Abstract
During both embryonic development and adult tissue regeneration, changes in chromatin structure driven by master transcription factors lead to stimulus-responsive transcriptional programs. A thorough understanding of how stem cells in the skeleton interpret mechanical stimuli and enact regeneration would shed light on how forces are transduced to the nucleus in regenerative processes. Here we develop a genetically dissectible mouse model of mandibular distraction osteogenesis—which is a process that is used in humans to correct an undersized lower jaw that involves surgically separating the jaw bone, which elicits new bone growth in the gap. We use this model to show that regions of newly formed bone are clonally derived from stem cells that reside in the skeleton. Using chromatin and transcriptional profiling, we show that these stem-cell populations gain activity within the focal adhesion kinase (FAK) signalling pathway, and that inhibiting FAK abolishes new bone formation. Mechanotransduction via FAK in skeletal stem cells during distraction activates a gene-regulatory program and retrotransposons that are normally active in primitive neural crest cells, from which skeletal stem cells arise during development. This reversion to a developmental state underlies the robust tissue growth that facilitates stem-cell-based regeneration of adult skeletal tissue.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data to support the conclusions in this manuscript can be found in the figures. All source data for graphs are available in the online version of the paper. Any other data can be requested from the corresponding authors. All ATAC-seq and RNA-seq data can be accessed from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) with accession number GSE104473.
References
Ilizarov, G. A. The tension-stress effect on the genesis and growth of tissues: part II. The influence of the rate and frequency of distraction. Clin. Orthop. Relat. Res. 239, 263–285 (1989).
Ilizarov, G. A. The tension-stress effect on the genesis and growth of tissues. Part I. The influence of stability of fixation and soft-tissue preservation. Clin. Orthop. Relat. Res. 238, 249–281 (1989).
Tahiri, Y., Viezel-Mathieu, A., Aldekhayel, S., Lee, J. & Gilardino, M. The effectiveness of mandibular distraction in improving airway obstruction in the pediatric population. Plast. Reconstr. Surg. 133, 352e–359e (2014).
McCarthy, J. G., Schreiber, J., Karp, N., Thorne, C. H. & Grayson, B. H. Lengthening the human mandible by gradual distraction. Plast. Reconstr. Surg. 89, 1–8 (1992).
Khansa, I. et al. Airway and feeding outcomes of mandibular distraction, tongue-lip adhesion, and conservative management in Pierre Robin sequence: a prospective study. Plast. Reconstr. Surg. 139, 975e–983e (2017).
Chan, C. K. et al. Identification and specification of the mouse skeletal stem cell. Cell 160, 285–298 (2015).
Adam, R. C. et al. Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice. Nature 521, 366–370 (2015).
Ge, Y. et al. Stem cell lineage infidelity drives wound repair and cancer. Cell 169, 636–650 (2017).
Frechin, M. et al. Cell-intrinsic adaptation of lipid composition to local crowding drives social behaviour. Nature 523, 88–91 (2015).
Bell, S. & Terentjev, E. M. Focal adhesion kinase: the reversible molecular mechanosensor. Biophys. J. 112, 2439–2450 (2017).
Rinkevich, Y., Lindau, P., Ueno, H., Longaker, M. T. & Weissman, I. L. Germ-layer and lineage-restricted stem/progenitors regenerate the mouse digit tip. Nature 476, 409–413 (2011).
Lehoczky, J. A., Robert, B. & Tabin, C. J. Mouse digit tip regeneration is mediated by fate-restricted progenitor cells. Proc. Natl Acad. Sci. USA 108, 20609–20614 (2011).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Hojo, H., Ohba, S., He, X., Lai, L. P. & McMahon, A. P. Sp7/Osterix is restricted to bone-forming vertebrates where it acts as a Dlx co-factor in osteoblast specification. Dev. Cell 37, 238–253 (2016).
Meyer, M. B., Benkusky, N. A. & Pike, J. W. The RUNX2 cistrome in osteoblasts: characterization, down-regulation following differentiation, and relationship to gene expression. J. Biol. Chem. 289, 16016–16031 (2014).
Prescott, S. L. et al. Enhancer divergence and cis-regulatory evolution in the human and chimp neural crest. Cell 163, 68–83 (2015).
