Abstract
Protein structures are dynamic and can explore a large conformational landscape1,2. Only some of these structural substates are important for protein function (such as ligand binding, catalysis and regulation)3,4,5. How evolution shapes the structural ensemble to optimize a specific function is poorly understood3,4. One of the constraints on the evolution of proteins is the stability of the folded ‘native’ state. Despite this, 44% of the human proteome contains intrinsically disordered peptide segments greater than 30 residues in length6, the majority of which have no known function7,8,9. Here we show that the entropic force produced by an intrinsically disordered carboxy terminus (ID-tail) shifts the conformational ensemble of human UDP-α-d-glucose-6-dehydrogenase (UGDH) towards a substate with a high affinity for an allosteric inhibitor. The function of the ID-tail does not depend on its sequence or chemical composition. Instead, the affinity enhancement can be accurately predicted based on the length of the intrinsically disordered segment, and is consistent with the entropic force generated by an unstructured peptide attached to the protein surface10,11,12,13. Our data show that the unfolded state of the ID-tail rectifies the dynamics and structure of UGDH to favour inhibitor binding. Because this entropic rectifier does not have any sequence or structural constraints, it is an easily acquired adaptation. This model implies that evolution selects for disordered segments to tune the energy landscape of proteins, which may explain the persistence of intrinsic disorder in the proteome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Frauenfelder, H., Sligar, S. G. & Wolynes, P. G. The energy landscapes and motions of proteins. Science 254, 1598–1603 (1991).
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Campbell, E. et al. The role of protein dynamics in the evolution of new enzyme function. Nat. Chem. Biol. 12, 944–950 (2016).
Boehr, D. D., Nussinov, R. & Wright, P. E. The role of dynamic conformational ensembles in biomolecular recognition. Nat. Chem. Biol. 5, 789–796 (2009).
Kumar, S., Ma, B., Tsai, C. J., Sinha, N. & Nussinov, R. Folding and binding cascades: dynamic landscapes and population shifts. Protein Sci. 9, 10–19 (2000).
Oates, M. E. et al. D2P2: database of disordered protein predictions. Nucleic Acids Res. 41, D508–D516 (2013).
van der Lee, R. et al. Classification of intrinsically disordered regions and proteins. Chem. Rev. 114, 6589–6631 (2014).
Papaleo, E. et al. The role of protein loops and linkers in conformational dynamics and allostery. Chem. Rev. 116, 6391–6423 (2016).
He, B. et al. Predicting intrinsic disorder in proteins: an overview. Cell Res. 19, 929–949 (2009).
Bickel, T., Jeppesen, C. & Marques, C. M. Local entropic effects of polymers grafted to soft interfaces. Eur. Phys. J. E 4, 33–43 (2001).
Bickel, T., Marques, C. & Jeppesen, C. Pressure patches for membranes: the induced pinch of a grafted polymer. Phys. Rev. E 62, 1124–1127 (2000).
Waters, J. T. & Kim, H. D. Calculation of a fluctuating entropic force by phase space sampling. Phys. Rev. E 92, 013308 (2015).
Carmichael, S. P. & Shell, M. S. Entropic (de)stabilization of surface-bound peptides conjugated with polymers. J. Chem. Phys. 143, 243103 (2015).
Ferreon, A. C., Ferreon, J. C., Wright, P. E. & Deniz, A. A. Modulation of allostery by protein intrinsic disorder. Nature 498, 390–394 (2013).
Hilser, V. J. An ensemble view of allostery. Science 327, 653–654 (2010).
Hilser, V. J. & Thompson, E. B. Intrinsic disorder as a mechanism to optimize allosteric coupling in proteins. Proc. Natl Acad. Sci. USA 104, 8311–8315 (2007).
Sugase, K., Dyson, H. J. & Wright, P. E. Mechanism of coupled folding and binding of an intrinsically disordered protein. Nature 447, 1021–1025 (2007).
Li, J. et al. Genetically tunable frustration controls allostery in an intrinsically disordered transcription factor. eLife 6, e30688 (2017).
Egger, S., Chaikuad, A., Kavanagh, K. L., Oppermann, U. & Nidetzky, B. Structure and mechanism of human UDP-glucose 6-dehydrogenase. J. Biol. Chem. 286, 23877–23887 (2011).
