Abstract
The two-membrane envelope is a defining feature of chloroplasts. Chloroplasts evolved from a Gram-negative cyanobacterial endosymbiont. During evolution, genes of the endosymbiont have been transferred to the host nuclear genome. Most chloroplast proteins are synthesized in the cytosol as higher-molecular-mass preproteins with an N-terminal transit peptide. Preproteins are transported into chloroplasts by the TOC and TIC (translocons at the outer- and inner-envelope membranes of chloroplasts, respectively) machineries1,2, but how TOC and TIC are assembled together is unknown. Here we report the identification of the TIC component TIC236; TIC236 is an integral inner-membrane protein that projects a 230-kDa domain into the intermembrane space, which binds directly to the outer-membrane channel TOC75. The knockout mutation of TIC236 is embryonically lethal. In TIC236-knockdown mutants, a smaller amount of the inner-membrane channel TIC20 was associated with TOC75; the amount of TOC–TIC supercomplexes was also reduced. This resulted in a reduced import rate into the stroma, though outer-membrane protein insertion was unaffected. The size and the essential nature of TIC236 indicate that—unlike in mitochondria, in which the outer- and inner-membrane translocons exist as separate complexes and a supercomplex is only transiently assembled during preprotein translocation3,4—a long and stable protein bridge in the intermembrane space is required for protein translocation into chloroplasts. Furthermore, TIC236 and TOC75 are homologues of bacterial inner-membrane TamB5 and outer-membrane BamA, respectively. Our evolutionary analyses show that, similar to TOC75, TIC236 is preserved only in plants and has co-evolved with TOC75 throughout the plant lineage. This suggests that the backbone of the chloroplast protein-import machinery evolved from the bacterial TamB–BamA protein-secretion system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in the article and its Supplementary Information. Full gel blots can be found in Supplementary Fig. 1. Any other data are available from the corresponding author upon reasonable request.
References
Shi, L. X. & Theg, S. M. The chloroplast protein import system: from algae to trees. Biochim. Biophys. Acta 1833, 314–331 (2013).
Paila, Y. D., Richardson, L. G. L. & Schnell, D. J. New insights into the mechanism of chloroplast protein import and its integration with protein quality control, organelle biogenesis and development. J. Mol. Biol. 427, 1038–1060 (2015).
Geissler, A. et al. The mitochondrial presequence translocase: an essential role of Tim50 in directing preproteins to the import channel. Cell 111, 507–518 (2002).
Yamamoto, H. et al. Tim50 is a subunit of the TIM23 complex that links protein translocation across the outer and inner mitochondrial membranes. Cell 111, 519–528 (2002).
Selkrig, J. et al. Discovery of an archetypal protein transport system in bacterial outer membranes. Nat. Struct. Mol. Biol. 19, 506–510 (2012).
Chu, C. C. & Li, H.-m. Protein import into isolated pea root leucoplasts. Front. Plant Sci. 6, 690 (2015).
Tzafrir, I. et al. Identification of genes required for embryo development in Arabidopsis. Plant Physiol. 135, 1206–1220 (2004).
Matsushima, R. et al. Amyloplast-localized SUBSTANDARD STARCH GRAIN4 protein influences the size of starch grains in rice endosperm. Plant Physiol. 164, 623–636 (2014).
Chen, Y. L., Chen, L. J. & Li, H.-m. Polypeptide transport-associated domains of the Toc75 channel protein are located in the intermembrane space of chloroplasts. Plant Physiol. 172, 235–243 (2016).
Heinz, E. & Lithgow, T. A comprehensive analysis of the Omp85/TpsB protein superfamily structural diversity, taxonomic occurrence, and evolution. Front. Microbiol. 5, 370 (2014).
Iqbal, H., Kenedy, M. R., Lybecker, M. & Akins, D. R. The TamB ortholog of Borrelia burgdorferi interacts with the β-barrel assembly machine (BAM) complex protein BamA. Mol. Microbiol. 102, 757–774 (2016).
Chen, L. J. & Li, H.-m. Stable megadalton TOC–TIC supercomplexes as major mediators of protein import into chloroplasts. Plant J. 92, 178–188 (2017).
Tu, S. L. et al. Import pathways of chloroplast interior proteins and the outer-membrane protein OEP14 converge at Toc75. Plant Cell 16, 2078–2088 (2004).
Li, H.-m. & Chen, L.-J. A novel chloroplastic outer membrane-targeting signal that functions at both termini of passenger polypeptides. J. Biol. Chem. 272, 10968–10974 (1997).