Tominaga, H. et al. CCAAT/enhancer-binding protein β promotes osteoblast differentiation by enhancing Runx2 activity with ATF4. Mol. Biol. Cell 19, 5373–5386 (2008).
Rogers, C. D., Phillips, J. L. & Bronner, M. E. Elk3 is essential for the progression from progenitor to definitive neural crest cell. Dev. Biol. 374, 255–263 (2013).
Santagati, F. & Rijli, F. M. Cranial neural crest and the building of the vertebrate head. Nat. Rev. Neurosci. 4, 806–818 (2003).
Kaukua, N. et al. Glial origin of mesenchymal stem cells in a tooth model system. Nature 513, 551–554 (2014).
Kaucka, M. et al. Analysis of neural crest-derived clones reveals novel aspects of facial development. Sci. Adv. 2, e1600060 (2016).
Kaucka, M. et al. Oriented clonal cell dynamics enables accurate growth and shaping of vertebrate cartilage. eLife 6, e25902 (2017).
Minoux, M. et al. Gene bivalency at Polycomb domains regulates cranial neural crest positional identity. Science 355, eaal2913 (2017).
Brinkley, J. F. et al. The FaceBase Consortium: a comprehensive resource for craniofacial researchers. Development 143, 2677–2688 (2016).
Attanasio, C. et al. Fine tuning of craniofacial morphology by distant-acting enhancers. Science 342, 1241006 (2013).
Tanaka, H. V. et al. A developmentally regulated switch from stem cells to dedifferentiation for limb muscle regeneration in newts. Nat. Commun. 7, 11069 (2016).
Sánchez Alvarado, A. Developmental biology: a cellular view of regeneration. Nature 460, 39–40 (2009).
Kragl, M. et al. Cells keep a memory of their tissue origin during axolotl limb regeneration. Nature 460, 60–65 (2009).
Purnell, B. A. Regrow like an axolotl. Science 355, 592 (2017).
Nacu, E., Gromberg, E., Oliveira, C. R., Drechsel, D. & Tanaka, E. M. FGF8 and SHH substitute for anterior–posterior tissue interactions to induce limb regeneration. Nature 533, 407–410 (2016).
Roensch, K., Tazaki, A., Chara, O. & Tanaka, E. M. Progressive specification rather than intercalation of segments during limb regeneration. Science 342, 1375–1379 (2013).
Kaufman, C. K. et al. A zebrafish melanoma model reveals emergence of neural crest identity during melanoma initiation. Science 351, aad2197 (2016).
Coufal, N. G. et al. L1 retrotransposition in human neural progenitor cells. Nature 460, 1127–1131 (2009).
Ivancevic, A. M., Kortschak, R. D., Bertozzi, T. & Adelson, D. L. LINEs between species: evolutionary dynamics of LINE-1 retrotransposons across the eukaryotic tree of life. Genome Biol. Evol. 8, 3301–3322 (2016).
Bridier-Nahmias, A. et al. Retrotransposons. An RNA polymerase III subunit determines sites of retrotransposon integration. Science 348, 585–588 (2015).
Mashanov, V. S., Zueva, O. R. & García-Arrarás, J. E. Retrotransposons in animal regeneration: overlooked components of the regenerative machinery? Mob. Genet. Elements 2, 244–247 (2012).
Baillie, J. K. et al. Somatic retrotransposition alters the genetic landscape of the human brain. Nature 479, 534–537 (2011).
Ueno, H. & Weissman, I. L. Clonal analysis of mouse development reveals a polyclonal origin for yolk sac blood islands. Dev. Cell 11, 519–533 (2006).
Wright, D. E., Wagers, A. J., Gulati, A. P., Johnson, F. L. & Weissman, I. L. Physiological migration of hematopoietic stem and progenitor cells. Science 294, 1933–1936 (2001).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Acknowledgements
We thank J. Wysocka for her review of the manuscript and helpful suggestions. We thank the Stanford Functional Genomics Facility, Stanford Cell Sciences Imaging Facility, Lorry Lokey Imaging Facility, and Stanford Shared FACS Facility Cores. We thank D. J. Hunter and D. Atashroo for their respective contributions to the design of the distraction device. This work was supported by the National Institutes of Health (NIH) grants R01DE026730 (to M.T.L. and R.C.R.), U24DE026914 (to M.T.L.) and K08DE024269 (to D.C.W.); the Child Health Research Institute (CHRI) at Stanford University (D.C.W.); The Hagey Laboratory for Pediatric Regenerative Medicine (M.T.L.); the Steinhart/Reed Award (M.T.L); the Gunn–Oliver Fund (M.T.L.); and NIH grant P50-HG007735 and the Scleroderma Research Foundation (H.Y.C.). H.Y.C. is an Investigator of the Howard Hughes Medical Institute.
