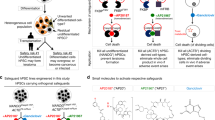
Abstract
Human pluripotent cell lines hold enormous promise for the development of cell-based therapies. Safety, however, is a crucial prerequisite condition for clinical applications. Numerous groups have attempted to eliminate potentially harmful cells through the use of suicide genes1, but none has quantitatively defined the safety level of transplant therapies. Here, using genome-engineering strategies, we demonstrate the protection of a suicide system from inactivation in dividing cells. We created a transcriptional link between the suicide gene herpes simplex virus thymidine kinase (HSV-TK) and a cell-division gene (CDK1); this combination is designated the safe-cell system. Furthermore, we used a mathematical model to quantify the safety level of the cell therapy as a function of the number of cells that is needed for the therapy and the type of genome editing that is performed. Even with the highly conservative estimates described here, we anticipate that our solution will rapidly accelerate the entry of cell-based medicine into the clinic.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Li, W. & Xiang, A. P. Safeguarding clinical translation of pluripotent stem cells with suicide genes. Organogenesis 9, 34–39 (2013).
Dobie, K., Mehtali, M., McClenaghan, M. & Lathe, R. Variegated gene expression in mice. Trends Genet. 13, 127–130 (1997).
Yagyu, S., Hoyos, V., Del Bufalo, F. & Brenner, M. K. Multiple mechanisms determine the sensitivity of human-induced pluripotent stem cells to the inducible caspase-9 safety switch. Mol. Ther. Methods Clin. Dev. 3, 16003–16008 (2016).
Kotini, A. G., de Stanchina, E., Themeli, M., Sadelain, M. & Papapetrou, E. P. Escape mutations, ganciclovir resistance, and teratoma formation in human iPSCs expressing an HSVtk suicide gene. Mol. Ther. Nucleic Acids 5, e284 (2016).
Preuß, E. et al. TK.007: A novel, codon-optimized HSVtk(A168H) mutant for suicide gene therapy. Hum. Gene Ther. 21, 929–941 (2010).
Diril, M. K. et al. Cyclin-dependent kinase 1 (Cdk1) is essential for cell division and suppression of DNA re-replication but not for liver regeneration. Proc. Natl Acad. Sci. USA 109, 3826–3831 (2012).
Santamaría, D. et al. Cdk1 is sufficient to drive the mammalian cell cycle. Nature 448, 811–815 (2007).
Satyanarayana, A. et al. Genetic substitution of Cdk1 by Cdk2 leads to embryonic lethality and loss of meiotic function of Cdk2. Development 135, 3389–3400 (2008).
Fillat, C., Carrió, M., Cascante, A. & Sangro, B. Suicide gene therapy mediated by the herpes simplex virus thymidine kinase gene/ganciclovir system: fifteen years of application. Curr. Gene Ther. 3, 13–26 (2003).
Tomicic, M. T., Thust, R. & Kaina, B. Ganciclovir-induced apoptosis in HSV-1 thymidine kinase expressing cells: critical role of DNA breaks, Bcl-2 decline and caspase-9 activation. Oncogene 21, 2141–2153 (2002).
Berger, C., Flowers, M. E., Warren, E. H. & Riddell, S. R. Analysis of transgene-specific immune responses that limit the in vivo persistence of adoptively transferred HSV-TK-modified donor T cells after allogeneic hematopoietic cell transplantation. Blood 107, 2294–2302 (2006).
Gertsenstein, M. et al. Efficient generation of germ line transmitting chimeras from C57BL/6N ES cells by aggregation with outbred host embryos. PLoS ONE 5, e11260 (2010).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
The International Stem Cell Initiative. Characterization of human embryonic stem cell lines by the International Stem Cell Initiative. Nat. Biotechnol. 25, 803–816 (2007).
Damjanov, I. & Andrews, P. W. Teratomas produced from human pluripotent stem cells xenografted into immunodeficient mice - a histopathology atlas. Int. J. Dev. Biol. 60, 337–419 (2016).