Heinz, E., Selkrig, J., Belousoff, M. J. & Lithgow, T. Evolution of the Translocation and Assembly Module (TAM). Genome Biol. Evol. 7, 1628–1643 (2015).
Josts, I. et al. The structure of a conserved domain of TamB reveals a hydrophobic β taco fold. Structure 25, 1898–1906.e5 (2017).
Paila, Y. D. et al. Multi-functional roles for the polypeptide transport associated domains of Toc75 in chloroplast protein import. eLife 5, e12631 (2016).
O’Neil, P. K. et al. The POTRA domains of Toc75 exhibit chaperone-like function to facilitate import into chloroplasts. Proc. Natl Acad. Sci. USA 114, E4868–E4876 (2017).
Komiya, T. et al. Interaction of mitochondrial targeting signals with acidic receptor domains along the protein import pathway: evidence for the ‘acid chain’ hypothesis. EMBO J. 17, 3886–3898 (1998).
Liu, L., McNeilage, R. T., Shi, L. X. & Theg, S. M. ATP requirement for chloroplast protein import is set by the Km for ATP hydrolysis of stromal Hsp70 in Physcomitrella patens. Plant Cell 26, 1246–1255 (2014).
Huang, P. K., Chan, P. T., Su, P. H., Chen, L. J. & Li, H.-m. Chloroplast Hsp93 directly binds to transit peptides at an early stage of the preprotein import process. Plant Physiol. 170, 857–866 (2016).
Perry, S. E., Li, H.-m. & Keegstra, K. In vitro reconstitution of protein transport into chloroplasts. Methods Cell Biol. 34, 327–344 (1991).
Chu, C. C. & Li, H.-m. Determining the location of an Arabidopsis chloroplast protein using in vitro import followed by fractionation and alkaline extraction. Methods Mol. Biol. 774, 339–350 (2011).
Cline, K., Werner-Washburne, M., Andrews, J. & Keegstra, K. Thermolysin is a suitable protease for probing the surface of intact pea chloroplasts. Plant Physiol. 75, 675–678 (1984).
Lubben, T. H. & Keegstra, K. Efficient in vitro import of a cytosolic heat shock protein into pea chloroplasts. Proc. Natl Acad. Sci. USA 83, 5502–5506 (1986).
Tu, S.-L. & Li, H.-m. Insertion of OEP14 into the outer envelope membrane is mediated by proteinaceous components of chloroplasts. Plant Cell 12, 1951–1960 (2000).
Chou, M. L., Chu, C. C., Chen, L. J., Akita, M. & Li, H.-m. Stimulation of transit-peptide release and ATP hydrolysis by a cochaperone during protein import into chloroplasts. J. Cell Biol. 175, 893–900 (2006).
Teng, Y. S. et al. Tic21 is an essential translocon component for protein translocation across the chloroplast inner envelope membrane. Plant Cell 18, 2247–2257 (2006).
Keegstra, K. & Yousif, A. E. Isolation and characterization of chloroplast envelope membranes. Methods Enzymol. 118, 316–325 (1986).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Ito, T. et al. A new resource of locally transposed Dissociation elements for screening gene-knockout lines in silico on the Arabidopsis genome. Plant Physiol. 129, 1695–1699 (2002).
Kuromori, T. et al. A collection of 11 800 single-copy Ds transposon insertion lines in Arabidopsis. Plant J. 37, 897–905 (2004).
Finn, R. D. et al. InterPro in 2017–beyond protein family and domain annotations. Nucleic Acids Res. 45, D190–D199 (2017).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Ochoa, D. & Pazos, F. Practical aspects of protein co-evolution. Front. Cell Dev. Biol. 2, 14 (2014).
Pazos, F. & Valencia, A. Similarity of phylogenetic trees as indicator of protein–protein interaction. Protein Eng. 14, 609–614 (2001).
Ochoa, D. & Pazos, F. Studying the co-evolution of protein families with the Mirrortree web server. Bioinformatics 26, 1370–1371 (2010).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Acknowledgements
We thank Y.-S. Teng for technical assistance at the initial stage of this project, M. Akita for the Cyanidioschyzon merolae 10D genomic DNA, the proteomics and imaging cores of the Institute of Molecular Biology and the proteomics core of the Institute of Biological Chemistry of Academia Sinica for technical assistance, ABRC for the tic236-1 and tic236-2 mutant seeds and the plant genome project of RIKEN Genomic Sciences Center for tic236-3 mutant seeds. This work was supported by the Ministry of Science and Technology (MOST 107-2321-B-001-001) and Academia Sinica of Taiwan (to H.-m.L.).
