Abstract
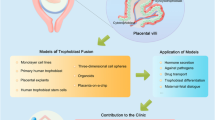
The placenta is the extraembryonic organ that supports the fetus during intrauterine life. Although placental dysfunction results in major disorders of pregnancy with immediate and lifelong consequences for the mother and child, our knowledge of the human placenta is limited owing to a lack of functional experimental models1. After implantation, the trophectoderm of the blastocyst rapidly proliferates and generates the trophoblast, the unique cell type of the placenta. In vivo, proliferative villous cytotrophoblast cells differentiate into two main sub-populations: syncytiotrophoblast, the multinucleated epithelium of the villi responsible for nutrient exchange and hormone production, and extravillous trophoblast cells, which anchor the placenta to the maternal decidua and transform the maternal spiral arteries2. Here we describe the generation of long-term, genetically stable organoid cultures of trophoblast that can differentiate into both syncytiotrophoblast and extravillous trophoblast. We used human leukocyte antigen (HLA) typing to confirm that the organoids were derived from the fetus, and verified their identities against four trophoblast-specific criteria3. The cultures organize into villous-like structures, and we detected the secretion of placental-specific peptides and hormones, including human chorionic gonadotropin (hCG), growth differentiation factor 15 (GDF15) and pregnancy-specific glycoprotein (PSG) by mass spectrometry. The organoids also differentiate into HLA-G+ extravillous trophoblast cells, which vigorously invade in three-dimensional cultures. Analysis of the methylome reveals that the organoids closely resemble normal first trimester placentas. This organoid model will be transformative for studying human placental development and for investigating trophoblast interactions with the local and systemic maternal environment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Microarray data for Fig. 2 and Extended Data Figs. 4 and 5 have been deposited in the ArrayExpress database at EMBL-EBI under accession number E-MTAB-6683. Illumina EPIC methylation array data for Fig. 2 and Extended Data Fig. 5 have been deposited in the ArrayExpress database at EMBL-EBI under accession number E-MTAB-7204. The mass spectrometry proteomics data for Extended Data Table 3 and Supplementary Table 2 have been deposited to the ProteomeXchange Consortium via the PRIDE36 partner repository (https://www.ebi.ac.uk/pride/archive/) with the dataset identifier PXD009118 and 10.6019/PXD009118. All other data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Burton, G. J. & Jauniaux, E. What is the placenta? Am. J. Obs. Gynecol 213, S6.e1–S6.e8 (2015).
Hamilton, W. J. & Boyd, J. D. Development of the human placenta in the first three months of gestation. J. Anat. 94, 297–328 (1960).
Lee, C. Q. et al. What is trophoblast? A combination of criteria define human first-trimester trophoblast. Stem Cell Reports 6, 257–272 (2016).
Kunath, T. et al. Developmental differences in the expression of FGF receptors between human and mouse embryos. Placenta 35, 1079–1088 (2014).
Paiva, P. et al. Human chorionic gonadotrophin regulates FGF2 and other cytokines produced by human endometrial epithelial cells, providing a mechanism for enhancing endometrial receptivity. Hum. Reprod. 26, 1153–1162 (2011).
Hempstock, J., Cindrova-Davies, T., Jauniaux, E. & Burton, G. J. Endometrial glands as a source of nutrients, growth factors and cytokines during the first trimester of human pregnancy: a morphological and immunohistochemical study. Reprod. Biol. Endocrinol. 2, 50 (2004).
Mühlhauser, J. et al. Differentiation and proliferation patterns in human trophoblast revealed by c-erbB-2 oncogene product and EGF-R. J. Histochem. Cytochem. 41, 165–173 (1993).
Sonderegger, S., Husslein, H., Leisser, C. & Knofler, M. Complex expression pattern of Wnt ligands and frizzled receptors in human placenta and its trophoblast subtypes. Placenta 28 (Suppl.), S97–S102 (2007).
Haider, S., Kunihs, V., Fiala, C., Pollheimer, J. & Knöfler, M. Expression pattern and phosphorylation status of Smad2/3 in different subtypes of human first trimester trophoblast. Placenta 57, 17–25 (2017).