Wu, S. M. & Hochedlinger, K. Harnessing the potential of induced pluripotent stem cells for regenerative medicine. Nat. Cell Biol. 13, 497–505 (2011).
Bianconi, E. et al. An estimation of the number of cells in the human body. Ann. Hum. Biol. 40, 463–471 (2013).
Mesnil, M. & Yamasaki, H. Bystander effect in herpes simplex virus-thymidine kinase/ganciclovir cancer gene therapy: role of gap-junctional intercellular communication. Cancer Res. 60, 3989–3999 (2000).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Cervantes, R. B., Stringer, J. R., Shao, C., Tischfield, J. A. & Stambrook, P. J. Embryonic stem cells and somatic cells differ in mutation frequency and type. Proc. Natl Acad. Sci. USA 99, 3586–3590 (2002).
Larson, J. S., Yin, M., Fischer, J. M., Stringer, S. L. & Stringer, J. R. Expression and loss of alleles in cultured mouse embryonic fibroblasts and stem cells carrying allelic fluorescent protein genes. BMC Mol. Biol. 7, 36 (2006).
Mortensen, R. M., Conner, D. A., Chao, S., Geisterfer-Lowrance, A. A. & Seidman, J. G. Production of homozygous mutant ES cells with a single targeting construct. Mol. Cell. Biol. 12, 2391–2395 (1992).
Koike, H. et al. Efficient biallelic mutagenesis with Cre/loxP-mediated inter-chromosomal recombination. EMBO Rep. 3, 433–437 (2002).
Yusa, K. et al. Genome-wide phenotype analysis in ES cells by regulated disruption of Bloom’s syndrome gene. Nature 429, 896–899 (2004).
Schwartz, S. D. et al. Human embryonic stem cell-derived retinal pigment epithelium in patients with age-related macular degeneration and Stargardt’s macular dystrophy: follow-up of two open-label phase 1/2 studies. Lancet 385, 509–516 (2015).
Mandai, M. et al. Autologous induced stem-cell-derived retinal cells for macular degeneration. N. Engl. J. Med. 376, 1038–1046 (2017).
Chong, J. J. H. et al. Human embryonic-stem-cell-derived cardiomyocytes regenerate non-human primate hearts. Nature 510, 273–277 (2014).
Barry, J., Hyllner, J., Stacey, G., Taylor, C. J. & Turner, M. Setting up a haplobank: issues and solutions. Curr. Stem Cell Rep. 1, 110–117 (2015).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Behringer, R., Gertsenstein, M., Nagy, K. V. & Nagy, A. Manipulating the Mouse Embryo: A Laboratory Manual 4th edn (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2013).
Mereau, A., Grey, L., Piquet-Pellorce, C. & Heath, J. K. Characterization of a binding protein for leukemia inhibitory factor localized in extracellular matrix. J. Cell Biol. 122, 713–719 (1993).
Park, J., Kusminski, C. M., Chua, S. C. & Scherer, P. E. Leptin receptor signaling supports cancer cell metabolism through suppression of mitochondrial respiration in vivo. Am. J. Pathol. 177, 3133–3144 (2010).
Klimanskaya, I. et al. Derivation and comparative assessment of retinal pigment epithelium from human embryonic stem cells using transcriptomics. Cloning Stem Cells 6, 217–245 (2004).
Sun, X. et al. Directed differentiation of human embryonic stem cells into thymic epithelial progenitor-like cells reconstitutes the thymic microenvironment in vivo. Cell Stem Cell 13, 230–236 (2013).
Luria, S. E. & Delbrück, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943).
Capizzi, R. L. & Jameson, J. W. A table for the estimation of the spontaneous mutation rate of cells in culture. Mutat. Res. 17, 147–148 (1973).
Luo, G. et al. Cancer predisposition caused by elevated mitotic recombination in Bloom mice. Nat. Genet. 26, 424–429 (2000).
Monnat, R. J. Jr. Molecular analysis of spontaneous hypoxanthine phosphoribosyltransferase mutations in thioguanine-resistant HL-60 human leukemia cells. Cancer Res. 49, 81–87 (1989).