Reviewer information
Nature thanks N. Pfanner, D. Schnell and S. Theg for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.-L.C., L.-J.C. and C.-C.C. performed crosslinking and immunoprecipitations; Y.-L.C. performed subplastid fractionation, electron microscopy and in vitro pull down; L.-J.C. and C.-C.C. performed alkaline extractions; L.-J.C. performed imports, mutant isolation and BN-PAGE; L.-J.C. and H.-m.L. performed sucrose density fractionations; P.-K.H. performed evolutionary analyses; J.-R.W. determined the length of TIC236 transit peptide; and H.-m.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of TOC75-interacting proteins.
Leucoplasts were isolated from four-day-old pea roots as described6 and treated with 1 mM DSP. Membrane fractions were solubilized with 1% decylmaltoside and immunoprecipitated with anti-TOC75 antibody. a, Immunoprecipitates were analysed by SDS–PAGE and stained with SYPRO Ruby. Gel shown is a representative of two technical repeats. b, Gel slices as numbered in a were excised and processed for in-gel trypsin digestion and analysed by an LTQ-Orbitrap XL spectrometer. The tandem mass spectrometry (MS/MS) raw data were processed by the extractmsn.exe program (Thermo) and searched against the ‘Pea RNA-Seq gene atlas’ (http://bios.dijon.inra.fr/FATAL/cgi/pscam.cgi) database using the Mascot Daemon 2.4.1 server. The top four protein hits in each band with the correct molecular mass range and more than 15 peptide matches are listed. c, Polypeptide sequence of pea TIC236 preprotein (PsCam044808). Peptides identified in liquid chromatography with MS/MS are marked in red.
Extended Data Fig. 2 Deduced polypeptide sequence of Arabidopsis TIC236 (At2g25660) and prediction of its secondary structure.
The transit peptide is shown in green, the transmembrane domain in purple and the DUF490 domain in orange. The peptide used to generate the mouse antibodies is underlined. Secondary-structure prediction is shown underneath the amino acid sequence and was generated using the JPred4 web server38. H, α-helix; E (blue), β-strand.
Extended Data Fig. 3 TIC236 has a cleavable transit peptide that is approximately 37 amino acids in length.
A C-terminally truncated clone that encodes residues 1 to 227 of EMB2410 was constructed. [35S]EMB2410(1–227) was synthesized as a protein about 23 kDa in size (lane 1) and, when imported into chloroplasts, it was processed into a mature protein slightly smaller than 20 kDa (lanes 3 and 5, blue arrows). To estimate the processing site, a series of N-terminally truncated clones were generated by mutating the initiation methionine to isoleucine, and then individually mutating residues 30, 35, 38 or 46 to methionine (labelled 30M, 35M, 38M and 46M, respectively). In vitro translation initiated from methionine at residue 38 produced a protein of approximately the same size as the mature protein (lane 6). Thus, the transit peptide of EMB2410 is estimated to be 37 residues in length and the calculated molecular mass of EMB2410 after transit peptide removal is 236 kDa (residues 38 to 2166). Data shown are representative of two independent experiments.
Extended Data Fig. 4 TIC236 is a member of the TOC–TIC supercomplexes.
a, b, Pea chloroplasts without (a) or with (b) (imported into chloroplasts under the conditions of 3 mM ATP at room temperature for 2 min) translocating [35S]prRBCS were solubilized by 1% decylmaltoside and analysed side-by-side in 15 to 45% linear sucrose density gradients. Fractions were analysed by SDS–PAGE followed by fluorography for [35S]prRBCS and immunoblotting for major translocon and control proteins. [35S]prRBCS sedimented in three peaks, two of which were near the top or middle of the gradient and most probably correspond to free [35S]prRBCS or [35S]prRBCS that is non-specifically bound to the RuBisCO complex during solubilization. The third peak was located further into the gradient, around fractions 16 to 20. Some TOC75, TOC159 and TIC110 also sedimented in these higher-density fractions, and TIC20 was detected only in fractions 16 to 18. These results suggest that fractions 16 to 18 hosted the TOC–TIC supercomplexes. Three control proteins (RBCL, IEP37 and CAB) were detected only at the top or centre of the gradient (see Fig. 1f and Extended Data Fig. 4c for the sedimentation pattern of CAB). The same results were observed when chloroplasts without translocating [35S]prRBCS were fractionated (a). c, The same experiment as shown in b but using Arabidopsis chloroplasts. d, Pea chloroplasts that contain [35S]prRBCS imported under 3 mM ATP at room temperature for 2 min were solubilized with 1% digitonin and analysed by 2D BN-PAGE followed by immunoblotting for TIC236, TOC75 and TIC20, or fluorography for [35S]prRBCS. The blue dot indicates degraded TIC236. The result from a similar experiment is presented in Fig. 1g. Here we show that the same results were obtained either with or without translocating [35S]prRBCS, and that the positions of the supercomplexes were further supported by the presence of translocating [35S]prRBCS. All data shown are representative of at least two independent experiments.