Gainey, P. A. & Phelps, C. F. Interactions of uridine-diphosphate glucose dehydrogenase with inhibitor uridine-diphosphate xylose. Biochem. J. 145, 129–134 (1975).
Neufeld, E. F. & Hall, C. W. Inhibition of UDP-d-glucose dehydrogenase by UDP-D-xylose - a possible regulatory mechanism. Biochem. Biophys. Res. Commun. 19, 456–461 (1965).
Beattie, N. R., Keul, N. D., Sidlo, A. M. & Wood, Z. A. Allostery and hysteresis are coupled in human UDP-glucose dehydrogenase. Biochemistry 56, 202–211 (2017).
Kadirvelraj, R., Sennett, N. C., Custer, G. S., Phillips, R. S. & Wood, Z. A. Hysteresis and negative cooperativity in human UDP-glucose dehydrogenase. Biochemistry 52, 1456–1465 (2013).
Kadirvelraj, R., Sennett, N. C., Polizzi, S. J., Weitzel, S. & Wood, Z. A. Role of packing defects in the evolution of allostery and induced fit in human UDP-glucose dehydrogenase. Biochemistry 50, 5780–5789 (2011).
Sennett, N. C., Kadirvelraj, R. & Wood, Z. A. Conformational flexibility in the allosteric regulation of human UDP-α-d-glucose 6-dehydrogenase. Biochemistry 50, 9651–9663 (2011).
Sennett, N. C., Kadirvelraj, R. & Wood, Z. A. Cofactor binding triggers a molecular switch to allosterically activate human UDP-α-d-glucose 6-dehydrogenase. Biochemistry 51, 9364–9374 (2012).
Kadirvelraj, R. et al. Hysteresis in human UDP-glucose dehydrogenase is due to a restrained hexameric structure that favors feedback inhibition. Biochemistry 53, 8043–8051 (2014).
Uversky, V. N. The intrinsic disorder alphabet. III. Dual personality of serine. Intrinsically Disord. Proteins 3, e1027032 (2015).
Theillet, F. X. et al. The alphabet of intrinsic disorder: I. Act like a pro: On the abundance and roles of proline residues in intrinsically disordered proteins. Intrinsically Disord. Proteins 1, e24360 (2013).
Busch, D. J. et al. Intrinsically disordered proteins drive membrane curvature. Nat. Commun. 6, 7875 (2015).
Balasubramaniam, D. & Komives, E. A. Hydrogen-exchange mass spectrometry for the study of intrinsic disorder in proteins. Biochim. Biophys. Acta 1834, 1202–1209 (2013).
Chen, J. Z. Y. Theory of wormlike polymer chains in confinement. Prog. Polym. Sci. 54–55, 3–46 (2016).
Smyda, M. R. & Harvey, S. C. The entropic cost of polymer confinement. J. Phys. Chem. B 116, 10928–10934 (2012).
Thielges, M. C., Chung, J. K., Axup, J. Y. & Fayer, M. D. Influence of histidine tag attachment on picosecond protein dynamics. Biochemistry 50, 5799–5805 (2011).
Wilkins, M. R. et al. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 112, 531–552 (1999).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
Brunger, A. T. Free R value: cross-validation in crystallography. Methods Enzymol. 277, 366–396 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Urzhumtsev, A., Afonine, P. V. & Adams, P. D. TLS from fundamentals to practice. Crystallogr. Rev. 19, 230–270 (2013).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Diederichs, K. & Karplus, P. A. Improved R-factors for diffraction data analysis in macromolecular crystallography. Nat. Struct. Biol. 4, 269–275 (1997).
Frieden, C. Kinetic aspects of regulation of metabolic processes. The hysteretic enzyme concept. J. Biol. Chem. 245, 5788–5799 (1970).
Laue, T. M., Shah, B. D., Ridgeway, T. M. & Pelletier, S. L. Analytical Ultracentrifugation. (Royal Society of Chemistry, London, 1992).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Schuck, P. On the analysis of protein self-association by sedimentation velocity analytical ultracentrifugation. Anal. Biochem. 320, 104–124 (2003).