Green, M. H. L., O’Neill, J. P. & Cole, J. Suggestions concerning the relationship between mutant frequency and mutation rate at the hprt locus in human peripheral T-lymphocytes. Mutat. Res. 334, 323–339 (1995).
Biancotti, J.-C. et al. The in vitro survival of human monosomies and trisomies as embryonic stem cells. Stem Cell Res. 9, 218–224 (2012).
Acknowledgements
We thank the TCP Transgenic Core and M. Gertsenstein for mouse line derivation; A. Bang for flow cytometry; the TCP Pathology Core and K. Harpal for histology analysis; M. Kownacka for providing MEFs; M. S. Shoichet for HAMC; N. Mitrousis for qPCR primers; S. Nurk for advice on Monte Carlo simulation; I. P. Michael, P. D. Tonge, B. V. Varga, C. He and R. El-Rass for experimental advice and J. S. Harding and K. C. Davidson for their proofreading of the manuscript, to J. S. Harding for the artwork in Fig. 4. This work was supported by CIHR Foundation Grant, Foundation Fighting Blindness, Canadian Research Chair and Medicine by Design (University of Toronto) funding to A.N.
Reviewer information
Nature thanks A. Hewitt, T. J. Kieffer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Q.L. designed and conducted most of the experiments, including targeting of mouse and human ES cells, teratoma analysis, mammary gland tumour analysis, neural in vitro differentiation, mutation rate calculation, as well as analysing the data and writing the manuscript. C.M. designed and conducted experiments, designed and constructed the CDK1–TK targeting cassette, targeted mouse and human ES cells, analysed safe-cell escapees, performed in vitro differentiation assays, analysed data, composed figures and wrote the manuscript. M.V.S. conducted the Monte Carlo simulations and analysed the data. I.G. performed the mathematical modelling. E.J.N. targeted human ES cells, differentiated RPEs and wrote the manuscript. S.H. performed the eye experiment and analysed the data. H.Y. performed in vitro differentiation into endoderm and analysed the data. C.K. performed in vitro differentiation into mesenchymal stem cells and analysed the data. P.Z. performed Southern blots. C.L. performed animal teratoma experiments. K.N. performed animal teratoma experiments, analysed the data and edited the manuscript. M.M. targeted human ES cells. H.-K.S. analysed teratoma histology. A.N. conceived and supervised the study, designed experiments, analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.N., C.M. and Q.L. are inventors on a patent application covering the SC technology (PCT/CA2016/050256). A.N. is a co-founder and shareholder of panCELLa Inc. C.M. is a senior scientist at panCELLa Inc. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generation, genotyping and characterization of mouse C57BL/6N C2 Cdk1–TK/Cdk1 and Cdk1–TK/Cdk1–TK ES cells.
a, Summary of the targeting steps used to generate mouse C2 Cdk1–TK/Cdk1 and Cdk1–TK/Cdk1–TK ES cells. b, Southern blot genotyping with internal TK–mCherry probe. c, Southern blot genotyping with mouse Cdk1 genomic probe. d, In vitro GCV dose–response killing curve of mouse C2 CDK1–TK/CDK1 ES cells. Data are mean ± s.d., n = 3. e, Teratoma formation efficiency of mouse C2 Cdk1–TK-PB/Cdk1, Cdk1–TK/Cdk1 and Cdk1–TK/Cdk1–TK ES cells.
Extended Data Fig. 2 Generation, genotyping and characterization of human H1 CDK1–TK/CDK1 and CDK1–TK/CDK1–TK ES cells.