Extended Data Fig. 5 The tic236-2 and tic236-3 mutants have reduced amounts of TIC236 RNA and TIC236 proteins.
a, Total RNA was isolated from tic236-2 and tic236-3 mutants and their corresponding wild types grown on MS plates for 14 days under 16-h light. Levels of TIC236 RNA expression were analysed by quantitative RT–PCR. Levels of UBQ10 RNA were analysed as normalization controls. Data are from two independent plant batches with three technical repeats for each batch, and are calculated by the Bio-Rad CFX Manager 3.1 software and shown as means ± s.e. The original Cq (quantification cycles) for the six samples are: Col (UBQ10): 20.82, 20.42, 20.38, 19.33, 19.42 and 19.31; Col (TIC236): 31.35, 31.9, 32.22, 30.64, 30.57 and 30.49; tic236-2 (UBQ10): 19.87, 19.86, 20.07, 18.64, 18.62 and 18.75; tic236-2 (TIC236): 34.6, 34.2, 34.8, 33.24, 32.77 and 32.96; No-0 (UBQ10): 19.21, 19.22, 19.27, 18.34, 18.36 and 18.37; No-0 (TIC236): 30.09, 30.41, 30.31, 29.42, 29.74 and 29.34; tic236-3 (UBQ10): 18.13, 17.96, 18.14, 17.63, 17.66 and 17.64; tic236-3 (TIC236): 32.51, 32.86, 32.47, 31.97, 32.62 and 32.1. b, Total proteins were extracted from 20-day-old plants grown on MS plates under 16-h light, and analysed by SDS–PAGE (20 μg of proteins per lane) and immunoblotting with the antibodies indicated at right. SPS, sucrose phosphate synthase; cpHSC70, chloroplast stromal HSC70. Data shown are representative of two independent experiments.
Extended Data Fig. 6 tic236-mutant chloroplasts accumulate more preproteins on the chloroplast surface and have reduced rates of protein import into the stroma.
a, [35S]prRBCS and [35S]prHSP93 were imported into chloroplasts isolated from 14-day-old tic236-3 mutant and the corresponding wild-type (No-0) plants, under 1 mM ATP at room temperature for 10 min. After import, half of the chloroplasts were further treated with thermolysin. Re-isolated intact chloroplasts were analysed by SDS–PAGE and the gels were stained with Coomassie blue. Fluorographs are shown above respective images from the same gel. b, [35S]Met-labelled prTIC40 and prHSP93 were incubated with isolated wild-type (WT) and tic236 (236) mutant chloroplasts under 1 mM ATP at room temperature for various amounts of time, as indicated above the gel. Chloroplasts were re-isolated and analysed by SDS–PAGE and the gels were stained with Coomassie blue. Fluorographs are shown above respective images from the same gel. The precursor, intermediate and mature forms of prTIC40 are indicated by pr, i and m, respectively. Quantification of the amount of imported mature proteins (mature plus intermediate proteins for TIC40), corrected for loading by quantifying the amount of RBCL (for prHSP93) or CAB (for prTIC40) and normalized to the amount imported in the wild-type chloroplasts at 20 min, is shown at right. c, tic236-2-knockdown mutant plants have similar amounts of major translocon components to wild-type plants (Col). Total leaf proteins were extracted from 20-day-old plants and analysed by SDS–PAGE and immunoblotting with the antibodies labelled at right. ‘1×’ represents 5 μg for the anti-TIC20 blot and 2.5 μg for other blots. Cytosolic SPS was also analysed as a control. Data shown as mean ± s.d. of three independent experiments (b); representative of three (c) and two (a) independent experiments.
