Abstract
Biosynthesis of glycogen, the essential glucose (and hence energy) storage molecule in humans, animals and fungi1, is initiated by the glycosyltransferase enzyme, glycogenin (GYG). Deficiencies in glycogen formation cause neurodegenerative and metabolic disease2,3,4, and mouse knockout5 and inherited human mutations6 of GYG impair glycogen synthesis. GYG acts as a ‘seed core’ for the formation of the glycogen particle by catalysing its own stepwise autoglucosylation to form a covalently bound gluco-oligosaccharide chain at initiation site Tyr 195. Precise mechanistic studies have so far been prevented by an inability to access homogeneous glycoforms of this protein, which unusually acts as both catalyst and substrate. Here we show that unprecedented direct access to different, homogeneously glucosylated states of GYG can be accomplished through a palladium-mediated enzyme activation ‘shunt’ process using on-protein C–C bond formation. Careful mimicry of GYG intermediates recapitulates catalytic activity at distinct stages, which in turn allows discovery of triphasic kinetics and substrate plasticity in GYG’s use of sugar substrates. This reveals a tolerant but ‘proof-read’ mechanism that underlies the precision of this metabolic process. The present demonstration of direct, chemically controlled access to intermediate states of active enzymes suggests that such ligation-dependent activation could be a powerful tool in the study of mechanism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Roach, P. J., Depaoli-Roach, A. A., Hurley, T. D. & Tagliabracci, V. S. Glycogen and its metabolism: some new developments and old themes. Biochem. J. 441, 763–787 (2012).
Adeva-Andany, M. M., González-Lucán, M., Donapetry-García, C., Fernández-Fernández, C. & Ameneiros-Rodríguez, E. Glycogen metabolism in humans. BBA Clin. 5, 85–100 (2016).
Roach, P. J. Are there errors in glycogen biosynthesis and is laforin a repair enzyme? FEBS Lett. 585, 3216–3218 (2011).
Zois, C. E., Favaro, E. & Harris, A. L. Glycogen metabolism in cancer. Biochem. Pharmacol. 92, 3–11 (2014).
Testoni, G. et al. Lack of glycogenin causes glycogen accumulation and muscle function impairment. Cell Metab. 26, 256–266 (2017).
Moslemi, A.-R. et al. Glycogenin-1 deficiency and inactivated priming of glycogen synthesis. N. Engl. J. Med. 362, 1203–1210 (2010).
Alonso, M. D., Lomako, J., Lomako, W. M. & Whelan, W. J. Tyrosine-194 of glycogenin undergoes autocatalytic glucosylation but is not essential for catalytic function and activity. FEBS Lett. 342, 38–42 (1994).
Hurley, T. D., Stout, S., Miner, E., Zhou, J. & Roach, P. J. Requirements for catalysis in mammalian glycogenin. J. Biol. Chem. 280, 23892–23899 (2005).
Gibbons, B. J., Roach, P. J. & Hurley, T. D. Crystal structure of the autocatalytic initiator of glycogen biosynthesis, glycogenin. J. Mol. Biol. 319, 463–477 (2002).
Chaikuad, A. et al. Conformational plasticity of glycogenin and its maltosaccharide substrate during glycogen biogenesis. Proc. Natl Acad. Sci. USA 108, 21028–21033 (2011).
Hurley, T. D., Walls, C., Bennett, J. R., Roach, P. J. & Wang, M. Direct detection of glycogenin reaction products during glycogen initiation. Biochem. Biophys. Res. Commun. 348, 374–378 (2006).
Chalker, J. M., Bernardes, G. J. L. & Davis, B. G. A “Tag-and-modify” approach to site-selective protein modification. Acc. Chem. Res. 44, 730–741 (2011).
Chalker, J. M., Wood, C. S. C. & Davis, B. G. A convenient catalyst for aqueous and protein Suzuki-Miyaura cross-coupling. J. Am. Chem. Soc. 131, 16346–16347 (2009).
Spicer, C. D. & Davis, B. G. Palladium-mediated site-selective Suzuki-Miyaura protein modification at genetically encoded aryl halides. Chem. Commun. 47, 1698–1700 (2011).
Spicer, C. D., Triemer, T. & Davis, B. G. Palladium-mediated cell-surface labeling. J. Am. Chem. Soc. 134, 800–803 (2012).
Spicer, C. D. & Davis, B. G. Rewriting the bacterial glycocalyx via Suzuki-Miyaura cross-coupling. Chem. Commun. 49, 2747–2749 (2013).
Dumas, A. et al. Self-liganded Suzuki-Miyaura coupling for site-selective protein PEGylation. Angew. Chem. Int. Edn Engl. 52, 3916–3921 (2013).
Li, J. & Chen, P. R. Moving Pd-mediated protein cross coupling to living systems. ChemBioChem 13, 1728–1731 (2012).