a, Generation of human H1 CDK1–TK/CDK1 and CDK1–TK/CDK1–TK ES cells. b, Southern blot genotyping of CDK1–TK/CDK1 clone Exc16, which was used in teratoma assays (Fig. 1c and Extended Data Fig. 7) and CDK1–TK/CDK1–TK clone Exc16-3C, which was used in the differentiation assays in Fig. 3. c, PCR genotyping of all the correct clones. d, Flow cytometry analysis of the CDK1–TK/CDK1 clone Exc16 and the CDK1–TK/CDK1–TK clone Exc16-3C. e, SybrGreen qPCR of human CDK1 expression in H1 wild-type cells, and cells expressing the CDK1–TK/CDK1 clone Exc16 and the CDK1–TK/CDK1–TK clone Exc16-3C. Data are mean ± s.d., n = 3. No significant difference was found between groups. f, TaqMan qPCR copy-number analysis of TKs of all clones with the correct genotype. g, The efficiency of teratoma formation in NSG mice using human H1 ES cells. h, Dose–response analysis of wild-type, CDK1–TK/CDK1 and CDK1–TK/CDK1–TK human H1 ES cells. Cells were treated with different GCV concentrations, dissociated and counted after seven days. Data are mean ± s.d., n = 3. Source data
Extended Data Fig. 3 Generation, genotyping and characterization of human CA1 CDK1–TK/CDK1 and CDK1–TK/CDK1–TK ES cells.
a, Generation of human CA1 CDK1–TK/CDK1 and CDK1–TK/CDK1–TK ES cells. b, Southern blot genotyping of human CA1 CDK1–TK/CDK1 and CDK1–TK/CDK1–TK ES cells. The plasmid concatemers are multiple copies of plasmid integration (including backbone). The ampicillin gene in the backbone contains a ScaI restriction enzyme site, which is consistent with the sizes of the band in Southern blots. c, Haematoxylin and eosin staining of a CDK1–TK/CDK1–TK CA1 ES-cell-derived teratoma. d, The efficiency of teratoma formation in NSG mice using human CA1 ES cells. e, Flow cytometry analysis shows a direct correlation between the number of CDK1–TK alleles and mCherry fluorescence levels.
Extended Data Fig. 4 Growth graphs of mouse and human ES-cell-derived teratomas.
a, Growth of teratomas derived from mouse heterozygous safe-cell ES cells (C2 Cdk1–TK/Cdk1) b, Adult mouse with stabilized subcutaneous tissue (safe-cell ES-cell-derived dormant teratoma), 2.5 months after GCV treatment. c, Growth of teratomas derived from mouse homozygous safe-cell ES cells (C2 Cdk1–TK/Cdk1–TK). d, Growth of teratomas derived from human heterozygous safe-cell ES cells (H1 CDK1–TK/CDK1, clone Exc16); daily GCV treatment. e, Examples of teratomas from human heterozygous safe-cell ES cells showing cyst formation, images of cystic teratomas at dissection are shown next to the corresponding growth line; daily GCV treatment. The graphs with two lines represent mice that had cells injected into both flanks. The graphs with one line represent mice that had cells injected into one flank. The GCV treatment regime varies among mice because each teratoma behaves differently; we started GCV when the teratoma size started to increase. f, Growth of teratomas derived from human homozygous safe-cell ES cells (H1 CDK1–TK/CDK1–TK), GCV treatment was every other day. Images of cystic teratomas are shown next to the corresponding growth line, cysts were drained after dissection to show the difference in tumour weight due to the fluid present in the tissue. Each graph represents one mouse. a, c, d, f, All replicates of these experiments are shown. Source data
Extended Data Fig. 5 Breast cancer transplantation assay using heterozygous and homozygous safe-cell mammary tumour cells.
a, Generation of mouse lines and experimental design. b, Growth of mammary gland tumours derived from mouse Cdk1/Cdk1 and Cdk1–TK/Cdk1 mammary epithelial cells with PBS or GCV treatment. c, Growth of mammary gland tumours derived from mouse CDK1–TK/CDK1–TK mammary epithelial cells with PBS or GCV treatment. b, c, The sample sizes of each group are indicated at the bottom of the graphs.
Extended Data Fig. 6 In vitro experiments with mouse C2 Cdk1–TK/Cdk1 and Cdk1–TK/Cdk1–TK ES cells and subsequent characterization of escapees.