Lee, C. Q. E. et al. A niche of trophoblast progenitor cells identified by integrin α2 is present in first trimester human placentas. Development 145, (2018).
Turco, M. Y. et al. Long-term, hormone-responsive organoid cultures of human endometrium in a chemically defined medium. Nat. Cell Biol. 19, 568–577 (2017).
Cambuli, F. et al. Epigenetic memory of the first cell fate decision prevents complete ES cell reprogramming into trophoblast. Nat. Commun. 5, 5538 (2014).
Yu, C. et al. GCMa regulates the syncytin-mediated trophoblastic fusion. J. Biol. Chem. 277, 50062–50068 (2002).
Frendo, J. L. et al. Direct involvement of HERV-W Env glycoprotein in human trophoblast cell fusion and differentiation. Mol. Cell. Biol. 23, 3566–3574 (2003).
Jauniaux, E., Hempstock, J., Teng, C., Battaglia, F. C. & Burton, G. J. Polyol concentrations in the fluid compartments of the human conceptus during the first trimester of pregnancy: maintenance of redox potential in a low oxygen environment. J. Clin. Endocrinol. Metab. 90, 1171–1175 (2005).
Petry, C. J. et al. GDF15 concentrations in maternal serum associated with vomiting in pregnancy: the Cambridge baby growth study. Preprint at https://www.bioRxiv.org/content/early/2017/11/17/221267 (2017).
Pijnenborg, R., Vercruysse, L. & Hanssens, M. The uterine spiral arteries in human pregnancy: facts and controversies. Placenta 27, 939–958 (2006).
Okae, H. et al. Derivation of human trophoblast stem cells. Cell Stem Cell 22, 50–63.e6 (2018).
Haider, S. et al. Self-renewing trophoblast organoids recapitulate the developmental program of the early human placenta. Stem Cell Reports 11, 537–551 (2018).
Robbins, J. R. & Bakardjiev, A. I. Pathogens and the placental fortress. Curr. Opin. Microbiol. 15, 36–43 (2012).
Burton, G. J., Jauniaux, E. & Charnock-Jones, D. S. The influence of the intrauterine environment on human placental development. Int. J. Dev. Biol. 54, 303–312 (2010).
Moffett, A. & Colucci, F. Co-evolution of NK receptors and HLA ligands in humans is driven by reproduction. Immunol. Rev. 267, 283–297 (2015).
Smith, G. C. First-trimester determination of complications of late pregnancy. J. Am. Med. Assoc. 303, 561–562 (2010).
Turco, M. Y. et al. Establishment and differentiation of long-term trophoblast organoid cultures from the human placenta. Protoc. Exchange https://doi.org/10.1038/protex.2018.121 (2018).
Nollet, F., Billiet, J., Selleslag, D. & Criel, A. Standardisation of multiplex fluorescent short tandem repeat analysis for chimerism testing. Bone Marrow Transplant. 28, 511–518 (2001).
Mossallam, G. I., Smith, A. G. & McFarland, C. Comparison of variable number tandem repeat and short tandem repeat genetic markers for qualitative and quantitative chimerism analysis post allogeneic stem cell transplantation. J. Egypt. Natl. Canc. Inst. 17, 103–113 (2005).
Turner, T. R. et al. Single molecule real-time DNA sequencing of HLA genes at ultra-high resolution from 126 International HLA and Immunogenetics Workshop cell lines. HLA 91, 88–101 (2018).
Hemberger, M., Udayashankar, R., Tesar, P., Moore, H. & Burton, G. J. ELF5-enforced transcriptional networks define an epigenetically regulated trophoblast stem cell compartment in the human placenta. Hum. Mol. Genet. 19, 2456–2467 (2010).
Morris, T. J. et al. ChAMP: 450k Chip Analysis Methylation Pipeline. Bioinformatics 30, 428–430 (2014).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Zhou, W., Laird, P. W. & Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 45, e22 (2017).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013).
Fortin, J.-P., Triche, T. J. Jr & Hansen, K. D. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 33, 558–560 (2017).
Karolchik, D. et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 32, D493–D496 (2004).