Yang, M., Li, J. & Chen, P. R. Transition metal-mediated bioorthogonal protein chemistry in living cells. Chem. Soc. Rev. 43, 6511–6526 (2014).
Jbara, M., Maity, S. K. & Brik, A. Palladium in the chemical synthesis and modification of proteins. Angew. Chem. Int. Ed. 56, 10644–10655 (2017).
Boeggeman, E. & Qasba, P. K. Studies on the metal binding sites in the catalytic domain of β1,4-galactosyltransferase. Glycobiology 12, 395–407 (2002).
Nielsen, M. M. et al. Substrate and metal ion promiscuity in mannosylglycerate synthase. J. Biol. Chem. 286, 15155–15164 (2011).
Young, T. S., Ahmad, I., Yin, J. A. & Schultz, P. G. An enhanced system for unnatural amino acid mutagenesis in E. coli. J. Mol. Biol. 395, 361–374 (2010).
Davis, L. & Chin, J. W. Designer proteins: applications of genetic code expansion in cell biology. Nat. Rev. Mol. Cell Biol. 13, 168–182 (2012).
Issoglio, F. M., Carrizo, M. E., Romero, J. M. & Curtino, J. A. Mechanisms of monomeric and dimeric glycogenin autoglucosylation. J. Biol. Chem. 287, 1955–1961 (2012).
Bazán, S., Issoglio, F. M., Carrizo, M. E. & Curtino, J. A. The intramolecular autoglucosylation of monomeric glycogenin. Biochem. Biophys. Res. Commun. 371, 328–332 (2008).
Laio, A. & Parrinello, M. Escaping free-energy minima. Proc. Natl Acad. Sci. USA 99, 12562–12566 (2002).
Ardèvol, A. & Rovira, C. Reaction mechanisms in carbohydrate-active enzymes: glycoside hydrolases and glycosyltransferases. insights from ab initio quantum mechanics/molecular mechanics dynamic simulations. J. Am. Chem. Soc. 137, 7528–7547 (2015).
Lee, S. S. et al. Mechanistic evidence for a front-side, SNi-type reaction in a retaining glycosyltransferase. Nat. Chem. Biol. 7, 631–638 (2011).
Ardèvol, A. & Rovira, C. The molecular mechanism of enzymatic glycosyl transfer with retention of configuration: evidence for a short-lived oxocarbenium-like species. Angew. Chem. Int. Edn Engl. 50, 10897–10901 (2011).
Carrizo, M. E., Miozzo, M. C., Goldraij, A. & Curtino, J. A. Purification of rabbit skeletal muscle proteoglycogen: studies on the glucosyltransferase activity of polysaccharide-free and -bound glycogenin. Glycobiology 7, 571–578 (1997).
Alonso, M. D., Lomako, J., Lomako, W. M. & Whelan, W. J. A new look at the biogenesis of glycogen. FASEB J. 9, 1126–1137 (1995).
Ashcroft, F. M., Rohm, M., Clark, A. & Brereton, M. F. Is type 2 diabetes a glycogen storage disease of pancreatic β cells? Cell Metab. 26, 17–23 (2017).
Lee, H.-M., Larson, D. R. & Lawrence, D. S. Illuminating the chemistry of life: design, synthesis, and applications of “caged” and related photoresponsive compounds. ACS Chem. Biol. 4, 409–427 (2009).
Li, J. et al. Palladium-triggered deprotection chemistry for protein activation in living cells. Nat. Chem. 6, 352–361 (2014).
Guibé, F. Allylic protecting groups and their use in a complex environment Part II: allylic protecting groups and their removal through catalytic palladium π-allyl methodology. Tetrahedron 54, 2967–3042 (1998).
Tsuji, J. New general synthetic methods involving π-allylpalladium complexes as intermediates and neutral reaction conditions. Tetrahedron 42, 4361–4401 (1986).
Tsuji, J. & Yamakawa, T. A convenient method for the preparation of 1-olefins by the palladium catalyzed hydrogenolysis of allylic acetates and allylic phenyl ethers with ammonium formate. Tetrahedron Lett. 20, 613–616 (1979).
Acknowledgements
This work was supported by grants from the EPSRC (DTA to C.D.S. and M.K.B.), MINECO (CTQ2017-85496-P to C.R.), AGAUR (2017SGR-1189 to C.R.), Spanish Structures of Excellence María de Maeztu (MDM-2017-0767 to C.R.), EU Horizon 2020 programme, Marie Skłodowska-Curie (67507) and the Royal Society (Wolfson Research Merit Award to B.G.D.). We thank G. Hemberg for useful discussions, P. G. Schultz for provision of initial pEVOL-pIPhe plasmid, and BSC-CNS for computer resources and technical support at the MareNostrum supercomputer (RES-QCM-2018-2-0025). The Structural Genomics Consortium is a registered charity (number 1097737) that receives funds from AbbVie, Bayer Pharma AG, Boehringer Ingelheim, Canada Foundation for Innovation, Eshelman Institute for Innovation, Genome Canada, Innovative Medicines Initiative (EU/EFPIA) (ULTRA-DD grant number 115766), Janssen, Merck & Co., Novartis Pharma AG, Ontario Ministry of Economic Development and Innovation, Pfizer, São Paulo Research Foundation-FAPESP, Takeda, and Wellcome Trust (092809/Z/10/Z).