Ortega, A., Amoros, D. & Garcia de la Torre, J. Prediction of hydrodynamic and other solution properties of rigid proteins from atomic- and residue-level models. Biophys. J. 101, 892–898 (2011).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 8, 275–282 (1992).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Ramachandran, G. N., Ramakrishnan, C. & Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J. Mol. Biol. 7, 95–99 (1963).
Word, J. M., Lovell, S. C., Richardson, J. S. & Richardson, D. C. Asparagine and glutamine: Using hydrogen atom contacts in the choice of side-chain amide orientation. J. Mol. Biol. 285, 1735–1747 (1999).
Huynh, K. & Partch, C. L. Analysis of protein stability and ligand interactions by thermal shift assay. Curr. Protoc. Protein Sci. 79, 28.9.1–28.9.14 (2015).
Fang, J. et al. Conformational dynamics of the Escherichia coli DNA polymerase manager proteins UmuD and UmuD′. J. Mol. Biol. 398, 40–53 (2010).
Wales, T. E. & Engen, J. R. Hydrogen exchange mass spectrometry for the analysis of protein dynamics. Mass Spectrom. Rev. 25, 158–170 (2006).
Johnson, B. et al. Dimerization controls Marburg virus VP24-dependent modulation of host antioxidative stress responses. J. Mol. Biol. 428, 3483–3494 (2016).
Chen, E. et al. Broadly neutralizing epitopes in the Plasmodium vivax vaccine candidate Duffy Binding Protein. Proc. Natl Acad. Sci. USA 113, 6277–6282 (2016).
Yan, Y., Grant, G. A. & Gross, M. L. Hydrogen–deuterium exchange mass spectrometry reveals unique conformational and chemical transformations occurring upon [4Fe-4S] cluster binding in the type 2 l-serine dehydratase from Legionella pneumophila. Biochemistry 54, 5322–5328 (2015).
Palmier, M. O. & Van Doren, S. R. Rapid determination of enzyme kinetics from fluorescence: overcoming the inner filter effect. Anal. Biochem. 371, 43–51 (2007).
Johnson, K. A. Fitting enzyme kinetic data with KinTek Global Kinetic Explorer. Methods Enzymol. 467, 601–626 (2009).
Johnson, K. A., Simpson, Z. B. & Blom, T. Global Kinetic Explorer: a new computer program for dynamic simulation and fitting of kinetic data. Anal. Biochem. 387, 20–29 (2009).
Johnson, K. A., Simpson, Z. B. & Blom, T. FitSpace explorer: an algorithm to evaluate multidimensional parameter space in fitting kinetic data. Anal. Biochem. 387, 30–41 (2009).
Acknowledgements
The authors thank A. P. Karplus, B. W. Matthews, S. N. Savvides and the members of the Z.A.W. laboratory for helpful discussions. We also thank R. Wang of Norclone for producing the R1 truncation constructs. This work was supported by the NIH National Institute of General Medicine grants R01GM114298 awarded to Z.A.W. and P41GM103422 awarded to M.L.G.
Reviewer information
Nature thanks J. Gsponer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
N.D.K. and Z.A.W. designed the study, analysed the data and composed the manuscript. N.D.K. performed the majority of the experiments. K.O. performed all the bioinformatic analyses and conducted the simulations of the ID-tail and analysed the results with S.C.H. S.C.H. contributed to the development of the entropic force model. E.T.S.B. performed HDX mass spectrometry experiments and interpreted the results with M.L.G. Both offered direction in HDX experiment design and contributed to this portion of the manuscript. M.L.G. further contributed with edits to the larger work. N.R.B. performed analysis and refinement of crystal structures. W.E.M. conducted several AUC experiments and performed thermal denaturation assays. R.K. solved Protein Data Bank entry 5VR8. R.S.P. contributed to the development of our kinetic model.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The crystal structures of UGDH(FL) and UGDH(ΔID) show no significant structural deviations, and structural evidence for UDP-Xyl binding in the NAD+ site.