Kay, R. G., Galvin, S., Larraufie, P., Reimann, F. & Gribble, F. M. Liquid chromatography/mass spectrometry based detection and semi-quantitative analysis of INSL5 in human and murine tissues. Rapid Commun. Mass Spectrom. 31, 1963–1973 (2017).
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, 11033 (2016).
Zhang, J. et al. PEAKS DB: de novo sequencing assisted database search for sensitive and accurate peptide identification. Mol. Cell. Proteomics 11, M111.010587 (2012).
Acknowledgements
The authors are grateful to patients for donating tissue for research. We thank D. Moore and staff at the Addenbrooke’s Hospital, Cambridge; J. Bauer, E. Clemente, E. Farnell, C. Reitter from Cambridge Genomic Services, Department of Pathology; K. Burling and Core Biochemical Assay Laboratory staff at NIHR Cambridge Biomedical Research Centre; I. Simonic; and T. Cindrova-Davies, H. W. Yung, B. D. Simons, B. K. Koo, M. Huch, Y. W. Loke and all members of the Moffett and Burton laboratories. This work was funded by Centre for Trophoblast Research, University of Cambridge and Medical Research Council (MR/L020041/1). A. Moffett holds a Wellcome Trust investigator award (200841/Z/16/Z). M.Y.T. received funding from EU Seventh Framework Programme for research, technological development and demonstration (PIEF-GA-2013-629785) and now holds a Royal Society Dorothy Hodgkin Fellowship. F.R. and F.M.G. hold Wellcome Trust joint investigator awards (106262/Z/14/Z and 106263/Z/14/Z), and funding from MRC-Metabolic Diseases Unit (MRC_MC_UU_12012/3), NIHR-Cambridge BRC and AZ/MedImmune.
Reviewer information
Nature thanks M. Roberts and the anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.Y.T. and L.G. designed and carried out all the core experiments and analysed the data; R.G.K. performed LC–MS/MS analysis; R.S.H. and M.P. performed microarray and DNA methylation analysis; A. McWhinnie and S.G.E.M. performed and analysed the STR and HLA typing; M.S.H. performed electron microscopy and confocal analysis; R.F. and H.S. assisted with IHC and ELISA experiments; L.E. performed qPCR experiments; A.S., F.M.G., F.R., M.H. and S.O. provided intellectual input to the study; M.H. performed the ELF5 methylation experiment; G.J.B. and A. Moffett assisted in analysis and interpretation of results and jointly supervised the project. M.Y.T., G.J.B. and A. Moffett wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Staining for signalling pathways in first trimester placenta and decidua.
a, Immunohistochemical staining of first trimester placenta (6–8 weeks gestation) for effectors of the following major signalling pathways: (1) WNT signalling through non-phosphorylated-β-catenin (S33/S37/T41); (2) TGFβ signalling through phosphorylated (p)-SMAD2 (S465/467) and p-SMAD3 (S423/425); (3) MAPK signalling through p-ERK1 and p-ERK2 (T202/Y204); and (4) p-STAT3 (Y705) signalling. Scale bars, 50 μm. Representative images from n = 8 for each antibody. BMP signalling through SMAD1, SMAD5 or SMAD8 was not possible to assess by immunohistochemistry. VCT cells and CCCs displayed membrane-localized staining of non-phosphorylated-β-catenin, whereas p-ERK1 and p-ERK2 was mostly found in the cytoplasm in both cell types. Cytoplasmic and nuclear signals for p-ERK1 and p-ERK2 were detected in the EVT. p-SMAD2 and p-SMAD3 staining also showed stronger nuclear signals in the EVT, suggesting a role for TGFβ signalling in differentiation, in accordance with a previous report9. Phosphorylated nuclear STAT3 was detected only in the EVT, again indicating involvement in their differentiation. The SCT was negative for all these signals. b, Summary of findings from a. Trophoblast cells from different regions of the placenta are represented as a circle with a nucleus (small inner circle). Black indicates strong staining, grey indicates faint staining, and white denotes not detected. Thicker circles indicate staining localized to the cell membrane. c, In situ hybridization for LGR5 on first trimester placental villi. LGR5 transcripts are detected in the VCT. Stroma is negative. Positive-control probe is for UBC; negative-control probe is for the bacterial gene dapB. Nuclei are counterstained with haematoxylin. Images are at ×10 magnification. Representative images from n = 2. d, Immunohistochemistry staining for R-spondin 1 in early first trimester (6–8 weeks gestation) and late first trimester (10–12 weeks gestation) decidual samples. Representative images from n = 2 for each tissue type. Images are at ×20 magnification. UG, uterine glands.
