Immunity in protochordates: the tunicate perspective. Front. Immunol. 8, 674 (2017).
Oren, M. et al. Marine invertebrates cross phyla comparisons reveal highly conserved immune machinery. Immunobiology 218, 484–495 (2013).
Timonen, T., Ortaldo, J. R. & Herberman, R. B. Characteristics of human large granular lymphocytes and relationship to natural killer and K cells. J. Exp. Med. 153, 569–582 (1981).
Ljunggren, H. G. & Kärre, K. In search of the ‘missing self’: MHC molecules and NK cell recognition. Immunol. Today 11, 237–244 (1990).
Smith, L. C. et al. in Invertebrate Immunity (ed. Söderhäll, K.) 260–301 (Springer, 2010).
Kawabata, S. in Invertebrate Immunity (ed. Söderhäll, K.) 122–136 (Springer, 2010).
Ballarin, L. Ascidian cytotoxic cells: state of the art and research perspectives. Inv. Surv. J. 9, 1–6 (2012).
Boyd, H. C. et al. Growth and sexual maturation of laboratory-cultured Monterey Botryllus schlosseri. Biol. Bull. 170, 91–109 (1986).
Taketa, D. A. & De Tomaso, A. W. Botryllus schlosseri allorecognition: tackling the enigma. Dev. Comp. Immunol. 48, 254–265 (2015).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Koh, P. W. et al. An atlas of transcriptional, chromatin accessibility, and surface marker changes in human mesoderm development. Sci. Data 3, 160109 (2016).
Burighel, P. & Brunetti, R. The circulatory system in the blastozooid of the colonial ascidian Botryllus schlosseri (Pallas). Italian J. Zool. 38, 273–289 (1971).
Zaniolo, G., Lane, N. J., Burighel, P. & Manni, L. Development of the motor nervous system in ascidians. J. Comp. Neurol. 443, 124–135 (2002).
Edri-Brami, M. et al. Glycans in sera of amyotrophic lateral sclerosis patients and their role in killing neuronal cells. PLoS ONE 7, e35772 (2012).
Sabbadin, A. & Graziani, G. Microgeographical and ecological distribution of colour morphs of Botryllus schlosseri (Ascidiacea). Nature 213, 815–816 (1967).
Cima, F. et al. Life history and ecological genetics of the colonial ascidian Botryllus schlosseri. Zool. Anz. 257, 54–70 (2015).
Laird, D. J. & Weissman, I. L. Continuous development precludes radioprotection in a colonial ascidian. Dev. Comp. Immunol. 28, 201–209 (2004).
Braden, B. P. et al. Vascular regeneration in a basal chordate is due to the presence of immobile, bi-functional cells. PLoS ONE 9, e95460 (2014).
Köster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Wolff, L. & Humeniuk, R. Concise review: erythroid versus myeloid lineage commitment: regulating the master regulators. Stem Cells 31, 1237–1244 (2013).
Rosental, B. et al. The effect of chemotherapy/radiotherapy on cancerous pattern recognition by NK cells. Curr. Med. Chem. 19, 1780–1791 (2012).
Potts, K. S. & Bowman, T. V. Modeling myeloid malignancies using zebrafish. Front. Oncol. 7, 297 (2017).
Moss, L. D. et al. Identification of phagocytic cells, NK-like cytotoxic cell activity and the production of cellular exudates in the coelomic cavity of adult zebrafish. Dev. Comp. Immunol. 33, 1077–1087 (2009).
Wei, S. et al. The zebrafish activating immune receptor Nitr9 signals via Dap12. Immunogenetics 59, 813–821 (2007).
Tang, Q. et al. Dissecting hematopoietic and renal cell heterogeneity in adult zebrafish at single-cell resolution using RNA sequencing. J. Exp. Med. 214, 2875–2887 (2017).
Han, Q. et al. Characterization of lamprey IL-17 family members and their receptors. J. Immunol. 195, 5440–5451 (2015).