Reviewer information
Nature thanks C. Tabin, L. Gerstenfeld and P. Scacheri for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
R.C.R. conceived the study and performed microsurgical procedures. R.C.R. and A.C.C. produced figures, wrote the manuscript, and performed ATAC-seq and RNA-seq experiments. A.C.C. performed ATAC-seq and RNA-seq analysis. R.C.R. and A.Sa. performed FACS isolation and experiments based on skeletal stem cells and progenitor cells. Y.W. assisted with ATAC analysis of transposable elements. T.L., O.M., M.L.L., C.D.M., M.P.M., E.Z.S., R.E.J., A.Sh., C.K.F.C. and D.C.W. generated key materials and executed multiple experiments. O.D.K. provided key materials and reagents and support. H.Y.C. and M.T.L. oversaw the work and provided support. All authors reviewed the manuscript and discussed the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Analysis of tissue sources of regeneration in distraction.
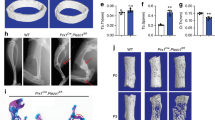
a, Experimental timeline of distraction model. b, Illustration of the tissue response (at POD15) in the mouse model of mandibular distraction (left), fracture (middle) and acute lengthening (right). c, Putative cellular sources of bone regeneration in mandibular distraction, including periosteum, endosteum and circulating progenitors. d, Experimental scheme for detecting circulating progenitor cells in mandibular distraction. GFP mice are surgically fused to their wild-type (WT) littermates through parabiosis. Peripheral blood chimaerism is confirmed via flow cytometry. After GFP-positive cells are confirmed in the WT through FACS, mandibular distraction is performed on the WT parabiont. FSC-A, forward scatter area; FSC-H, forward scatter height. e, Upon detection of 1/1 blood chimaerism, WT parabionts undergo mandibular distraction according to the timeline outlined. f, Quantification of the haematopoietic fraction of GFP-positive cells obtained from distraction calluses (n = 4 biological replicates per time point). g, Representation of the Rainbow reporter construct at the R26 locus and the colours produced by random recombination. h, One-year tracing of mandibles under normal homeostasis (uninjured), with confocal micrographs of whole-mount periosteum one year after recombination. Clones are shown with coloured dotted outlines. The white dotted line demarcates skeletal muscle (upper left quadrant) from the periosteum. The view is a buccal-to-lingual view of the posterior periosteum overlying the body of the mandible. n = 50 clones, with 16–151 cells per clone. i, Confocal micrograph of the transverse mandible section from a Rainbow mouse at POD15 after targeted labelling of the periosteum for subsequent mandibular distraction (n = 8). Coloured outlines indicate single clones; the white dotted outline indicates mandibular bone (mb) at the distraction site. n refers to the number of animals in each independent experiment.
Extended Data Fig. 2 Skeletal lineage tracing demonstrates labelling of bone regenerate in mandibular distraction.
