Abstract
The arms race between bacteria and the phages that infect them drives the continual evolution of diverse anti-phage defences. Previously described anti-phage systems have highly varied defence mechanisms1,2,3,4,5,6,7,8,9,10,11; however, all mechanisms rely on protein components to mediate defence. Here we report a chemical anti-phage defence system that is widespread in Streptomyces. We show that three naturally produced molecules that insert into DNA are able to block phage replication, whereas molecules that target DNA by other mechanisms do not. Because double-stranded DNA phages are the most numerous group in the biosphere and the production of secondary metabolites by bacteria is ubiquitous12, this mechanism of anti-phage defence probably has a major evolutionary role in shaping bacterial communities.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Cumby, N., Edwards, A. M., Davidson, A. R. & Maxwell, K. L. The bacteriophage HK97 gp15 moron element encodes a novel superinfection exclusion protein. J. Bacteriol. 194, 5012–5019 (2012).
Scholl, D., Adhya, S. & Merril, C. Escherichia coli K1’s capsule is a barrier to bacteriophage T7. Appl. Environ. Microbiol. 71, 4872–4874 (2005).
Molineux, I. J. Host–parasite interactions: recent developments in the genetics of abortive phage infections. New Biol. 3, 230–236 (1991).
Chopin, M. C., Chopin, A. & Bidnenko, E. Phage abortive infection in lactococci: variations on a theme. Curr. Opin. Microbiol. 8, 473–479 (2005).
Tock, M. R. & Dryden, D. T. The biology of restriction and anti-restriction. Curr. Opin. Microbiol. 8, 466–472 (2005).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Goldfarb, T. et al. BREX is a novel phage resistance system widespread in microbial genomes. EMBO J. 34, 169–183 (2015).
Ofir, G. et al. DISARM is a widespread bacterial defence system with broad anti-phage activities. Nat. Microbiol. 3, 90–98 (2018).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Stern, A. & Sorek, R. The phage–host arms race: shaping the evolution of microbes. BioEssays 33, 43–51 (2011).
Seed, K. D. Battling phages: how bacteria defend against viral attack. PLoS Pathog. 11, e1004847 (2015).
Davies, J. Specialized microbial metabolites: functions and origins. J. Antibiot. 66, 361–364 (2013).
Koskella, B. & Brockhurst, M. A. Bacteria–phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol. Rev. 38, 916–931 (2014).
Komatsu, M., Uchiyama, T., Omura, S., Cane, D. E. & Ikeda, H. Genome-minimized Streptomyces host for the heterologous expression of secondary metabolism. Proc. Natl Acad. Sci. USA 107, 2646–2651 (2010).
Vetsigian, K., Jajoo, R. & Kishony, R. Structure and evolution of Streptomyces interaction networks in soil and in silico. PLoS Biol. 9, e1001184 (2011).
Barka, E. A. et al. Taxonomy, physiology, and natural products of Actinobacteria. Microbiol. Mol. Biol. Rev. 80, 1–43 (2015).
Parisi, B. & Soller, A. Studies on the antiphage activity of daunomycin. Giorn. Microbiol. 12, 183–194 (1964).
Morita, J., Tanaka, A., Komano, T. & Oki, T. Inactivation of phage ϕX174 by anthracycline antibiotics, alacinomycin A, doxorubicin and daunorubicin. Agric. Biol. Chem. 43, 2629–2631 (1979).
Lucas, X. et al. StreptomeDB: a resource for natural compounds isolated from Streptomyces species. Nucleic Acids Res. 41, D1130–D1136 (2013).
Thaker, M. N. et al. Identifying producers of antibacterial compounds by screening for antibiotic resistance. Nat. Biotechnol. 31, 922–927 (2013).
Smith, P. B., Tomfohrde, K. M., Rhoden, D. L. & Balows, A. API system: a multitube micromethod for identification of Enterobacteriaceae. Appl. Microbiol. 24, 449–452 (1972).
Furlan, R. L. et al. DNA-binding properties of cosmomycin D, an anthracycline with two trisaccharide chains. J. Antibiot. 57, 647–654 (2004).
Waksman, S. A., Geiger, W. B. & Reynolds, D. M. Strain specificity and production of antibiotic substances. VII. Production of actinomycin by different Actinomycetes. Proc. Natl Acad. Sci. USA 32, 117–120 (1946).
Boulanger, P. & Letellier, L. Ion channels are likely to be involved in the two steps of phage T5 DNA penetration into Escherichia coli cells. J. Biol. Chem. 267, 3168–3172 (1992).
Boulanger, P. & Letellier, L. Characterization of ion channels involved in the penetration of phage T4 DNA into Escherichia coli cells. J. Biol. Chem. 263, 9767–9775 (1988).
Casjens, S. R. & Gilcrease, E. B. Determining DNA packaging strategy by analysis of the termini of the chromosomes in tailed-bacteriophage virions. Methods Mol. Biol. 502, 91–111 (2009).
Ackermann, H. W. 5500 phages examined in the electron microscope. Arch. Virol. 152, 227–243 (2007).