Extended Data Fig. 2 Culture components tested for the establishment of long-term organoid cultures of trophoblast from human placentas.
a, Growth factors (HGF, PGE2, Y-27632 and nicotinamide) were added as supplements to basal TOM that contains EGF, CHIR99021, R-spondin 1, A83-01 and FGF2 (Supplementary Table 1a, b). Bright-field images of placental digests at passage 1, day 7. The cystic structures that appear with the addition of nicotinamide (red asterisks) are contaminating maternal glandular organoids. Representative images from n = 2. Conditions containing factors that did not show growth are not included. Scale bars, 500 μm. b, Trophoblast organoid cultures at passages 2 and 10 with continuous culture. Representative images from n = 3. Scale bars, 500 μm. c, Analysis of genetic stability of cultures (n = 2) with comparative genomics hybridization (CGH) array. Shown is a representative whole-genome array CGH plot generated with Agilent Cytogenomics software. Genomic DNA from late passage (passage 8) trophoblast organoids is compared to genomic DNA from an early passage (passage 2). Each spot is a single probe. Plotted are the log ratios of the average signal intensity of each probe on the y axis along its position on the chromosomes (1–22, X and Y) on the x axis. A log signal ratio of 0 represents equivalent copy number in the samples. No significant DNA copy number abnormalities were identified. d, Live imaging of trophoblast organoid cultures (n = 2) passaged for more than 6 months and then frozen, thawed and exposed to Mitotracker Red. Functional mitochondria are visible showing that the cells are healthy (white arrowheads). Scale bars, 50 μm (whole organoid) and 10 μm (individual cells). e, Organoids derived from the same placental cell isolate using either trophoblast or decidual organoid medium (TOM or ExM, respectively) demonstrate that matched placental (fetal) and decidual (maternal) organoids can be derived from one sample. Representative bright-field images from n = 3. Scale bars, 500 μm.
Extended Data Fig. 3 Trophoblast organoids retain characteristic features of first trimester trophoblast in vivo.
a, Immunofluorescence images by confocal microscopy of GATA3, KRT7, EGFR and DAPI on trophoblast organoids (representative images from n = 3). EGFR stains both VCT cells and the surface of the SCT, as reported in vivo7. The basement membrane is around the outside of the organoids, with syncytial masses present in the centre. Scale bar, 50 μm. b, Immunohistochemical staining for the transcription factor TFAP2A shows uniform expression on trophoblast organoids (representative image from n = 20). Scale bars, 50 μm. c, Gating strategy used for flow cytometric analysis of single, live cells from trophoblast organoids. d, qPCR analysis of ELF5 in trophoblast organoids (n = 5) compared to whole placental villi (n = 8) and placental stromal cells (n = 5). Graph shows expression levels relative to the geometric mean of the three housekeeping genes TBP, TOP1 and HPRT1. The mean ELF5 expression is shown for each sample group. e, Bisulfite sequencing of the ELF5 promoter region (−379 bp to −28 bp upstream of the transcription start site) of trophoblast organoids from two different placentas (TOrg_3 and TOrg_6) and matched maternal blood leukocytes (positive control). The relative percentage of methylated cytosine residues (filled circles) is indicated. f, qPCR analysis for miRNAs miR525-3p, miR526-3p and miR517-5p from the C19MC cluster on trophoblast organoids (n = 6), choriocarcinoma lines JEG-3 and JAR cells (positive controls) and PBMC (low expression/negative control). Graph shows relative expression levels of each organoid culture to the housekeeping gene RNU48.
