Abstract
The proteasome is an ATP-dependent, 2.5-megadalton molecular machine that is responsible for selective protein degradation in eukaryotic cells. Here we present cryo-electron microscopy structures of the substrate-engaged human proteasome in seven conformational states at 2.8–3.6 Å resolution, captured during breakdown of a polyubiquitylated protein. These structures illuminate a spatiotemporal continuum of dynamic substrate–proteasome interactions from ubiquitin recognition to substrate translocation, during which ATP hydrolysis sequentially navigates through all six ATPases. There are three principal modes of coordinated hydrolysis, featuring hydrolytic events in two oppositely positioned ATPases, in two adjacent ATPases and in one ATPase at a time. These hydrolytic modes regulate deubiquitylation, initiation of translocation and processive unfolding of substrates, respectively. Hydrolysis of ATP powers a hinge-like motion in each ATPase that regulates its substrate interaction. Synchronization of ATP binding, ADP release and ATP hydrolysis in three adjacent ATPases drives rigid-body rotations of substrate-bound ATPases that are propagated unidirectionally in the ATPase ring and unfold the substrate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-9215 (the combined EA refined with CP-ATPase mask), EMD-9216 (whole EA1), EMD-9217 (whole EA2), EMD-9218 (whole EB), EMD-9219 (whole EC1), EMD-9220 (whole EC2), EMD-9221 (whole ED1), EMD-9222 (whole ED2), EMD-9223 (RP of EA1), EMD-9224 (RP of EA2), EMD-9225 (RP of EB), EMD-9226 (RP of EC1), EMD-9227 (RP of EC2), EMD-9228 (RP of ED1) and EMD-9229 (RP of ED2). Each EMDB entry includes three maps: (1) the low-pass-filtered map without amplitude correction as a default; (2) the low-pass-filtered map with amplitude correction by a negative B-factor shown in Extended Data Table 1; and (3) the raw map without any post-processing such as low-pass-filtering and amplitude correction. Coordinates are available from the RCSB Protein Data Bank under accession codes 6MSB (whole EA1), 6MSD (whole EA2), 6MSE (whole EB), 6MSG (whole EC1), 6MSH (whole EC2), 6MSJ (whole ED1) and 6MSK (whole ED2). Raw data are available from the corresponding author upon request.
References
Livneh, I., Cohen-Kaplan, V., Cohen-Rosenzweig, C., Avni, N. & Ciechanover, A. The life cycle of the 26S proteasome: from birth, through regulation and function, and onto its death. Cell Res. 26, 869–885 (2016).
Finley, D., Chen, X. & Walters, K. J. Gates, channels, and switches: elements of the proteasome machine. Trends Biochem. Sci. 41, 77–93 (2016).
Bard, J. A. M. et al. Structure and function of the 26S proteasome. Annu. Rev. Biochem. 87, 697–724 (2018).
Groll, M. et al. Structure of 20S proteasome from yeast at 2.4 Å resolution. Nature 386, 463-471 (1997).
Lu, Y. et al. Conformational landscape of the p28-bound human proteasome regulatory particle. Mol. Cell 67, 322–333.e6 (2017).
Shi, Y. et al. Rpn1 provides adjacent receptor sites for substrate binding and deubiquitination by the proteasome. Science 351, aad9421 (2016).
Verma, R., Oania, R., Graumann, J. & Deshaies, R. J. Multiubiquitin chain receptors define a layer of substrate selectivity in the ubiquitin-proteasome system. Cell 118, 99–110 (2004).
Verma, R. et al. Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26S proteasome. Science 298, 611–615 (2002).
Smith, D. M., Fraga, H., Reis, C., Kafri, G. & Goldberg, A. L. ATP binds to proteasomal ATPases in pairs with distinct functional effects, implying an ordered reaction cycle. Cell 144, 526–538 (2011).
Chen, S. et al. Structural basis for dynamic regulation of the human 26S proteasome. Proc. Natl Acad. Sci. USA 113, 12991–12996 (2016).