Hirano, M., Das, S., Guo, P. & Cooper, M. D. The evolution of adaptive immunity in vertebrates. Adv. Immunol. 109, 125–157 (2011).
Han, Y., Huang, G., Zhang, Q., Yuan, S. & Liu, J. The primitive immune system of amphioxus provides insights into the ancestral structure of the vertebrate immune system. Dev. Comp. Immunol. 34, 791–796 (2010).
Rhodes, C. P., Ratcliffe, N. A. & Rowley, A. F. Presence of coelomocytes in the primitive chordate amphioxus (Branchiostoma lanceolatum). Science 217, 263–265 (1982).
Muñoz-Chápuli, R., Carmona, R., Guadix, J. A., Macías, D. & Pérez-Pomares, J. M. The origin of the endothelial cells: an evo-devo approach for the invertebrate/vertebrate transition of the circulatory system. Evol. Dev. 7, 351–358 (2005).
Lanot, R., Zachary, D., Holder, F. & Meister, M. Postembryonic hematopoiesis in Drosophila. Dev. Biol. 230, 243–257 (2001).
Meister, M. & Lagueux, M. Drosophila blood cells. Cell. Microbiol. 5, 573–580 (2003).
Nakayama, K., Nomoto, A. M. & Nishijima, M. Morphological and functional characterization of hemocytes in the giant clam Tridacna crocea. J. Invertebr. Pathol. 69, 105–111 (1997).
Acknowledgements
We thank C. Lowe, C. Anselmi, I. Dimov, S. Karten, C. Patton, J. Thompson, P. Lovelace, R. Voskoboynik, N. Fernhoff, W.-J. Lu, P. Chu, K. Weiskopf, M. Oren, B. Wang, J. Lee, B. Compton, K. Uhlinger, T. Naik and T. Storm for technical advice and help. This study was supported by NIH grants R56AI089968, R01AG037968 and RO1GM100315 (to I.L.W., S.R.Q., and A.V.), the Virginia and D. K. Ludwig Fund for Cancer Research, a grant from the Siebel Stem Cell Institute and a Stinehart-Reed grant (to I.L.W.). L.M. was supported by PRIN - Prot. 2015NSFHXF. B.R. was supported by a Postdoctoral Fellowship of the Human Frontier Science Program Organization LT000591/2014-L, NIH Immunology training grant 5T32AI07290-28 and NIH Hematology training grant T32 HL120824-03.
Reviewer information
Nature thanks M. D. Cooper, W. Jeffery, G. Litman and J. Pascual-Anaya for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Conception and design: B.R., A.V., M.K., S.R.Q. and I.L.W.; mariculture: K.J.I. and K.J.P.; flow cytometry and sorting: B.R.; CyTOF screening and cluster analysis: B.R., S.-Y.C. and G.P.N.; RNA isolation and library preparation: B.R., K.J.P., R.S. and A.V.; sequencing: J.O., G.M. and N.F.N.; sequencing analysis and development of analytical tools: M.K., J.S, A.M.N. and S.R.Q.; immunological assays: B.R.; microscopy and experimental design: B.R., D.M.C., K.J.I., D.N.C. and A.V.; electron microscopy: L.M.; BHF localization to the membrane and serum validation: J.M.T., B.R. and A.V.; transplantation: B.R.; writing of manuscript: B.R., A.V., M.K., T.R., K.J.P. and I.L.W.; technical support and conceptual advice: N.F.N., D.M.C., A.M.N., T.R., G.P.N., S.R.Q., A.V. and I.L.W.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
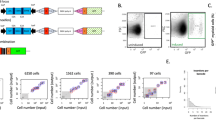
Extended Data Fig. 1 B. schlosseri cell sorting workflow.