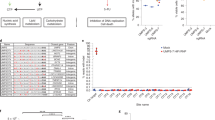
a, Experimental design: mCherry+ cells were selected by sorting to ensure that the starting cell population did not contain escapees. These cells were plated on six-well plates (200 cells per well, in a total of 36 wells) and allowed to grow to 14 cell doublings (this was estimated by counting cells in sample wells). The 36 cultures were then resuspended to a single-cell suspension and each was plated in a 15-cm plate (4 × 106 cells). One day after plating, selection with GCV was started and maintained until escapee colonies appeared. b, Escapee numbers obtained in 36 independent cultures growing from Cdk1–TK/Cdk1 and Cdk1–TK/Cdk1–TK ES cells. c, PCR to determine the presence of TK. d, TaqMan copy number qPCR analysis of Akap7, Sim1 and Cdk1 junction of exon 8 and 3′ UTR, Neurod, Cdk1, TK transgene and Abca on mouse chromosome 10. Data are the copy number calculated by CopyCaller Software v.2.1 and the error bars indicate the range from the minimum to the maximum number. n = 3. The same colour in the background of c and d indicates that they are from the same independent culture. n.d., not determined. e, qPCR to compare TK expression level in Cdk1–TK/Cdk1 escapee clone 2A and C2 wild-type ES cells. Data are mean ± s.e.m., n = 3. f, Summary of the copy number analysis of mouse Cdk1–TK/Cdk1 escapees. a, b, Experiments were repeated twice on a smaller scale but with similar results. d, Experiments were repeated twice with similar results.
Extended Data Fig. 7 Schematics of the possible mutation types affecting the CDL–SU allele.
a, Three types of mutations that could affect the CDL–SU allele. b, Safe-cell allele transition considered in the modelling and Monte Carlo simulations.
Extended Data Fig. 8 Quality control of batches generated from single human H1 CDK1–TK/CDK1–TK ES cells.
a, The drop of SCL due to aliquoting from a pool of cells relative to non-aliquoted batches of the same size. b, Schematics of the alleles in the CDK1–TK/CDK1–TK human ES cells used in the quality control. c, Workflow schematic of performing quality control (QC) on several ES cell batches. d, An example of the flow cytometry for the quality control of nine clonally derived batches. e, An example of PCR for the quality control of nine clonally derived batches.
Extended Data Fig. 9 Mouse and human safe-cell homozygous CDK1–TK/CDK1–TK cells demonstrate pluripotency.
All experiments were performed using the same clone of mouse C2 or human H1 (Exc16-3C) CDK1–TK/CDK1–TK cells. a, Bright-field photograph showing mouse homozygous Cdk1–TK/Cdk1–TK ES cell morphology. b, Bright-field photograph showing human homozygous CDK1–TK/CDK1–TK ES cell morphology. c, Haematoxylin and eosin staining of a mouse Cdk1–TK/Cdk1–TK ES-cell-derived teratoma. d, Haematoxylin and eosin staining of a human CDK1–TK/CDK1–TK ES-cell-derived teratoma. e, An adult Cdk1–TK/Cdk1–TK mouse. f, OCT4 and NANOG staining of human CDK1–TK/CDK1–TK ES cells. g, qPCR characterization of CDK1–TK/CDK1–TK ES cell differentiation into RPE cells. Data are mean ± s.d., n = 3. h, Bright-field picture of human CDK1–TK/CDK1–TK ES-cell-derived RPE cells. i, ZO1 staining of human CDK1–TK/CDK1–TK ES-cell-derived RPE cells. j, Human CDK1–TK/CDK1–TK ES-cell-derived adipocytes, chondrocytes and osteocytes. k, SOX17 and FOXA2 staining of human CDK1–TK/CDK1–TK ES-cell-derived definitive endoderm. l, OCT4 (also known as POU5F1), NANOG, SOX17, FOXA2 and HOXA3 qPCR characterization of ES cell (day 0) differentiation into definitive endoderm (day 4) and pharyngeal pouch endoderm (day 8). Data are mean ± s.d., n = 3. a–d, f–i, Experiments were repeated three times with similar results. j–l, Experiments were repeated twice with similar results.