Zhu, Y. et al. Structural mechanism for nucleotide-driven remodeling of the AAA-ATPase unfoldase in the activated human 26S proteasome. Nat. Commun. 9, 1360 (2018).
Eisele, M. R. et al. Expanded coverage of the 26S proteasome conformational landscape reveals mechanisms of peptidase gating. Cell Rep. 24, 1301–1315 (2018).
Wehmer, M. et al. Structural insights into the functional cycle of the ATPase module of the 26S proteasome. Proc. Natl Acad. Sci. USA 114, 1305–1310 (2017).
Schweitzer, A. et al. Structure of the human 26S proteasome at a resolution of 3.9 Å. Proc. Natl Acad. Sci. USA 113, 7816–7821 (2016).
Lasker, K. et al. Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach. Proc. Natl Acad. Sci. USA 109, 1380–1387 (2012).
Lander, G. C. et al. Complete subunit architecture of the proteasome regulatory particle. Nature 482, 186–191 (2012).
da Fonseca, P. C., He, J. & Morris, E. P. Molecular model of the human 26S proteasome. Mol. Cell 46, 54–66 (2012).
Huang, X., Luan, B., Wu, J. & Shi, Y. An atomic structure of the human 26S proteasome. Nat. Struct. Mol. Biol. 23, 778–785 (2016).
Luan, B. et al. Structure of an endogenous yeast 26S proteasome reveals two major conformational states. Proc. Natl Acad. Sci. USA 113, 2642–2647 (2016).
Beck, F. et al. Near-atomic resolution structural model of the yeast 26S proteasome. Proc. Natl Acad. Sci. USA 109, 14870–14875 (2012).
Śledź, P. et al. Structure of the 26S proteasome with ATP-γS bound provides insights into the mechanism of nucleotide-dependent substrate translocation. Proc. Natl Acad. Sci. USA 110, 7264–7269 (2013).
Ding, Z. et al. High-resolution cryo-EM structure of the proteasome in complex with ADP-AlFx. Cell Res. 27, 373-385 (2017).
Saeki, Y., Isono, E. & Toh-E, A. Preparation of ubiquitinated substrates by the PY motif-insertion method for monitoring 26S proteasome activity. Methods Enzymol. 399, 215–227 (2005).
Verma, R., McDonald, H., Yates, J. R., III & Deshaies, R. J. Selective degradation of ubiquitinated Sic1 by purified 26S proteasome yields active S phase cyclin-Cdk. Mol. Cell 8, 439–448 (2001).
Lu, Y., Lee, B. H., King, R. W., Finley, D. & Kirschner, M. W. Substrate degradation by the proteasome: a single-molecule kinetic analysis. Science 348, 1250834 (2015).
Galan, J.-M. & Haguenauer-Tsapis, R. Ubiquitin Lys63 is involved in ubiquitination of a yeast plasma membrane protein. EMBO J. 16, 5847-5854 (1997).
Whitby, F. G. et al. Structural basis for the activation of 20S proteasomes by 11S regulators. Nature 408, 115-120 (2000).
Zhang, F. et al. Structural insights into the regulatory particle of the proteasome from Methanocaldococcus jannaschii. Mol. Cell 34, 473–484 (2009).
Worden, E. J., Dong, K. C. & Martin, A. An AAA motor-driven mechanical switch in Rpn11 controls deubiquitination at the 26S proteasome. Mol. Cell 67, 799–811.e8 (2017).
Pathare, G. R. et al. Crystal structure of the proteasomal deubiquitylation module Rpn8–Rpn11. Proc. Natl Acad. Sci. USA 111, 2984–2989 (2014).
Worden, E. J., Padovani, C. & Martin, A. Structure of the Rpn11–Rpn8 dimer reveals mechanisms of substrate deubiquitination during proteasomal degradation. Nat. Struct. Mol. Biol. 21, 220–227 (2014).