Extended Data Fig. 7 TIC236 directly binds TOC75 and Arabidopsis tic236-mutant chloroplasts have reduced amounts of TOC–TIC supercomplexes.
a, GST–DUF490 was incubated with a quarter amount of POTRA1–POTRA2–POTRA3–His6 (POTRA1+2+3–His6), POTRA1–POTRA2–His6 (POTRA1+2–His6) or POTRA1–His6. Proteins pulled down by metal-affinity resin were analysed by SDS–PAGE and fluorography. Equal moles of POTRA proteins were loaded among the lanes. The amount of GST–DUF490 pulled down was quantified. b, Membranes from pea chloroplasts treated with 0.5 mM SMCC were solubilized by 1% LDS. The clarified supernatant (input) was immunoprecipitated with anti-TOC75 or the preimmune serum and protein A beads, analysed by SDS–PAGE and immunoblotting, and hybridized to anti-TOC75 or anti-TIC236 antibodies. Similar results were obtained for Arabidopsis chloroplasts (see Fig. 3b). c, We attempted to directly compare the amount of TOC–TIC supercomplexes in mutant and wild-type chloroplasts. However, although we could detect TOC–TIC supercomplex on one-dimensional BN-PAGE immunoblots using anti-TOC75 antibodies12, blotting efficiencies from native gels were variable and not quantitative. Consequently, we used imported [35S]TOC34 to indirectly reflect the amounts of the translocon complexes. [35S]TOC34 was imported into wild-type (WT) and tic236-2 (236) mutant chloroplasts and analysed by BN-PAGE or SDS–PAGE. The two images in the BN-PAGE panel are from the same gel with different exposure times. In wild-type chloroplasts, TOC34 was detected in the 1.25- and 1-megadalton supercomplexes and in a complex about 242 kDa in size (arrowhead). In the tic236-mutant chloroplasts, the same complexes were detected but their amounts were reduced, and the amount of unassembled TOC34 migrating below 200 kDa (bracket) increased. Because the tic236-knockdown mutation does not affect the insertion of TOC34 into the outer membrane (Fig. 2e), these results suggest that a lower amount of the supercomplexes was present in the tic236-mutant chloroplasts. However, we cannot exclude the possibility that mutant chloroplasts were defective in the assembly of TOC34 into supercomplexes. The composition of the 242-kDa, TOC34-containing complex also remains to be determined. Data shown as mean ± s.d. of three independent experiments (a); representative of two (b, c) independent experiments.
Extended Data Fig. 8 Lineage distribution of TIC236 and specificity of the three anti-TIC236 antibody preparations.
a, Phylogenetic relationships of sequences of the DUF490 of TIC236, from bacteria to higher plants. Bootstrap values from 1,000 replicates are indicated. The 0.2 scale shows substitution distance. b, Specificity of the three anti-TIC236 antibody preparations. Pea and Arabidopsis (At) total chloroplast proteins were analysed by SDS–PAGE (10 μg per lane) and immunoblotting, and hybridized to the three antibody preparations against TIC236—a mouse anti-serum against a synthetic peptide corresponding to residues 1957 to 1988 of Arabidopsis TIC236 (anti-peptide), a rabbit antiserum against the DUF490 domain of pea TIC236 (anti-peaDUF490), and a rabbit antiserum against the DUF490 domain of Arabidopsis TIC236 (anti-AtDUF490) (labelled ‘immu’)—as well as their corresponding preimmune sera (labelled ‘preimmu’). Data shown are representative of at least two independent experiments.
Supplementary information
Supplementary Information
This file contains additional discussions on Omp85, the amoeba Paulinilla chromatophora and independent translocation across TOC and TIC, Supplementary References and Supplementary Fig. 1, the full gel blots.
Rights and permissions
About this article
Cite this article
Chen, YL., Chen, LJ., Chu, CC. et al. TIC236 links the outer and inner membrane translocons of the chloroplast. Nature 564, 125–129 (2018). https://doi.org/10.1038/s41586-018-0713-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0713-y
Keywords
This article is cited by
-
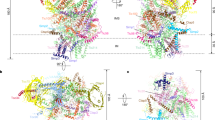
Architecture of chloroplast TOC–TIC translocon supercomplex
Nature (2023)
-
Path unveiled for protein entry into chloroplasts
Nature (2023)
-
Absence of photosynthetic state transitions in alien chloroplasts
Planta (2019)
-
The TOC GTPase Receptors: Regulators of the Fidelity, Specificity and Substrate Profiles of the General Protein Import Machinery of Chloroplasts
The Protein Journal (2019)
-
Exit route evolved into entry path in plants
Nature (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.