a, Outline of cell purification process. Unsorted cells (light microscopy) are loaded into a FACS (i) and sorted, and this is followed by morphological observation (ii). Cells were labelled with diverse markers and screened using CyTOF (iii) for differential labelling. On the basis of SPADE cluster analysis (iv) markers were selected for FACS gating (v) before a final sort was performed (vi; c). b, Sorting based on FCS/SSC in the lower panel, and natural fluorescence in the upper panels. The analysis is after gating propidium iodide (PI)-negative cells (live cells). The specific excitation laser and optical filter for emission measurements were as follows (excitation laser in nm and filters stated as long pass (LP) and band pass (BP)): 488 nm for FSC and SSC, 488 nm (550LP 575/25BP)- PE, 405 nm (450/50BP)-Pacific Blue, 633 nm (660/20BP)-APC, 633 nm (690LP 710/50BP)-APC-Cy5.5. Nomenclature based on published work3,26. Experiment was performed three times. Scale bars, 20 μm. c, Sorting panels of 34 cell populations using FSC, SSC (central panel), CD49d, CD57, concanavalin-A (ConA), BHF and AP, after gating PI-negative cells (live cells). Central panel is FCS/SSC, from which additional populations were differentiated. The specific excitation laser and optical filter for emission measurements were as follows: 488 nm (505LP 530/30BP)-AP, 488 nm (755LP 780/60BP)-CD49d, 405 nm (450/50BP)-CD57, 633 nm (660/20BP)-ConA, 633 nm (755LP 780/60BP)-BHF. d, Hierarchy of sorted cell populations by main parameters of differentiation. For example, the control populations used in Fig. 2b–e (CP19, 20, 21) are all included in CP18. CP18 is derived from CP9. e, H&E staining of the end point cell populations isolated in a rectangle of original population by FSC/SSC. Live imaging was done three times; H&E was one experiment with three replicates. Colour key (c, d) for different populations identified in this study and the figures that further describe them.
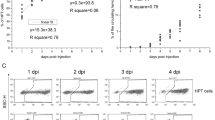
Extended Data Fig. 2 Screen for differentiation markers of cell populations for FACS-based sorting.
a, b, Examples of antibodies screened by CyTOF mass cytometry with B. schlosseri cells analysed by 2D mass spectrometry. The CyTOF screen was performed once. a, Examples of antibodies considered as nonspecific binders because different antibodies showed the same binding patterns. b, Examples of antibodies considered as specific binders because a cell population was bound by one antibody but not the other (red rectangle). c, d, Examples of validation screen by flow cytometry of antibodies with specific binding by CyTOF, done in 2D fluorescence excited by the same laser owing to autofluorescence. Negative FACS validation was done twice and positive three times. c, Negative validation of CCR2 labelled with PE. d, Positive validation of CD57 labelled with Pacific Blue. e, Flow cytometry of live B. schlosseri cells labelled with anti-BHF mouse serum. Anti-mouse Alexa Fluor-647 was used as a secondary antibody. f, Example of positive differential labelling by the lectin concanavalin-A in PE-Cy7 by FACS. The experiment was performed three times. g, Confocal imagery of membrane concanavalin-A labelling with Alexa Fluor-633 in red and Hoechst DNA labelling in blue. The experiment was performed once. Scale bar, 20 µm.
Extended Data Fig. 3 B. schlosseri cell population clustering based on transcriptome analysis.
a, We used 250 genes with the largest weights in the first 11 principal components (explaining 90% of the variance in mean-adjusted log-transformed gene counts) to cluster the different cell populations in a heat map. b, Transcriptome sequencing of B. schlosseri cell populations compared to FACS analysis. 2D projection of the distances between transcriptomes of cell populations based on all differential genes. Lines are drawn between the nearest two neighbours. Blue, FACS adjacency of the populations in the differentiation panel; red, genetic level proximity not predicted by FACS panel adjacency. The proximities of twenty (of thirty) genetic level cellular populations were predicted by FACS. Widths of lines are inversely proportional to distances.