Dambacher, C. M., Worden, E. J., Herzik, M. A., Martin, A. & Lander, G. C. Atomic structure of the 26S proteasome lid reveals the mechanism of deubiquitinase inhibition. eLife 5, e13027 (2016).
Zhang, X. & Wigley, D. B. The ‘glutamate switch’ provides a link between ATPase activity and ligand binding in AAA+ proteins. Nat. Struct. Mol. Biol. 15, 1223–1227 (2008).
He, J. et al. The structure of the 26S proteasome subunit Rpn2 reveals its PC repeat domain as a closed toroid of two concentric α-helical rings. Structure 20, 513–521 (2012).
Snoberger, A., Brettrager, E. J. & Smith, D. M. Conformational switching in the coiled-coil domains of a proteasomal ATPase regulates substrate processing. Nat. Commun. 9, 2374 (2018).
Glynn, S. E., Martin, A., Nager, A. R., Baker, T. A. & Sauer, R. T. Structures of asymmetric ClpX hexamers reveal nucleotide-dependent motions in a AAA+ protein-unfolding machine. Cell 139, 744–756 (2009).
Kim, Y. C., Snoberger, A., Schupp, J. & Smith, D. M. ATP binding to neighbouring subunits and intersubunit allosteric coupling underlie proteasomal ATPase function. Nat. Commun. 6, 8520 (2015).
Horwitz, A. A. et al. ATP-induced structural transitions in PAN, the proteasome-regulatory ATPase complex in Archaea. J. Biol. Chem. 282, 22921–22929 (2007).
Puchades, C. et al. Structure of the mitochondrial inner membrane AAA+ protease YME1 gives insight into substrate processing. Science 358, eaao0464 (2017).
Ripstein, Z. A., Huang, R., Augustyniak, R., Kay, L. E. & Rubinstein, J. L. Structure of a AAA+ unfoldase in the process of unfolding substrate. eLife 6, e25754 (2017).
Gates, S. N. et al. Ratchet-like polypeptide translocation mechanism of the AAA+ disaggregase Hsp104. Science 357, 273–279 (2017).
Han, H., Monroe, N., Sundquist, W. I., Shen, P. S. & Hill, C. P. The AAA ATPase Vps4 binds ESCRT-III substrates through a repeating array of dipeptide-binding pockets. eLife 6, e31324 (2017).
Deville, C. et al. Structural pathway of regulated substrate transfer and threading through an Hsp100 disaggregase. Sci. Adv. 3, e1701726 (2017).
Monroe, N., Han, H., Shen, P. S., Sundquist, W. I. & Hill, C. P. Structural basis of protein translocation by the Vps4-Vta1 AAA ATPase. eLife 6, e24487 (2017).
Enemark, E. J. & Joshua-Tor, L. Mechanism of DNA translocation in a replicative hexameric helicase. Nature 442, 270–275 (2006).
Thomsen, N. D. & Berger, J. M. Running in reverse: the structural basis for translocation polarity in hexameric helicases. Cell 139, 523–534 (2009).
Wang, X. et al. Mass spectrometric characterization of the affinity-purified human 26S proteasome complex. Biochemistry 46, 3553–3565 (2007).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zhu, Y., Ouyang, Q. & Mao, Y. A deep convolutional neural network approach to single-particle recognition in cryo-electron microscopy. BMC Bioinformatics 18, 348 (2017).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Wu, J. et al. Massively parallel unsupervised single-particle cryo-EM data clustering via statistical manifold learning. PLoS ONE 12, e0182130 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
The PyMOL Molecular Graphics System Version 1.8 (Schrödinger, LLC).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Acknowledgements
We thank Y. Saeki for the plasmids expressing Sic1PY and WW-HECT; H. Huang for proteasome-expressing cell lines; D. Yu, J. Xu, Y. Ma, C. Fan and J. Jackson for technical support; and S. Elsasser for critical reading of the manuscript. This work was funded in part by an Intel Corporation academic grant, the Thousand Talents Plan of China, National Natural Science Foundation of China grant nos. 11774012 and 91530321, the Peking-Tsinghua Center for Life Sciences (Y.M.), NIH grant GM43601 (D.F.) and an Edward Mallinckrodt, Jr. Foundation award (Y.L.). The cryo-EM data were collected from the Electron Microscopy Laboratory and Cryo-EM Platform at Peking University. Initial cryo-EM screening was performed in part at the Center for Nanoscale Systems at Harvard University supported by the National Science Foundation under NSF award no. 1541959 and NIH grant AI100645. Data processing was performed in part in the Sullivan cluster, which is funded in part by a gift from Mr. and Mrs. Daniel J. Sullivan, Jr., and in the Weiming No.1 and Life Science No. 1 High-Performance Computing Platform at Peking University.