a, Experimental scheme for using tamoxifen (TMX; administered at 9 weeks of age, before surgery at 10 weeks) to induce recombination in Sox9creERT2;R26mT/mG mice to carry out skeletal lineage tracing, with isolation of mandibles at POD29 (n = 6). b, Confocal micrograph (lingual mandible) of the distraction callus in Sox9creERT2;R26mT/mG mice after TMX induction. Filters are shown in the following order from left to right: mT (mTomato, background), mG (mGFP, Sox9 lineage), mTmG (merged) and merged (mTmG with DAPI) (n = 6). c, As for b, but for the buccal mandible. d, Experimental scheme for tamoxifen induction of Sox9creERT2;R26mT/mG mice during the early phase of distraction (POD5–10) for lineage tracing of Sox9+ cells during distraction. Mandibles were isolated at POD29 for confocal microscopy (n = 4). i.p., intraperitoneal. e, Confocal micrographs showing the contribution of the skeletal lineage (mGFP, Sox9 lineage) to new bone formed in mandibular distraction. f, Low-magnification images of the Sox9creERT2;R26Rainbow mandible during distraction at POD8, including bright-field, eGFP, mCerulean and mCherry filters under a dissection microscope. The induced Rainbow-coloured clones (white arrowheads) are anterior and posterior to the site of distraction (dotted white box) between the pin holes (dotted white circles) (n = 8). g, As for f, but for POD12. h, As for Fig. 1l, but for POD12 (n = 5). i, Schematic showing the vantage point for clonal analysis of the skeletal-lineage (Sox9creERT2;R26Rainbow) contribution to regeneration during distraction, through the use of whole-mount specimens from mandibular callus microdissection. DO, distraction osteogenesis. j–l, Quantification of clones (whole-mount imaging) that are positive for each fluorophore per high-pass filter (h.p.f.; ×40 magnification) two weeks after tamoxifen induction (one injection per day, five days total) plus distraction until POD12 using ActincreERT2;R26Rainbow mice (j, k) and Sox9creERT2;R26Rainbow mice (l). Means ± s.d. are shown Scale bars, 200 µm (b, c, e), 1 mm (f). n refers to the number of animals in each independent experiment.
Extended Data Fig. 3 Dynamics of SSCs and progenitor cells in parabiosis and distraction.
a, The skeletal stem cell lineage, showing the immunological phenotype of each cell. CP, chondroprogenitor; OP, osteoprogenitor. b, FACS isolation of single SSCs for evaluation of serial colony-forming potential in vitro. Secondary (left) and tertiary (right) colonies are shown. c, Experimental scheme for detecting circulating cells in mandibular distraction at POD10. d, FACS gating of Thy1−6C3− cells (light brown box) for detection of circulating SSCs (green box in e) and progenitor cells (green box in f) in mandibular distraction. e, FACS gating shows an absence of circulating (GFP+) SSCs in mandibular distraction (n = 8; mean = 2.02 ± 0.78). f, As for e, but for BCSPs (n = 8; mean = 1.83 ± 0.91). g–i, FACS isolation from uninjured (g; 38.0% αVβ3-positive events within the gate) versus fracture (h; 15.0%) and distraction (i; 58.8%) conditions reveal expansion of the SSC hierarchy (left column) in response to distraction by POD10. Representative of three independent experiments. n refers to the number of animals in each independent experiment.
Extended Data Fig. 4 Changes in gene regulation in response to distraction.
a, Schematic showing FACS isolation of stem and progenitor cells from uninjured, fractured and distracted mandibles for ATAC-seq (assay for accessibility of chromatin, which is shown with purple spheres and yellow lines, to depict nucleosomes) and RNA-seq (shown with a polyA strand). Samples were collected in duplicate for ATAC-seq at POD10 and in duplicate for RNA-seq at PODs 5, 10 and 15. b, PCA showing PC1 and PC2 for all ATAC-seq data (n = 164,266 peaks). UI, uninjured. c, Venn diagram showing the number of differential peaks between the fracture and diffraction conditions in both SSCs and BCSPs. Overlap between these (shared peaks; twofold change with P < 0.05 in both SSCs and BCSPs) is shown in the centre. d, Cluster dendrogram for ATAC-seq, showing clustering of samples based on all ATAC-seq peaks from SSCs, BCSPs and OPs. e, Example loci (Mmp15 and Runx2, with their locations on chromosomes 8 and 17, respectively, shown at the top), revealing accessibility that is distraction-specific (Mmp15) and fracture-specific (Runx2), respectively. The height of the genome browser tracks shows the number of reads normalized by read depth and overall peak enrichment in the library. f, PCA showing PC1 and PC2 for all RNA-seq data at PODs 5,10 and 15 (n = 17,491 genes). g, Cluster dendrogram for RNA-seq, showing clustering of samples based on all genes from SSCs, BCSPs and OPs on PODs 5, 10 and 15. h, Volcano plot showing differential gene expression between d-SSCs at PODs 10 and 15 and f-SSCs at PODs 10 and 15. Red dots represent genes that are significantly differentially expressed with an adjusted P-value cut-off of 0.05 (DESeq2). Differentially expressed genes that are upregulated in f-SSCs (Fx) are shown in blue (fold change greater than 1.5; Padj less than 0.05), and differentially expressed genes that are upregulated in d-SSCs (DO) are shown in red. i, Significantly enriched GO terms for genes upregulated in d-SSCs at PODs 10 and 15, from GREAT version 3.0.0, with P values (one-sided binomial) corrected using the Benjamini–Hochberg correction. j, Significantly enriched GO terms for genes upregulated in f-SSCs at PODs 10 and 15 from GREAT, with P values (one-sided binomial) corrected using the Benjamini–Hochberg correction. k, As for h, but for BCSPs. l, As for i, but for BCSPs. m, As for j, but for BCSPs. n, As for h, but for OPs. o, As for i, but for OPs. p, As for j, but for OPs.