Extended Data Fig. 4 Gene expression in B. schlosseri cell populations.
a, Enrichment scores of the top nine systems (left), nine tissues (centre) and ten cell types (right) of annotated genes upregulated in CP33, CP34 and CP25 using the GeneAnalytics tool (compared to human). In systems the haematopoietic system has the highest score, in tissues the blood has the highest score, and within the cells, the granulocytes and HSC have the highest scores. b–d, Geneset Activity Analyses using the Gene Expression Commons tool on a mouse haematopoiesis model of different B. schlosseri cell populations. b, Analysis of CP8 (pigment cells) based on 12 significantly upregulated genes, showing that CP8 is part of the haematopoietic system with gene activity resembling known cell types. c, Analysis of CP3 (small cells) based on 235 significantly upregulated genes, showing that this population is not part of the haematopoietic system. On the bottom: enrichment scores of the top ten tissues using CP3 genes by GeneAnalytics tool compared to human; the highest score is for the testes, suggesting that this population is a gonadal population. d, Analysis of CP19 (enriched with morula cells) based on 96 upregulated genes (P < 0.25), showing that CP19 has gene activity resembling cells in the lymphoid lineage using Geneset analysis.
Extended Data Fig. 5 Subendostylar sinuses as an HSC niche.
a, Reduction in DiO fluorescence suggesting cell proliferation, three weeks after transplantation. The experiment was performed once with two pools for each population from five animals each. b, Candidate HSC (cHSC) population and a control population (CP3) were isolated, labelled with DiD and transplanted into CFSE-labelled compatible colonies; in vivo tracing of transplanted cell migration was used to identify niches. There were no cells detected for CP3 (0/4) or the uninjected colonies (0/4) in the subendostylar sinus, whereas 5/6 colonies injected with cHSC cells showed significant localization of the cHSCs to the subendostylar sinus. P = 0.048, Fisher’s exact test, two-tailed. Although in the cell islands 4/4 were positive with CP3, 5/6 positive with cHSC, and 2/4 positive in the uninjected colonies, there are high levels of autofluorescent cells in the cell islands. Full image panels of Fig. 2g. c, Transverse sections of an adult zooid counterstained with toluidine blue (top two left) where the endostyle (green arrowhead) and endo-niche (blue arrowhead) are enlarged (scale bar, 30 μm). Electron microscopy section of the same animal’s endostyle and endo-niche (right and bottom, enlarged). Yellow arrowheads indicate cells with haemoblast (HSC) morphology that are enriched within the endo-niche (scale bar, 5 μm). The experiment was performed three times. Full image panels of Fig. 2b. d, Immunohistochemistry with antibodies against phospho-histone H3 (PHH3), suggesting that there are mitotic cells in the endostyle region in the developing primary bud and also in the adult zooid endostyle.
Extended Data Fig. 6 Gene expression in an HSC niche: the endostyle.
a, Comparison between the transcriptome sequence data from 10 samples of dissected endostyles and the transcriptome data for 34 whole colonies revealed a list of 327 genes that were significantly upregulated in the endostyle and showed homology to genes expressed in human haematopoietic bone marrow. Heat map includes the top 100 (by log(FC)) of the bone-marrow-associated endostyle genes. b, Geneset Activity Analysis of top 200 genes upregulated in the B. schlosseri endostyle associated with the blood system, found by RNA-seq (this study) using the Gene Expression Commons tool on a mouse haematopoiesis model. The enriched populations are bone marrow stromal cells and HSCs. c, Similar analysis, but for C. robusta based on previous in situ work21, revealed enriched mouse bone marrow stromal cells as well, based on 188 genes that are expressed in the C. robusta endostyle and are associated with the blood system.
Extended Data Fig. 7 Discovery of a myeloid lineage phagocytic population.