Reviewer information
Nature thanks C. Hill and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.D. purified proteins, conducted biochemical analysis and prepared samples for imaging. Y.D., S.Z., Z.W., X.L., W.L.W., Y.Z. and S.S.-M. collected data. S.Z. and Z.W. processed data and refined the maps. Y.D., S.Z., Y.L. and D.F. contributed to structural analysis and manuscript preparation. Y.M. conceived and supervised this study, devised the methods, performed atomic modelling, analysed the structures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization and structure determination of the substrate-engaged human proteasome.
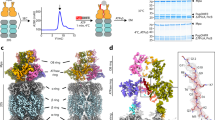
a, FPLC purification of the human 26S proteasome on Superose 6 10/300 GL. The dashed box shows the fraction taken for structural analysis. b, Native PAGE analysis of proteasome purified as in a. c, SDS–PAGE analysis of proteasome purified as in a. d, SDS–PAGE analysis of Sic1PY purified through a Superdex 75 column. e, SDS–PAGE analysis of WW-HECT purified through Superdex 75. b–e, Stained with Coomassie blue. f, SDS–PAGE/western blot analysis of polyubiquitinated Sic1PY. After ubiquitination, the samples were applied to an SDS–polyacrylamide gel, followed by western blotting with anti-T7 antibody. The result suggests that almost all Sic1PY is ubiquitinated. g, Native PAGE analysis of proteasomes crosslinked to PUb-Sic1PY. Left, no-substrate control. With the addition of the PUb-Sic1PY, the proteasome ran slower than without PUb-Sic1PY, indicating that the substrate had been captured by the proteasome but not totally degraded when samples were prepared for cryo-EM analysis. h, SDS–PAGE/western blot analysis of the degradation of PUb-Sic1PY, which was visualized with anti-T7 antibody. This experiment confirms that PUb-Sic1PY is readily degraded by our proteasome samples. i, Typical cryo-EM micrograph of the substrate-engaged human proteasome after motion correction. All experiments in a–i were repeated independently at least three times with similar results. j, Power spectrum evaluation of the micrograph shown in i. k, Gallery of unsupervised class averages calculated by ROME54 using machine-learning-based clustering. l, Local resolution estimation calculated by ResMap55 on seven maps refined by focusing the mask on the RP component. m, Local resolution estimation on seven maps refined by focusing the mask on the CP and ATPase components. n, Gold-standard FSC plots of eight maps calculated without masking the separately refined half-maps. o, Gold-standard FSC plots of the maps refined by focusing on the CP and ATPase components, calculated with masking of the raw half-maps. p, Gold-standard FSC plots of the maps refined by focusing on the RP subcomplex, calculated with masking of the raw half-maps. q, Model-map FSC plots calculated by Phenix57 between each refined map and its corresponding atomic model. For each state, the maps refined by differential masking were merged in Fourier space into a single map, which was used for the model–map FSC calculation. The same colour code is used in n–q.
Extended Data Fig. 2 Focused classification to separate the seven conformational states.
The diagram illustrates the four major steps of our hierarchical focused classification strategy. Further detailed iterations of classification in each step are omitted for clarity.