a–c, Structural overlay (root mean square deviation (r.m.s.d) = 0.385 Å), comparing the UGDH(FL) (cyan) and UGDH(ΔID) (grey) E* hexamers (a, b) and monomers (c). PDB entries for UGDH(FL) and UGDH(ΔID) are 4RJT and 5W4X, respectively (Extended Data Table 1). d, Crystal structure of native UGDH with UDP-Xyl bound in the active site. Difference density map (F0 – Fc) of UDP-Xyl (chain B) calculated at 2.0 Å resolution and contoured at 3.5σ. The map was calculated after omitting the UDP-Xyl and subjecting the model to simulated annealing. e–l, UDP-Xyl can also bind weakly to the NAD+-binding site of native UGDH. Difference electron density maps (F0 – Fc) were calculated as in d. The uracil and xylose in the NAD+-binding sites were contoured at 3.5 and 3σ for chain A (e and f, respectively), chain B (g and h, respectively), chain D (i and j, respectively) and chain E (k and l, respectively). Chains C and F do not contain UDP-Xyl in the NAD+-binding site. UDP-Xyl binding in the NAD+ site is the source of mixed inhibition observed in the UGDH(FL-dimer) and UGDH(ΔID-dimer) constructs. (see Supplementary Information, Section 1). PDB entry: 5VR8 (this work, Extended Data Table 1).
Extended Data Fig. 2 Steady-state kinetic analysis of all UGDH constructs.
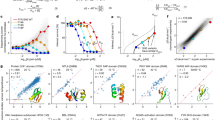
a–w, Inhibition studies with the allosteric inhibitor UDP-Xyl. Data from two or three independent rate curves were globally fit to equation (7) (or equation (8) for dimers c–f) using nonlinear regression (n ≥ 26 data points). See Extended Data Table 2 for the specific number of data points and fit parameters. Dashed lines indicate 95% confidence intervals. x–kk, NAD+ substrate-saturation curves fit to equation (6) using nonlinear regression (n ≥ 10 independent data points). See Extended Data Table 3 for the specific number of data points used in global fitting.
Extended Data Fig. 3 Sedimentation velocity studies of the UGDH constructs.
a–s, Plots of the c(s) distributions with oligomeric species labelled as H (hexamer), T (tetramer), D (dimer) or M (monomer). The R2 mutant (h) shows no change in UDP-Xyl affinity (Fig. 2c and Extended Data Table 2), yet shows evidence of a less stable hexamer. Panel s was included to show that the hexamer in h is less stable partly owing to the K465D substitution in the UGDH(R2) construct. The K465D substitution introduces an unfavourable negative charge near E460 in the hexamer interface, which may reduce the stability. t, Relative distributions, s values (S) and r.m.s.d. values for all sedimentation velocity experiments.
Extended Data Fig. 4 The ID-tail is conserved in vertebrates.
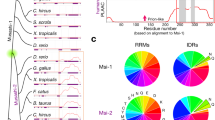
a, ClustalO sequence alignment of all vertebrate UGDH ID-tail regions (79 total). Residues are coloured by type, where blue is positive charge (K, R, H), red is negative charge (D, E), peach is hydrophobic (A, V, L, I, M), orange is aromatic (F, W, Y), green is hydrophilic (S, T, N, Q), yellow is cysteine (C), and magenta is special (P, G). b, The ID-tail was extensively randomized and modified. Sequences of UGDH (FL, R1, R2, −Lys, −Pro, and Ser), aligned by position and coloured by residue type. c, Relative evolutionary rate of UGDH residues from the alignment of 79 vertebrate sequences. The ID-tail (red dots) begins at residue 465 and displays an approximately threefold higher rate of divergence than the folded portion of the protein (black dots). For clarity, only a small, representative segment of the folded protein is shown (residues 460–464). All rates were scaled such that the average rate is 1.0 across the entire dataset.
Extended Data Fig. 5 Exhaustive Monte Carlo simulations constraining the ID-tail.