Extended Data Fig. 10 Representative images of eyes transplanted with both safe-cell RPE and safe-cell ES cells, and only with safe-cell RPE cells.
a, b, Fundoscopy, optical coherence tomography and fluorescence imaging of eyes transplanted with safe-cell RPE and safe-cell ES cells (four-week GCV treatment). The absence of a mCherry signal indicates that ES cell growth has not occurred. b, Bottom, images of the green fluorescence channel are included to illustrate that the observed signal in the red fluorescence channel is actually autofluorescence. This experiment was repeated four times in four mice with similar results. c, Histological analysis of the eye presented in d. d, Fundoscopy, optical coherence tomography and fluorescence imaging of eyes transplanted with safe-cell RPE and safe-cell ES cells (PBS treatment). This experiment was repeated twice in two mice with similar results. e, f, Fundoscopy, optical coherence tomography and fluorescence imaging of eyes transplanted with safe-cell RPE and safe-cell ES cells, mCherry signal is detectable and indicates ES cell growth. GCV treatment began three weeks post-injection following an initial PBS treatment. This experiment was repeated four times in three mice with similar results. g, Fundoscopy, optical coherence tomography and fluorescence imaging of eyes receiving only safe-cell RPE cells (four-week GCV treatment). This demonstrates that GCV treatment did not affect the RPE cells. This experiment was repeated five times in three mice with similar results. h, Fundoscopy, optical coherence tomography and fluorescence imaging of eyes receiving only safe-cell RPE cells (four-week PBS treatment). This experiment was repeated six times in three mice with similar results. i, Fundoscopy, optical coherence tomography and fluorescence imaging of eyes receiving only HAMC (four-week GCV treatment). This experiment was repeated twice in one mouse with similar results.
Supplementary information
Supplementary Information
This file contains Supplementary Equation 1: Detailed mathematical steps used to calculate the effect of aliquoting on SCL as illustrated in Extended Data Fig. 8a
Supplementary Table
This file contains Supplementary Table 1. To generate a list of CDLCDEL candidates, we cross-referenced genes whose knock-out has an early embryonic lethal phenotype in mice (www.informatics.jax.org/phenotypes.shtml) with data from a genome-wide CRISPR/Cas9 mutagenesis screen for essential genes in human cancer cell lines41. We found 167 genes with a high-fitness score (< -1.0) that also have a known early-embryonic lethal phenotype
Supplementary Table
This file contains Supplementary Table 2: List of DNA primers that were used in this study
Supplementary Table
This file contains Supplementary Table 3: List of antibodies that were used in this study
Supplementary Table
This file contains Supplementary Table 4: Calculation of the LOH rate at the human CDK1 locus with human H1 CDK1-TK/CDK1-TK ES cells. a, Summary table. b, Detailed cell number for each individual clone
Video 1
: Bright field video of beating cardiomyocytes differentiated from CDK1-TK/CDK1-TK human H1 ES cells
Rights and permissions
About this article
Cite this article
Liang, Q., Monetti, C., Shutova, M.V. et al. Linking a cell-division gene and a suicide gene to define and improve cell therapy safety. Nature 563, 701–704 (2018). https://doi.org/10.1038/s41586-018-0733-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0733-7
Keywords
This article is cited by
-
Soluble CX3CL1-expressing retinal pigment epithelium cells protect rod photoreceptors in a mouse model of retinitis pigmentosa
Stem Cell Research & Therapy (2023)
-
Cell-based passive immunization for protection against SARS-CoV-2 infection
Stem Cell Research & Therapy (2023)
-
Regenerating human skeletal muscle forms an emerging niche in vivo to support PAX7 cells
Nature Cell Biology (2023)
-
Culture-expanded mesenchymal stromal cell therapy: does it work in knee osteoarthritis? A pathway to clinical success
Cellular & Molecular Immunology (2023)
-
Immune-privileged tissues formed from immunologically cloaked mouse embryonic stem cells survive long term in allogeneic hosts
Nature Biomedical Engineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.