Extended Data Fig. 3 Cryo-EM maps and quality assessment.
a, The five refined cryo-EM maps that are not shown in the main figures. b, Typical central cross-sections of the density maps for each of the four subcomplexes (the α-ring, the β-ring and the ATPase ring in state EA, and the lid in state ED1). c, Typical nucleotide-binding pocket densities from the EA1 map. The ATP density is compared with the ADP density in two orthogonal perspectives. The magnesium density next to the nucleotide is labelled. d, Typical densities of secondary structures and substrate in the proteasome superimposed with their atomic models.
Extended Data Fig. 4 Key structural features that differentiate the seven conformational states.
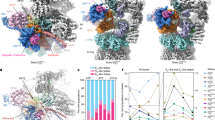
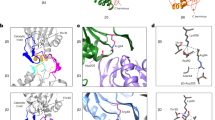
a, Ubiquitin densities in states EA1 (left) and EA2 (right). The T1 site is labelled by fitting the yellow cartoon representation of the NMR structure (RCSB Protein Data Bank (PDB) ID 2N3U) of the yeast Rpn1 T1 element in complex with two ubiquitins into our density, showing that the ubiquitin on human RPN1 is bound to a site very close to the yeast Rpn1 T1 site6. The density maps are low-pass-filtered to 8 Å to show the ubiquitin features clearly, owing to the lower local resolution of the ubiquitin density in these maps. b, The ubiquitin–RPN11–RPT5 interface observed at high resolution in state EB is also observed in state EA2, albeit at lower local resolution. The EA2 density is shown as a transparent surface. c, Comparison of the Ins1 loop of RPN11 in different states. d, Comparison of the RPN11 structures in states EA2, EB and EC1 around the zinc-binding site and Ins1 region with that in the crystal structure (PDB ID: 5U4P) of a ubiquitin-bound Rpn11–Rpn8 complex from yeast29. e, Close-up comparison of the RPN11 Ins1 structure between state EB and 5U4P (left two panels) and between state EC1 and 5U4P (right two panels) in two orthogonal perspectives, showing a 5 Å displacement of the Ins1 β-hairpin in EB relative to 5U4P or EC1. This displacement is not observed between EC1 and 5U4P, suggesting that the Ins1 β-hairpin tilt in EB is mostly to optimize the coordination of the isopeptide bond with the zinc ion. f, Comparison of the RP structures of EA and EB. g, Comparison of lid subcomplex conformations among all states. h, Comparison of ATPase ring structures between two successive states. The structures are aligned together against their CP in f–h. i, Side views of the structural comparison of the AAA ring between EC1 (colour) and EB (grey). The large AAA subdomain of RPT1 was used to align the two AAA-ring structures together. A 40° out-of-plane rotation of the large AAA subdomain of RPT1 relative to the AAA ring is observed during disengagement of RPT1–RPT2 from the substrate. The right panel, rotated vertically against the left panel, shows that the out-of-plane rotation in RPT1 is more substantially amplified in its anticlockwise neighbouring ATPases than in its clockwise neighbours. Red arrows mark the centre of the AAA ring. j, Structural comparison between ED1 (colour) and EC2 (grey) in which the large AAA subdomain of RPT1 is used to align the two AAA-ring structures. A small 5° out-of-plane rotation of the large AAA subdomain of RPT1 relative to the AAA ring is observed during the re-engagement of RPT1–RPT2 with the substrate. k, Structural comparison between ED1 (colour) and EC2 (grey), by using the large AAA subdomain of RPT5 to align the two AAA-ring structures. A 30° out-of-plane rotation of the large AAA subdomain of RPT5 relative to the AAA ring is observed during disengagement of RPT5 from the substrate. The right panel, rotated vertically against the left panel, shows that the out-of-plane rotation in RPT5 is amplified in its anticlockwise neighbouring ATPases more substantially than in its clockwise neighbours. Red arrows mark the centre of the AAA ring. l, Structural comparison between ED2 (colour) and ED1 (grey) in which the large AAA subdomain of RPT5 is used to align the two AAA-ring structures. An 8° out-of-plane rotation of the large AAA subdomain of RPT5 relative to the AAA ring is observed during re-engagement of RPT5 with the substrate.