Watve, M. G., Tickoo, R., Jog, M. M. & Bhole, B. D. How many antibiotics are produced by the genus Streptomyces? Arch. Microbiol. 176, 386–390 (2001).
Acknowledgements
We thank G. Wright for providing the WAC strains and D. Bona for technical assistance. We thank A. Davidson for many useful discussions and comments on the manuscript, M. Meneghini and W. Navarre for reading the manuscript and A. Edwards for his long-standing support. This work was supported by operating grants from the Natural Sciences and Engineering Research Council (RGPIN-2017-06898) and Canadian Institutes of Health Research (CIHR; MOP-136845) to K.L.M. and CIHR (MOP-57684) to J.R.N. M.D.-I. was supported by a Studentship from the Canadian Cystic Fibrosis Foundation.
Reviewer information
Nature thanks M. Clokie and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.K. performed and analysed most of the biological experiments, M.D.-I. performed mass spectrometry and analytical profile index 20E analyses, Z.D. performed the high-throughput E. coli drug screen, and S.H., A.I.W., and I.M. assisted with phage isolation and plaque experiments. J.R.N. and K.L.M. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
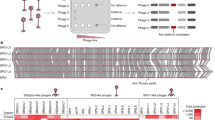
Extended Data Fig. 1 Characterization of Streptomyces phages used in this study.
a, Host range profiles of the Streptomyces phages. Streptomyces phages were isolated from dirt samples from a number of different geographical locations as noted on the right. The host range of these phages was determined by plating on the panel of ten Streptomyces strains listed at the top (n = 3 biological replicates). Green boxes denote strains in which each phage was able to form plaques. Phages φScoe2 and φScoe25 were used in the initial screen of the 48 WAC extracts for anti-phage activity. b, Negatively stained phages were examined using TEM on a Talos L120C. Scale bars are indicatd at the bottom of each image. Each of the phages belongs to the Siphoviridae family, members of which have long non-contractile tails and an icosahedral head that contains a dsDNA genome. TEM grids were prepared once for each phage and two images were taken for each.
Extended Data Fig. 2 Effect of daunorubicin, doxorubicin and spent medium from S. peucetius on phage propagation and S. coelicolor growth.
a, Determination of the minimum inhibitory concentration of doxorubicin on phages φScoe2 (black bars) and φScoe25 (grey bars) propagated in S. coelicolor. Phage titres were determined following propagation in S. coelicolor in the presence of doxorubicin at a range of concentrations that varied from 2.5 μM to 40 μM. Because 10 μM was the lowest concentration at which full inhibition of phage φScoe25 was observed, this was the baseline concentration selected for further experiments. Data are mean ±s.d.; n = 3 independent biological replicates. b, Determination of the minimum inhibitory concentration of daunorubicin on phages φScoe2 (black bars) and φScoe25 (grey bars) propagated in S. coelicolor. Phage titres were determined in S. coelicolor in the presence of daunorubicin at a range of concentrations that varied from 2.5 μM to 40 μM as in a. Data are mean ±s.d.; n = 3 independent biological replicates. c, Determination of the effects of 10 μM daunorubicin, 10 μM doxorubicin and spent medium from S. peucetius that contained doxorubicin and daunorubicin on the growth of S. coelicolor. The number of colony-forming units that are present after overnight growth of S. coelicolor in the presence of each of these compounds shows that they do not significantly decrease the growth of S. coelicolor. Data are mean ±s.d.; n = 3 independent biological replicates. d, Quantification of doxorubicin produced by S. peucetius. A standard curve was generated with commercially available doxorubicin by quantifying the ions that represent the proton adduct of the species (n = 3 independent experiments). Curves were then used to extrapolate the concentration of doxorubicin in S. peucetius culture supernatants after three and four days of growth. e, Quantification of daunorubicin produced by S. peucetius. A standard curve was generated with commercially available daunorubicin by quantifying the ions that represent the proton adduct of the species (n = 3 independent experiments). Curves were used to extrapolate the concentration of daunorubicin in S. peucetius culture supernatants after three and four days of growth. f, Effect of spent mannitol–soy and DNB media from S. peucetius on the propagation of phage φScoe2 after 1–4 days of S. peucetius growth. This graph reflects replicates of the phage spotting assay shown in Fig. 2b. Data are mean ±s.d.; n = 3 independent biological replicates. g, The effects of natural and synthetic compounds on the propagation of phages φScoe2 and φScoe25 were determined by overnight propagation of each phage in S. coelicolor in the presence of each compound at its specified concentration. This graph reflects replicates of the phage spotting assay shown in Fig. 2a. Data are mean ±s.d.; n = 3 independent biological replicates.