Extended Data Fig. 5 Expression of core signalling genes during distraction.
a, Heat map showing the expression of Wnt, Notch, bone morphogenetic protein (BMP) and FGF family members, as well as integrins, matrix metalloproteinases (MMPs) and tissue inhibitors of metalloproteinases (TIMPs), in SSCs. Values from RNA-seq in transcripts per million (t.p.m.) are z-scored for each gene to reflect differences across conditions and time. Only genes that are expressed (t.p.m. > 3) in at least one sample are shown. b, As for a, but for BCSPs. c, As for a, but for OPs. d, Heat map showing expression (in t.p.m.) of Runx-family transcription factors in POD5 SSCs, and in POD10 and POD15 f-SSCs and d-SSCs. e, As for d, but for Dlx-family transcription factors. f, Heat map showing accessibility from ATAC-seq in SSCs and BCSPs at Runx2 chromatin immunoprecipitation (ChIP)-seq peaks from MC3T3 preosteoblasts;12,673 binding sites from ChIP-seq are shown41. Plotted are the row z-scored normalized counts from SSC and BCSP ATAC-seq data. g, As for f, but for Dlx5 ChIP-seq data38. ChIP-seq binding sites (24,365) from MC3T3 preosteoblasts are shown.
Extended Data Fig. 6 Expansion of SSCs and progenitor cells depends on FAK.
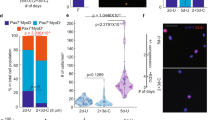
a, Timeline for FAK inhibition and conditional genetic knockout during distraction, with respective control conditions, for µCT analysis (Fig. 3a–d) at mid-consolidation (POD29). b, Pentachrome staining of transverse section from mandibles treated with DMSO during distraction osteogenesis and collected at POD15. The white dotted line indicates cortical bone at the osteotomy site; the red dotted line indicates bone that newly formed in response to distraction. Representative of three independent replicates. c, As for b, but for FAKi treatment. The green dotted line indicates cartilage that newly formed in response to FAKi during distraction osteogenesis. d, As for b, but for POD29. e, As for c, but for POD29. f, Schematic showing FACS isolation of SSCs and BCSPs from mandible calluses for subsequent in vitro analyses. g, h, FACS isolation of SSCs and BCSPs from distraction control mandibles receiving DMSO injections (g; 55.5%) versus FAK-inhibitor injections (FAKi) (h; 41.7%), revealing diminished expansion of the SCC hierarchy (first column, black gate) in response to FAK inhibition by POD10. The ratio of SSCs (red gate) to their committed bone progenitors (BCSPs, green gate) during distraction was substantially disrupted by FAK inhibition, such that the proportion of SSCs was higher. Representative of three independent replicates. i, Quantification of the frequency of d-SSCs, d-BCSPs and d-OPs within mandibular calluses collected at POD10 after FAKi or control injections (n = 6; error bars indicate s.d. from the mean). j, Representative fluorescence micrographs showing phosphorylated-FAK activity (left column, green) in d-SSCs after treatment with DMSO (top) or with FAK inhibitor (bottom). Right column, phospho-FAK fluorescence merged with DAPI fluorescence. Representative of three independent replicates. k, Quantification of d-SSC colony formation in vitro in response to FAKi and in a FAK(cKO), compared with their respective control conditions (n = 15 per condition; ***P < 0.001, Tukey’s multiple comparisons; means ± s.d.). Direct comparison of colony formation in FAKi and FAK(cKO) conditions was not significantly different. n refers to the number of animals in each independent experiment.