Extended Data Fig. 5 The RP–CP interface in different states.
a, Comparison of the RP–CP interface and RPT C-terminal tail insertions into the α-pockets of the CP in different states. The cryo-EM densities of the RP–CP interfaces are shown as a grey surface representation. The red dashed circles highlight the densities of the RPT C-terminal tails. b, Close-up views and comparison of the RPT C-tail densities superimposed with the atomic models in different states. The cryo-EM densities of the RPT C-tails are shown in blue mesh representation. The atomic models of the RPT C-tails are shown in stick representation. The CP structures are shown as grey cartoon representations.
Extended Data Fig. 6 Substrate densities in different states.
a, Close-up views of two typical substrate densities observed in the CP chamber in state EA. Left, the substrate density directly contacting the proteolytically active Thr1 in subunit β2. Right, a long substrate density at the seam between two β4 subunits inside the CP. b, The overall ATPase ring density of state EB (left) and a close-up view of the substrate density (right). c, The overall ATPase ring density of states EC1,2 (left) and a close-up view of the substrate density (right). d, The overall ATPase ring density of states ED1,2 (left) and a close-up view of the substrate density (right). All close-up views were directly screen-copied from Coot56 after atomic modelling into the density maps without modification.
Extended Data Fig. 7 Nucleotide densities in all states.
The nucleotide densities fitting with atomic models are shown in blue mesh. All close-up views were directly screen-copied from Coot56 after atomic modelling into the density maps without modification. At the contour level commonly used for atomic modelling, the potential nucleotide densities in the apo-like subunits mostly disappear, although they can appear as partial nucleotide shapes at a much lower contour level.
Extended Data Fig. 8 Geometries of nucleotide-binding pockets and nucleotide-driven intrasubunit conformational changes of AAA domains.
a, Comparison of the nucleotide-binding pockets of six ATPases in all states illustrates a common pattern in the geometry of the nucleotide-binding sites. Each row shows the geometry of the nucleotide-binding pocket of one ATPase in all six states. In each panel showing an ATP or ADP-bound state, one red dashed line marks the distance from the β- or γ-phosphate of the nucleotide to the arginine finger of the adjacent ATPase, and the other line marks the distance from the same phosphate to the Walker B motif. In the case of apo-like states, the red lines extend to the proline of the Walker A motif rather than to the phosphate groups. These geometries indicate the potential reactivity of these sites33. When the ATPase is positioned in the middle of the pore-loop staircase, but not at the lowest position, the nucleotide-binding pockets are tightly packed regardless of whether ATP or ADP is bound. By contrast, when the ATPase is either in the lowest position of the substrate-pore loop staircase or disengaged from the substrate, the nucleotide-binding pocket is rather open regardless of whether it is ADP-bound or free of nucleotide. b–e, Superpositions of the AAA domain structures of RPT1 (b), RPT2 (c), RPT3 (d) or RPT4 (e) from six distinct states aligned against their large AAA subdomains. RPT1 assumes two major conformations and RPT2 assumes three.
Extended Data Fig. 9 Changes in lid–base interactions are associated with ATP hydrolysis events through long-range allosteric regulation.