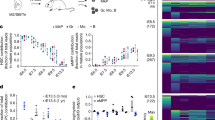
a, FACS analysis of B. schlosseri cells that are fluorescently positive in one of three phagocytosis assays performed: (first and second from left) phagocytosis of green fluorescent beads, (third) phagocytosis of V. diazotrophicus labelled with AF647, and (fourth) allogeneic phagocytosis. Three phagocytic populations were identified: amoebocytes (A), myeloid cells (M), and large phagocytes (LP). The experiment was repeated twice. The myeloid cells were the main contributors to phagocytosis, contributing more than 40% to each of the phagocytosis assays. The large phagocytes contribute mainly to allogeneic phagocytosis compared to the other assays. b, Live images of the three isolated phagocytic populations. The experiment was performed three times. Scale bar, 20 μm. c, We isolated the three main phagocytic populations and a small cell population (CP3) as a control, and incubated each one with fluorescent beads to validate the engulfment capacity of each population. The experiment was repeated twice. Plots show FACS analysis of green fluorescent bead phagocytosis by sorted populations. d, Amoebocytes, myeloid cells, and large phagocytes all had significantly higher engulfment rates than the small cell population. Moreover, amoebocytes and myeloid cells had significantly higher cell percentages than the large phagocyte population. Percentage analysis was carried out on two samples for each sorted population. Unpaired t-test, two-tailed; *P < 0.05, data shown as mean. e, Representative confocal images of the three phagocytic populations after engulfment of beads. Scale bar, 20 μm. f, ImageStream analysis confirmed that the three phagocytic populations assayed engulfed the beads. The positive cells had mainly the morphology of amoebocytes, myeloid cells, or large phagocytes. The experiment was performed once on ImageStream. Representative images of the three phagocytic populations after engulfment of beads. Scale bar, 7 μm.
Extended Data Fig. 8 Cytotoxicity and the two morphs of morula cells at PORs.
a, Cytotoxicity assays of isolated LGL cells compared to small cells, and to isolated morula cells (MCs). In both cases the LGL cells had significantly higher cytotoxicity compared to the other cell types. The experiment with isolated cells was performed twice with triplicates. Unpaired t-test, two-tailed; *P = 0.003, **P = 0.0013; data shown as mean. b, LGL cells were isolated (upper left) and incubated overnight either in syngeneic (upper right) or in allogeneic challenge (lower left). FSC/SSC analysis of LGL cell (lower population) and morula cells (upper population). Insets, sample light microscopy images of the cells after incubation for each treatment. Lower right, analysis of LGL and morula cells in syngeneic compared to allogeneic challenge. The experiment was performed once with duplicates and validated by light microscopy. Bars show mean. c, H&E-stained section of B. schlosseri colonies undergoing rejection. In the ampule (AMP) the inactivated form of cytotoxic morula cells/large granular lymphocyte-like cells (LGL) can be observed (top). On the other hand, the activated form with the brown pigmentation of morula cells can be observed at POR (bottom). d, Confocal imagery of phagocytosis assays to validate the allogeneic engulfment. Colonies are labelled with CFSE in green and with DiD in red after allogeneic phagocytosis assay. Large phagocytic cells can be seen after engulfment of allogeneic cells or vesicles. Validation of allogeneic phagocytosis by confocal imaging was performed twice. Scale bar, 20 μm. e, Example of cytotoxicity assay with different effector to target (E:T) ratios, where the targets are compatible or rejecting colony cells to the effector colony. In the rejecting colony, specific lysis is significantly higher. The experiment was performed three times with triplicates. ANOVA two-factor with replication; *P = 0.0015; data shown as mean ± s.d.
Supplementary information
Supplementary Table 1
Gene expression by populations. Reads per kilobase of transcript per million for each gene of each isolated endpoint population (Extended Data Fig. 1c-d). On the right side is the results of the differential expression analysis for each populations or cluster of populations with adjusted (using Benjamini-Hochberg) p-values (computed using exact tests assuming a negative binomial model) <0.05 and <0.25. Differentially up-regulated=1, differentially down-regulated=-1, no differential expression=0. Each population was analyzed once compared to the rest.