a, Dashed lines outline the traditional, generously allowed regions of the Ramachandran plot, whereas the red circles identify the conformations used in the Monte Carlo simulations. b, The ranges of ϕ and ψ angles depicted in a. The 10 × 10° bins are centred on the first and last numbers in the range. For example, in region 1, the first ϕ, ψ bin (−155°, 90°) represents the ϕ range −155° to −145° and the ψ range 85° to 95°. c, Ratio of ID-tail conformations constrained (Ω2) to the number of conformations when the ID-tail is unconstrained (Ω1). The entropic costs of confining tails of each length were calculated using equation (1). d, e, The results of the 10-residue ID-tail simulations, shown in a surface representing the hexamer-building interface (orange and grey dimers) with the adjacent ID-tail (cyan) that was fixed during simulations. Also depicted is a representative sampling of 20 allowed Ω2 conformations (blue sticks) from the 4,503 identified in the Monte Carlo simulation. f, The same view as in e, but without the protein surface. g–i, Same as in d–f, but now including a sampling of 20 of the 3,002 Ω1 conformations (red sticks) that clash with the fixed adjacent ID-tail (not depicted for clarity). j–l, Same as g–i, but including 750 of the 142,607 Ω1 conformations (tan sticks) that clash with the protein surface.
Extended Data Fig. 6 The ID-tail induces global changes in the structure and dynamics of UGDH.
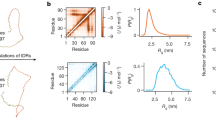
a, The per cent deuterium uptake of the ID-tail peptide region (residues 457–492; red closed circles) saturates rapidly, consistent with an unfolded peptide31. For comparison, two peptides corresponding to the well-ordered α9 helix region (open blue squares and circles) saturate slowly. Data are mean ± s.d. of independently replicated time points (n = 4). For some points, the standard deviation is less than the dimensions of the data symbol. b, The normalized cumulative changes in the hydrogen–deuterium exchange rates (UGDH(FL-dimer) –UGDH(ΔID-dimer)). Most of the kinetics measurements consisted of six independently replicated time points (n = 4), processed to give the mean exchange (red, blue or green bars). Approximately 5% of the data displayed low signal:noise or was missing, and in those cases the means were derived from four or more time points. Results were normalized by dividing by the number of measurements. The propagation error for each peptide is equal to the square root of the sum of all squared standard deviation values for the collective measurement of UGDH(FL-dimer) and UGDH(ΔID-dimer).
Extended Data Fig. 7 Transient-state analysis of UGDH(FL) and UGDH(ΔID).
a, b, Transient-state analysis of UDP-Xyl binding kinetics using intrinsic protein fluorescence. Six independent progress curves (coloured traces) at different inhibitor concentrations were globally fit (black line) to the allosteric inhibition model (see Fig. 4b) for UGDH(FL) and UGDH(ΔID). Each progress curve was replicated (n ≥ 4) with similar results, and the final kinetic model was refined against the averaged progress curves (see e for fit parameters). c, d, Confidence contour plots depicting how constrained each globally fit parameter is relative to the others, for all progress curves in a and b (parameters are listed in e). e, Table of the microscopic rate constants from global fitting of the progress curves described in a and b. The best fit and s.e.m. were obtained from global nonlinear regression based on the numerical integration of rate equations for the described model (see main text and Methods). Upper and lower limits were obtained from the confidence contour analysis. Kd = (K1K2K3)−1, where Kn = kn/k−n. f, Enzyme hysteresis is observed as a lag in progress curves. Representative progress curves (of n = 6 independent measurements) for both UGDH(FL) (cyan) and UGDH(ΔID) (grey) are fit to equation (4) (black line). Curves are displayed with the y axis offset for clarity. Final results for all replicate curves are displayed in Fig. 4a.
Supplementary information
Supplementary Information
This file contains Section 1: hUGDH Dimer [FL and ΔID] Mixed Inhibition
Rights and permissions
About this article
Cite this article
Keul, N.D., Oruganty, K., Schaper Bergman, E.T. et al. The entropic force generated by intrinsically disordered segments tunes protein function. Nature 563, 584–588 (2018). https://doi.org/10.1038/s41586-018-0699-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0699-5
Keywords
This article is cited by
-
Fuzzy recognition by the prokaryotic transcription factor HigA2 from Vibrio cholerae
Nature Communications (2024)
-
The molecular basis for cellular function of intrinsically disordered protein regions
Nature Reviews Molecular Cell Biology (2024)
-
DNA binding redistributes activation domain ensemble and accessibility in pioneer factor Sox2
Nature Communications (2024)
-
Structure of SpoT reveals evolutionary tuning of catalysis via conformational constraint
Nature Chemical Biology (2023)
-
Intrinsically disordered protein biosensor tracks the physical-chemical effects of osmotic stress on cells
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.