Extended Data Fig. 7 Cellular dynamics during distraction osteogenesis in FAK(cKO) mice.
a, b, FACS isolation of cells from control mice (a, Sox9creERT2;Ptk2fl/fl, no TMX) and FAK(cKO) mice (b, Sox9creERT2;Ptk2fl/fl, TMX treatment) shows that the SSC hierarchy (first column, black gate) is disrupted similarly in FAK(cKO) mice and in FAKi (Extended Data Fig. 6h). In the FAK(cKO) mice, the proportion of downstream multipotent BCSPs (green gate) compared with SSCs (orange and red gates) is lower than in controls. Representative of three independent replicates. c, Experimental strategy for clonal analysis of FAK(cKO) Rainbow contribution to regeneration in response to distraction, using whole-mount specimens from callus microdissection of mandibles in Sox9creERT2;R26Rainbow mice. d, Vantage point for the acquisition of the confocal images of whole-mount specimens shown in Fig. 3g, h. e, Experimental strategy for clonal analysis in ActincreERT2;R26Rainbow mice. f, Quantification of average clone size at POD12 during distraction osteogenesis in ActincreERT2;R26Rainbow mice (n = 10; mean ± s.d.). g, Whole-mount imaging at the site of distraction at POD12 in ActincreERT2;R26Rainbow mice. The view is a lateral-to-medial view of callus overlying the distraction site (indicated by the white dotted outline overlying the distraction gap). The distraction gap contains large clones with a migratory spreading phenotype (n = 4 for each of three independent replicates). h, Quantification of clones (whole-mount imaging) that are positive for each fluorophore per h.p.f. (×40 magnification) two weeks after tamoxifen induction (one injection per day, five days total) plus distraction until POD12, using Sox9creERT2;R26Rainbow mice (left) and FAK(cKO)Rainbow mice (right) (n = 10; mean ± s.d.). n refers to the number of animals in each independent experiment.
Extended Data Fig. 8 Involvement of neural crest transcription factors in SSCs during distraction.
a, Motifs enriched in FAK-R and FAK-NR sites in SSCs and BCSPs. The size of each diamond represents the negative P value for enrichment (one-sided binomial, Benjamini–Hochberg correction); colours represent the percentage of sites in target/percentage of sites in background. b, Heat map showing expression of Sox-family transcription factors. Colours represent z-scored t.p.m. values from RNA-seq at PODs 5, 10 and 15 for d-SSCs and f-SSCs. Only factors with t.p.m. greater than or equal to 3 in at least one sample are shown. c, As for b, but for Ets-family transcription factors. d, The Sox10 locus (with its location on chromosome 15 shown at the top), showing accessibility at the promoter that is distraction-specific (red), congruent with the expression data in b and the RNA-seq track below. The signal for tracks is normalized by read depth and overall peak enrichment in the library. Tracks are representative of two biological replicates. e, Experimental scheme for tamoxifen induction of Sox10creERT2;R26mT/mG mice during the early and mid-distraction phase (POD5–10) to trace the Sox10+ lineage in a distraction-specific context. Mandibles were collected for confocal microscopy at POD29 (n = 4). f, Confocal micrograph of Sox10creERT2;R26mT/mG mandible at POD29, demonstrating the capability of Sox10+ cells to give rise to the distraction regenerate. Filters are, from left to right, mT (mTomato, background), mG (mGFP, Sox10+ lineage), mTmG (merged) and merged (mTmG with DAPI). g, Confocal micrograph of Sox10creERT2;R26mT/mG mandible at POD29, showing the presence of Sox10-lineage cells within surrounding callus and periosteum to give rise to the regenerate. Scale bars, 200 µm (b, c). n refers to the number of animals in each independent experiment.