Supplementary Table 2
Genes sets used throughout the study. List of the genes used in the analysis in Gene Commons Tool and the GeneAnalytics. If gene name appears more than once it means that there are more than one B. schlosseri gene model that is associated with this gene and was upregulated.
Supplementary Table 3
GeneAnalytics analysis of 42 genes upregulated in the Botryllus cHSC population that known to be expressed in human hematopoietic stem cells. Analysis include tissues and cells that express these genes and pathways, biological process, molecular functions, phenotypes and diseases that are associated with these genes.
Supplementary Table 4
GeneAnalytics analysis of 327 genes upregulated in the endostyle that known to be expressed in human hematopoietic bone marrow. Analysis include tissues and cells that express these genes and pathways, biological process, molecular functions, phenotypes and diseases that are associated with these genes.
Supplementary Table 5
Data source for the figures. Figure 2. a, Gene list for analysis. c, Pigment cell analysis. f, FACS differentiation analysis. g, Transplantation localization analysis. Figure 3. b, Genes of CP31. e, Allogeneic phagocytosis and cytotoxicity. f, Anti-BHF blocking cytotoxicity. Extended Data Figure 4 gene lists used in the analysis. a, CP25,33,34. b, CP8. c, CP3. d, CP19. Extended Data Figure 6 gene lists used in the analysis. b, Botryllus endostyle. c, Ciona Endostyle. Extended Data Figure 7. d, Beads phagocytosis of sorted cells. Extended Data Figure 8. a, Cytotoxicity of LGL compared to other cells. b, Differentiation from LGL to Morula upon allogeneic challenge. e, Cytotoxicity of compatible and rejecting with different effector to target ratios.
Video 1
Budding cycle and take-over. Three day time lapse acquisition of buds’ development in two B. schlosseri colonies (colony 1 and colony 2). At time lapse’s 35th hour point, the zooids of colony 2 are getting absorbed and replaced by their buds. Taken on a Kyence BZ-X700 every 18 minutes during a 60 hour period.
41586_2018_783_MOESM8_ESM.mp4
Video 2Vascular fusion between two compatible B. schlosseri colonies. Time-lapse acquisition of vascular fusion between two B. schlosseri colonies (colony 1 and colony 2). At the time lapse’s 3 hour and 10 minute mark the ampullae in the center touch, than at the 6 hour 40 minute mark they fuse. Movies of blood flowing from one colony to the other are shown after. Taken on a Kyence BZ-X700 every 10 minutes during an 18 hour period.
Video 3
Differential labeling of colonies. Live colonies labeled on ibidi 35mm micro dishes with either CFSE (green) or cell tracer far red (red). Also an example of fused colonies. Taken by confocal microscopy.
Rights and permissions
About this article
Cite this article
Rosental, B., Kowarsky, M., Seita, J. et al. Complex mammalian-like haematopoietic system found in a colonial chordate. Nature 564, 425–429 (2018). https://doi.org/10.1038/s41586-018-0783-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0783-x
Keywords
This article is cited by
-
Articulating the “stem cell niche” paradigm through the lens of non-model aquatic invertebrates
BMC Biology (2022)
-
Immunoglobulin-like receptors and the generation of innate immune memory
Immunogenetics (2022)
-
Differentiation of endostyle cells by Nkx2-1 and FoxE in the ascidian Ciona intestinalis type A: insights into shared gene regulation in glandular- and thyroid-equivalent elements of the chordate endostyle
Cell and Tissue Research (2022)
-
Multiple Alr genes exhibit allorecognition-associated variation in the colonial cnidarian Hydractinia
Immunogenetics (2022)
-
Crayfish hemocytes develop along the granular cell lineage
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.