a, b, Long-range association between RPN1 and RPN2 through a looping structure from RPN2 (residues 820–871) observed in states ED1 (a) and ED2 (b). c, Comparison of the RPN1–RPN2 long-range association between these two states shows a marked, 12 Å movement of RPN1 relative to the CP. In both states (ED1 and ED2), the RPN1 toroidal domain and the CC domain of the RPT1–RPT2 dimer together form a surface cavity into which a short helix from RPN2 is inserted34. This helix resides in the middle of a long loop (residues 820–871) emanating from the toroidal domain of RPN2. The long-range association of RPN1 and RPN2 seems to stabilize a larger interface formed between RPN1–RPN2 and RPT1–RPT2. However, such a quaternary architecture is not observed in other states (EA–C). In states EC1,2, the RPN1 density is considerably blurred, reflecting strong motions that potentially break the long-range RPN1–RPN2 association (Fig. 1b, Extended Data Fig. 3a). Thus, the specific RPN1 conformation in each state appears to be highly coordinated with the hydrolytic cycle of the ATPase ring, and is controlled by RPN1’s interactions with RPN2 in a long-range fashion. d, Comparison of the interactions of the CC domain of RPT4–RPT5 with RPN9 and RPN10 in states EC1,2 and ED1,2. e, Close-up views of the CC domain of RPT4–RPT5 in contact with RPN9 in states EC1,2 and ED1, and of this CC domain’s contact switching to RPN10 in state ED2. These observations are consistent with a recent study35.
Extended Data Fig. 10 Expanded model of the complete cycle of substrate processing by the human 26S proteasome.
The cartoon summarizes the concept of three principal modes of coordinated ATP hydrolysis observed in the seven states and our proposal of how they regulate the complete cycle of substrate processing by the proteasome holoenzyme. Coordinated ATP hydrolysis in modes 1, 2 and 3 features hydrolytic events in two oppositely positioned ATPases11,36, in two consecutive ATPases9,37,38, and in only one ATPase at a time39,40,43,44,45,46, respectively. Substrate processing undergoes three major steps before CP gate opening for processive translocation: (1) ubiquitin recognition; (2) simultaneous deubiquitylation and substrate engagement with the AAA-ATPase ring; and (3) translocation initiation, which involves multiple simultaneous events, including ubiquitin release, ATPase repositioning and switching of the RPT C-tail insertion pattern. In some cases, the initiation of translocation may precede deubiquitylation. In steps 1 and 2, the ATPases follow mode-1 ATP hydrolysis. In step 3, they follow mode-2 ATP hydrolysis. After the gate is open, the AAA-ATPases hydrolyse ATP in mode 3, in which only one nucleotide is hydrolysed at a time.
Supplementary information
Supplementary Information
This file contains Supplementary Discussions regarding “Power stroke”, “Long-range quaternary allosteric regulation”, and “Functional asymmetry” along with Supplementary References.
Video 1: Substrate processing by the human 26S proteasome.
The complete dynamic process of the substrate engagement, deubiquitylation and translocation within the human 26S holoenzyme. The video is directly interpolated from the seven atomic structures of the substrate-bound human proteasome. The video shows smooth motions of the entire 26S holoenzyme, intuitively suggesting that the seven conformational states are on the pathway of substrate processing by the proteasome
Video 2: Substrate interactions with the axial channel of the proteasomal ATPase ring.
The closeup view of the dynamic interactions between the substrate and the ATPase ring during substrate processing, showing how the differential rotations are driven by ATP hydrolysis and how such conformational changes mechanically translate the substrate toward the core particle (below the ATPase but not shown for clarity). The video is directly interpolated from the seven atomic structures of the substrate-bound human proteasome
Rights and permissions
About this article
Cite this article
Dong, Y., Zhang, S., Wu, Z. et al. Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature 565, 49–55 (2019). https://doi.org/10.1038/s41586-018-0736-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0736-4
This article is cited by
-
Improving resolution and resolvability of single-particle cryoEM structures using Gaussian mixture models
Nature Methods (2024)
-
Characterizing ATP processing by the AAA+ protein p97 at the atomic level
Nature Chemistry (2024)
-
Visualizing chaperone-mediated multistep assembly of the human 20S proteasome
Nature Structural & Molecular Biology (2024)
-
Molecular basis of SAP05-mediated ubiquitin-independent proteasomal degradation of transcription factors
Nature Communications (2024)
-
Protein degradation: expanding the toolbox to restrain cancer drug resistance
Journal of Hematology & Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.