Extended Data Fig. 9 NCC transcriptional networks in d-SSCs.
a, Diagram showing the developmental origin of NCC-derived SSCs. e, embryonic day of development. b, Clustering of SSC and NCC samples using all accessible sites in SSCs and NCCs merged. c, Heat map showing all accessible sites that are significantly differentially accessible between uninjured SSCs and NCC progenitors. Colour represents the log2-transformed normalized read counts within peak regions. d, GO terms enriched in sets of genes near peaks that are more accessible in NCC and d-SSCs than in uninjured SSCs. GO terms shown are those that are significantly enriched after false discovery rate correction in GREAT (two-sided binomial P value shown). e, Heat map showing all accessible sites that are differentially accessible between f-SSCs and d-SSCs. The sites that are FAK-R or FAK-NR are highlighted along the right. Colour represents the log2-transformed normalized read counts within peak regions. f, Quantification of immunofluorescence staining for each of the NCC markers observed on RNA-seq analysis and evaluated in this figure, including nuclear Zeb2 (n = 15, ***P < 0.001), nuclear Twist2 (n = 15, *P < 0.05), Cdc42 (n = 6, P < 0.05), S100a (n = 6, ***P < 0.001) and Plp1 (n = 6, ***P < 0.001) (Student’s t-test; shown are means ± s.d.). n refers to the number of animals in each independent experiment.
Extended Data Fig. 10 Controlled mechanical advancement of the lower jaw unlocks neural crest potential for regeneration of the mandible.
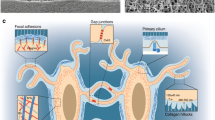
a, At the tissue level, clonality within the mandible during distraction (middle) is observed to occur in a highly linear and directional manner in parallel to the vector of distraction. By contrast, clonality observed in fracture (top) and distraction plus FAKi (bottom) was highly mesenchymal with less apparent organization in its morphology, indicating a nondirectional clonal proliferation. b, The cellular level: the developmental origin of the NCC-derived SSCs of the mandible (top), and the postnatal SSCs of the mandible (bottom) that are present in our experiments. During distraction the d-SSC (shown in pink) demonstrates plasticity and takes on an NCC- derived signature, whereas the f-SSC (purple) retains its postnatal SSC characteristics with no NCC signature. In the absence of FAK signalling, the d-SSC FAKi (green) reverts functionally and epigenomically to the fracture state without emergence of the NCC signature. ‘NCC Prog’ indicates the premigratory (e8.5) NCC progenitor population; ‘NCC Md’ indicates the postmigratory (e10.5) NCC population arriving within the mandible. c, At the chromatin level, distraction induces a gain in accessibility at promoters (P) of FAK-responsive NCC craniofacial genes through the activation of their enhancers (E), with a parallel gain in accessibility of retroviral elements near NCC-specific craniofacial enhancers. Thus mechanotransduction during mandible distraction unlocks FAK-responsive craniofacial enhancers, potentially through retrotransposons, enacting a developmental NCC program in d-SSCs similar to that of the e10.5 NCC Md population (b). This does not occur under fracture conditions (top) or during distraction plus FAKi (bottom). These differential epigenomic responses correlate with the degree of clonality and patterning seen in Rainbow mice (a) that occurs in response to distraction. Circled P represents phosphorylation.
Supplementary information
Supplementary Table 1
FAK-responsive and non-responsive sites in SSCs. a, FAK-responsive sites and nearest genes. b, FAK-nonresponsive sites and nearest genes. c, GO terms enriched in genes nearest to FAK-responsive sites. P-values (one-sided binomial) are corrected using the Benjamini-Hochberg correction. d, Same as c for FAK-nonresponsive sites.
Rights and permissions
About this article
Cite this article
Ransom, R.C., Carter, A.C., Salhotra, A. et al. Mechanoresponsive stem cells acquire neural crest fate in jaw regeneration. Nature 563, 514–521 (2018). https://doi.org/10.1038/s41586-018-0650-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0650-9
Keywords
This article is cited by
-
All-in-one porous membrane enables full protection in guided bone regeneration
Nature Communications (2024)
-
Denervation during mandibular distraction osteogenesis results in impaired bone formation
Scientific Reports (2023)
-
Premature aging of skeletal stem/progenitor cells rather than osteoblasts causes bone loss with decreased mechanosensation
Bone Research (2023)
-
Impaired function of skeletal stem cells derived from growth plates in ovariectomized mice
Journal of Bone and Mineral Metabolism (2023)
-
Cranial Neural Crest Cells Contribution to Craniofacial Bone Development and Regeneration
Current Osteoporosis Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.