Extended Data Fig. 4 Hierarchical clustering of microarray data comparing placental villi, trophoblast organoids and placental stromal cells.
a, Unsupervised hierarchical clustering analysis of global gene expression profiles by microarray of first trimester placental villi (Pl) (n = 8), trophoblast organoids (TOrg) (n = 5), placental stromal cells (Str) (n = 5) and decidual organoids (DOrg) (n = 3). Analysis was based on 12,673 probes. The expression profiles of trophoblast organoids cluster with first trimester placental samples, whereas decidual organoids and placental stromal cells cluster in a separate tree. b, The top 20 genes contributing to PC1 and PC2 in the principal component analysis plot from Fig. 2e. The top genes contributing to PC1 are trophoblast-specific genes such as CGB3, GATA3 and PSG6, indicating that these genes separate the trophoblast organoid and placental villous samples from the two potentially contaminating, non-trophoblast samples (decidual organoids and placental stromal cells). The top genes contributing to PC2 are epithelial genes such as CLDN3, TACSTD2 and KRT23. The organoids only contain trophoblast, but cells of the villous core (stromal, Hofbauer and endothelial cells) are also present in the placental samples. c, Immunohistochemistry of placental villi and trophoblast organoids stained for KRT23 shows expression in all trophoblast cells in vivo and in vitro. The experiment was repeated independently three times. Scale bars, 50 μm and 20 μm (insets). d, Clustered heat map of differentially expressed genes between trophoblast organoids, placental villi and placental stroma with an absolute base-2 logarithmic fold change of 2 (adjusted P < 0.05). e, Immunohistochemistry of placental villi and trophoblast organoids stained for CCNE1 shows expression in trophoblast cells in vivo and in vitro. Scale bars, 50 μm and 20 μm (insets). The experiment was repeated independently three times.
Extended Data Fig. 5 Expression profiles of transcription factors in trophoblast organoids and placental villi.
a, Heat map highlighting transcription factors from the differentially expressed genes between placental villi, trophoblast organoids and placental stromal cells. b, Heat map of genes from the ELF family of transcription factors and syncytial genes, GCM1 and ERVW-1. ELF3 and ELF5 both show moderate expression levels across the organoids and placental samples, and very low or no expression in the stromal samples. ELF4 and ELF1 are similar in all samples. There is very high expression of ELF1 in placentas and organoids. Similarly, both ERVW-1 and GCM1 are expressed at higher levels in placentas and organoids in agreement with qPCR data (Fig. 3d). c, Genomic mapping of the methylation array probes to the ELF5 gene. The height of the bars indicates the methylation level from 0 (unmethylated) to 1.0 (fully methylated). d, Methylation of the ELF5 promoter shows hypomethylation in the organoid and placental samples. e, Distribution of methylcytosine across the promoters of the 10 trophoblast gatekeeper genes identified previously in mice12. The organoids (Org) and placental (Pl) samples show very similar methylation patterns across all 10 gene promoters that are distinct from the control brain (CL) and maternal blood (MB). Box plots comprise minimum: 1.5× interquartile range, bottom: first quartile, middle: median, top: third quartile, maximum: 1.5× interquartile range. *P < 0.01, Pearson’s correlation coefficient. f, Table for results in e showing Pearson’s correlation coefficient (R), number of CpG islands or probes compared and P values. g, Chord plot representing terms from the Gene Ontology analysis of upregulated genes in trophoblast organoids.
Extended Data Fig. 6 Trophoblast organoids proliferate and structurally resemble first trimester placental villi.