Extended Data Fig. 3 Detection of doxorubicin and daunorubicin in spent medium from S. peucetius.
a, Doxorubicin production in mannitol–soy broth was confirmed by ultra-performance liquid chromatography–tandem mass spectrometry with comparison to a doxorubicin standard. The observed fragmentation from a sample grown in doxorubicin-producing permissive medium adhered to the doxorubicin structure with minimal mass errors. Complementary fragments (1 and 6) with their losses highlighted in red further support the doxorubicin structure. In addition, the fragmentation observed from extracts matches a doxorubicin commercial standard (n = 1 experiment). b, Daunorubicin production in mannitol–soy broth was confirmed by ultra-performance liquid chromatography–tandem mass spectrometry with comparison to a daunorubicin standard. The observed fragmentation from a sample grown in daunorubicin-producing permissive medium adhered to the daunorubicin structure with minimal mass errors. Complementary fragments (1 and 6) with their losses highlighted in red further support the daunorubicin structure. In addition, the fragmentation observed from extracts matches a daunorubicin commercial standard (n = 1 experiment).
Extended Data Fig. 4 Effect of active WAC extracts on the growth of S. coelicolor and diversity of secondary metabolites produced.
a, S. coelicolor was incubated overnight in the presence of each of the fourteen WAC extracts that showed anti-phage activity. The majority of extracts had no effect on bacterial cell growth. Two extracts, WAC170 and WAC178, inhibited growth of S. coelicolor approximately tenfold. Data are mean ±s.d.; n = 3 independent biological replicates. Statistically different samples compared to the untreated sample by ANOVA are indicated (from left to right, *P = 0.0117, *P = 0.0188, **P = 0.0022, ****P < 0.0001, **P = 0.0029). b, Growth of the 14 WAC strains for which anti-phage activity was detected on MYM solid medium reveals the diversity of secondary metabolite pigment production (n = 3 biologically independent replicates).
Extended Data Fig. 5 Elucidation of active secondary metabolites in WAC288.
Cosmomycin D was determined to be the active metabolite within the WAC288 extract following a bioactivity-guided fractionation strategy (n = 3 biological replicates). The chromatogram (A494 nm) of the HPLC purification indicates the cosmomycin D fraction collected for bioactivity assays and structure elucidation. The cosmomycin D structure was confirmed by high-resolution tandem mass spectrometry of its associated proton adduct (1,189.5869 m/z). The observed fragmentation supports the cosmomycin D structure with minimal mass errors for each fragment (n = 1 experiment). Key fragment losses are annotated with their associated peak, and their losses are highlighted in red.
Extended Data Fig. 6 Elucidation of active secondary metabolites in WAC240.
The active molecule in the WAC240 extract was determined by a bioactivity-guided fractionation approach (n = 3 biological replicates). The chromatogram (A444 nm) of the final HPLC purification step highlights the actinomycin D fraction collected. Structure elucidation was accomplished via high-resolution tandem mass spectrometry of the actinomycin D proton adduct (1255.6329 m/z) following HPLC purification (n = 1 experiment). All of the fragments support the actinomycin D structure with minimal mass errors, and the key fragment losses are highlighted in the structures in red. Many of the annotated fragments are complementary (for example, 1 and 11), further supporting that the parent ion is actinomycin D.
Extended Data Fig. 7 Effect of daunorubicin on the life cycle of the E. coli phage λ.
a, Determination of the working concentration of daunorubicin for E. coli phage λ. Phage λ was propagated in the presence of daunorubicin at concentrations that varied from 2.5 μM to 40 μM and the resulting phage titres were determined by spotting serial dilutions of the lysate onto a lawn of E. coli BW25113. Because daunorubicin at 15 μM provided a 105-fold decrease in phage titre and was similar in concentration to the 10-μM concentration used in the Streptomyces phage assays, it was chosen as the working concentration for phage λ. Data are mean ±s.d.; n = 3 independent biological replicates. b, Phage λ was propagated in E. coli BW25113 for 6 h in the absence and presence of 15 μM daunorubicin and the resulting phages were enumerated by plating serial dilutions of the supernatant on BW25113. When the phage was pre-incubated with daunorubicin, there was no significant decrease in phage titre. Induction of a temperature-sensitive λ lysogen in the presence of daunorubicin also did not decrease the phage titre. This graph reflects replicates of the phage spotting assay shown in Fig. 3c. Data are mean ±s.d.; n = 3 biological replicates. The statistically different sample compared to the untreated sample by ANOVA is indicated (****P < 0.0001).
Supplementary information
Rights and permissions
About this article
Cite this article
Kronheim, S., Daniel-Ivad, M., Duan, Z. et al. A chemical defence against phage infection. Nature 564, 283–286 (2018). https://doi.org/10.1038/s41586-018-0767-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0767-x
Keywords
This article is cited by
-
Conservation and similarity of bacterial and eukaryotic innate immunity
Nature Reviews Microbiology (2024)
-
Phages overcome bacterial immunity via diverse anti-defence proteins
Nature (2024)
-
Complete genomes and comparative analyses of Streptomyces phages that influence secondary metabolism and sporulation
Scientific Reports (2023)
-
Viral lysing can alleviate microbial nutrient limitations and accumulate recalcitrant dissolved organic matter components in soil
The ISME Journal (2023)
-
The highly diverse antiphage defence systems of bacteria
Nature Reviews Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.