a, A schematic diagram of a normal placental villus in vivo compared to a trophoblast organoid. The basement membrane (BM) beneath the VCT is contiguous with the stromal villous core in vivo and with the Matrigel in vitro. The SCT contacts maternal blood in the intervillous space in vivo, and forms in the centre of the organoids. b, Immunohistochemical staining for TP63 in first trimester placenta and trophoblast organoids (representative images from n = 14). TP63 is expressed in VCT. Scale bars, 50 μm and 20 μm (insets). c, Representative images of TP63, Ki67 and DAPI staining in trophoblast organoids by confocal microscopy (n = 3). Cells on the outside of the organoids are TP63+ and Ki67+. Scale bars, 20 μm. d, Confocal microscopy images of trophoblast organoid stained for EdU, EPCAM and DAPI, showing fewer proliferating cells (white arrowheads) as the organoids enlarge. Scale bars, 50 μm. Representative images from n = 3. e, Immunohistochemical staining for the SCT markers CD46 and CD71 in first trimester placenta and trophoblast organoids (representative images from n = 20). CD46 and CD71 stain the syncytial brush border. Scale bars, 50 μm and 20 μm (insets). f, Carnegie stage 5b embryo (approximately 9 days after fertilization) from the Carnegie Collection at the early lacunar stage (number 8171). Courtesy of A. Enders and the Centre for Trophoblast Research (https://www.trophoblast.cam.ac.uk/Resources/enders). Arrows point to examples of cavities that appear in the primitive syncytium owing to fluid accumulation before the coelomic cavity and the embryo have fully developed. g, Similar cavities in placental tissue samples from first trimester (6–9 weeks gestation) and in syncytium in the centre of trophoblast organoids. Boxed areas are shown at higher magnification (bottom). Scale bars, 200 μm (top) and 50 μm (bottom). Similar morphology seen in at least five early placental villi and in all organoids. ICM, inner cell mass; Pr.Syn., primitive syncytium; Str, stromal core.
Extended Data Fig. 7 Trophoblast organoids grown in TOM and EVTM.
a, Confocal image of organoid stained for F-actin, DAPI and HLA-G with a merged image. A few isolated cells stain for HLA-G (arrowheads) at the periphery of the organoid. Scale bar, 50 μm. Representative images from n = 3. b, Phase-contrast images from time-lapse videos of trophoblast organoids grown for 16 h in TOM when EVT differentiation does not occur (top) and no invasive cells are visible. An organoid (middle) and a placental villous explant (bottom) exposed to EVTM are shown for comparison. Arrows indicate cells migrating, and arrowheads denote the visible tracks made as the cells invade through the Matrigel. For time-lapse videos of these cultures, see Supplementary Videos 1, 4 and 5. Scale bars, 200 μm.
Supplementary information
Supplementary Methods
Detailed methods for GDF15 ELISA, preparation of samples for LC-MS/MS analysis of trophoblast organoid supernatants and reverse transcription and real-time quantitative RT-PCR (qRT-PCR)
Supplementary Table 1
Derivation of human trophoblast organoids. a, Reagents tested for derivation of human trophoblast organoids. Each reagent was added to basal trophoblast organoid medium (EGF, CHIR99021, A83-01, FGF2, R-spondin 1) on placental digests in triplicate for each culture. Overall growth of cultures (by appearance of organoid/spheroid like structures) were compared to basal condition medium and scored as positive, mildly positive, no effect or negative effect at 7 days after first passage of cultures. Negative effect may be due to either inhibition of proliferation or stimulation of differentiation, or both. Summary of results are based on experiments repeated with at least 2 independent placental digests. References for the source of growth factors and their targets in trophoblast refer to human reproductive tissues during the first trimester only. Abbreviations: na, not applicable; VCT, villous cytotrophoblast; CCC, cell column cytotrophoblast; EVT, extravillous trophoblast; SCT, syncytiotrophoblast. b, Medium composition of Trophoblast Organoid Medium (TOM) for the establishment and long term culture of trophoblast organoids from human placental samples. c, Summary of derived trophoblast organoid cultures and their characterisation. d, Information on antibodies used for this study. Abbreviations: TOrg, Trophoblast organoids; g.a., gestational age; LC-MS. Liquid chromatography mass spectometry; EVT, extravillous trophoblast
Supplementary Table 2
LC-MS/MS analysis of trophoblast organoids supernatants. Result table showing proteins identified by the PEAKS software in a, the trophoblast organoid medium (TOM) and the trophoblast organoid supernatants (b-g). Search results for each corresponding trophoblast organoid supernatants (n=6) are b, TOrg_2; c, TOrg_3; d, TOrg_5; e, TOrg_10; f, TOrg_12; g, TOrg_14. Table h contains a summary of all peptides identified (Supp Tables 2a-g). Protein group and protein ID columns indicate if peptide matches can be assigned to more than one protein. The accession refers to the Swissprot database protein identifier code. The -10lgP is the degree of confidence that the PEAKS software has assigned each peptide match, and protein -10lgP is the weighted sum of the peptides assigned to that protein. Coverage% refers to the proportion of the primary amino acid sequence of each protein that is identified in the experiment. Intensity refers to the relative abundance of each protein, as assigned by the PEAKS software spectral counting approach. #Peptides refers to the number of peptide matches assigned to a protein, whilst #Unique refers to the number of peptides that are assigned solely to that protein group. #Spec refers to the number of peptide MS/MS spectra matched against a particular protein. PTM refers to an of the specified post translational modifications that were identified within a peptide/protein match. In table h, peptide sequence indicates the amino acids in a particular protein, and whether it is a unique hit or not. The mass of each peptide is assigned in daltons, along with the accuracy of the peptide mass assignment compared to the theoretical mass (measured in ppm). The intensity of each peptide match in the samples a-g and the number of mass spectra for each peptide are displayed. The amino acid position of the start and end of each peptide is also included. The PTM’s assigned to each peptide is displayed, as well as the Ascore – the confidence assigned to the PTM match by the PEAKS software
Supplementary Video 1. Generation and migration of HLAG+ cells from trophoblast organoids
Trophoblast organoids placed in EVT differentiation medium (EVTM) generate HLAG+ cells. These cells migrate through Matrigel in 3D to form tracks. Time lapse microscopy taken at day 7 of culture with an image taken every 30 min for between 16-20 h. Representative video of trophoblast organoid in EVTM from three independently derived cultures.
Supplementary Video 2. Generation and migration of HLAG+ cells from trophoblast organoids
Trophoblast organoids placed in EVT differentiation medium (EVTM) generate HLAG+ cells. These cells migrate through Matrigel in 3D to form tracks. Time lapse microscopy taken at day 7 of culture with an image taken every 30 min for between 16-20 h. Representative video of trophoblast organoid in EVTM from three independently derived cultures.
Supplementary Video 3. Growth of trophoblast organoids
Trophoblast organoids grown in trophoblast organoid medium (TOM). Time lapse microscopy taken at day 7 of culture with an image taken every 30 min for between 16-20 h. Representative video of trophoblast organoid in TOM from three independently derived cultures.
Supplementary Video 4. Growth of trophoblast organoids
Trophoblast organoids grown in trophoblast organoid medium (TOM). Time lapse microscopy taken at day 7 of culture with an image taken every 30 min for between 16-20 h. Representative video of trophoblast organoid in TOM from three independently derived cultures.
Supplementary Video 5. Generation and migration of HLAG+ cells from placental villous explants
Placental villi isolated from samples at gestational age 6-9 weeks were placed in EVT differentiation medium (EVTM). Cells that migrate through Matrigel in 3D to form tracks are present. Time-lapse microscopy taken at day 7 of culture with an image taken every 30 min for between 16-20 h. Experiment was repeated independently with two different patient samples.
Supplementary Video 6. Generation and migration of HLAG+ cells from placental villous explants
Placental villi isolated from samples at gestational age 6-9 weeks were placed in EVT differentiation medium (EVTM). Cells that migrate through Matrigel in 3D to form tracks are present. Time-lapse microscopy taken at day 7 of culture with an image taken every 30 min for between 16-20 h. Experiment was repeated independently with two different patient samples.
Rights and permissions
About this article
Cite this article
Turco, M.Y., Gardner, L., Kay, R.G. et al. Trophoblast organoids as a model for maternal–fetal interactions during human placentation. Nature 564, 263–267 (2018). https://doi.org/10.1038/s41586-018-0753-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0753-3
This article is cited by
-
Exposure to high dose of polystyrene nanoplastics causes trophoblast cell apoptosis and induces miscarriage
Particle and Fibre Toxicology (2024)
-
Trophoblast stem cell-based organoid models of the human placental barrier
Nature Communications (2024)
-
Unraveling the molecular mechanisms driving enhanced invasion capability of extravillous trophoblast cells: a comprehensive review
Journal of Assisted Reproduction and Genetics (2024)
-
Deciphering the Epigenetic Landscape: Placental Development and Its Role in Pregnancy Outcomes
Stem Cell Reviews and Reports (2024)
-
Maternal obesity and placental function: impaired maternal–fetal axis
Archives of Gynecology and Obstetrics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.