Author information
Authors and Affiliations
Contributions
M.K.B., S.S.L., C.D.S., W.W.Y. and B.G.D. designed the project. M.K.B., T.M. and M.A.G. carried out chemical synthesis, protein modification reactions, enzymatic assays and associated analysis. H.J.B., S.S.L., C.D.S., M.K.B., T.M. and M.A.G. carried out protein expression. H.J.B. performed protein expression optimization and crystallography experiments. L.R., J.I.-F. and C.R. performed computational experiments. M.K.B., C.R., W.W.Y. and B.G.D. wrote the manuscript. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biological and chemical methods for glycogen assembly.
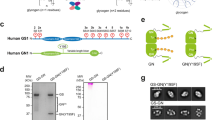
a, Mechanism of glycogen biosynthesis in eukaryotes. The glycosyltransferase enzyme glycogenin (GYG; top left) catalyses its successive, stepwise autoglucosylation at Tyr195 to form a short enzyme-bound maltooligosaccharide, which undergoes further extension and branching catalysed by glycogen synthase (GYS) and glycogen branching enzyme (GBE) to form mature glycogen particles. Intermediate glucosylation states of glycogenin cannot be isolated in homogeneous form via conventional biological methods, hindering precise mechanistic studies. b, The formation of glycogen from GYG placed in the context of the entire glycogenesis pathway (beginning from free glucose) and glycogenolysis pathway. A wide range of pathological conditions (text in red) are known to arise from malfunction of one of the many enzymes (text in black) involved. c, An alternative chemical method, involving site-selective ligation of sugar-mimic moieties to a GYG species bearing an amino acid ‘tag’ at the 195 site, may allow direct access to defined, homogeneous GYG glucosylation states corresponding to intermediates along the autoglucosylation pathway. This could, in turn, enable study of GYG mechanism with increased precision, through analysis of individual autoglucosylation steps.
Extended Data Fig. 2 Development of chemically addressable GYG scaffold as a strategy for mechanistic investigation.
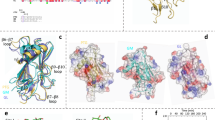
a, GYG-pIPhe195 enzyme (top, middle), which lacks a native glycosyl acceptor and thus cannot undergo glucosylation, represents a suitable substrate for Suzuki–Miyaura cross-coupling to a range of boronic acids (boxed), to generate potentially active enzyme species that mimic defined GYG glycoforms. In this way, inactive GYG-pIPhe195 might be activated through C–C bond-forming ligation allowing pre-determined, ‘shunted’ access to intermediate catalyst states of GYG. b, Expression of GYG-195pIPhe in Escherichia coli using a polyhistidine tag removable by TEV cleavage was confirmed by LC-MS (shown; y axis, m/z ratio) and showed structural similarity to wild-type enzyme. Similar LC-MS spectra were obtained for at least three analogous expressions. Circular dichroism plots (one shown, with ellipticity versus wavelength) are mean averages of three successive measurements of the same sample (n = 1). c, d, Overlay (c) and (d) enlarged view of the chain A acceptor arm for the structures of GYG-Y195X (Mn2+ + UDP; this study) (red), GYG-Phe (Mn2+ + UDPG) (yellow), GYG-WT (Glc4 + UDP) (blue), and GYG-WT (Glc6 + UDP) (green). Inset to d, 2Fo − Fc electron density map for Y195pIPhe of the acceptor arm. The acceptor arm in chain B is disordered (marked by red-dashed line in c) and probably adopts multiple conformations to accommodate the equivalent pIPhe group. e, Evidence of dimer formation for GYG in solution. Left, SEC-SAXS signal plot. Each orange point represents the integrated area of the ratio of the sample SAXS curve to the estimated background. Each blue point shows radius of gyration (Rg) estimated from the Guinier region for each frame. Right, main panel, logarithmic intensity plot of subtracted and merged SAXS frames. Cyan circles represent averaged buffer frames subtracted from averaged sampled frames. Black circles represent the median of the buffer frames subtracted from the averaged sample frames. Above, aligned, averaged and refined DAMMIN ab initio model (grey) superimposed with the dimeric GYG1 crystal structure (PDB 6eqj) using supcomb. SAXS analysis was performed from 16 independent scattering measurements of one biological sample.
Extended Data Fig. 3 Effect of Suzuki–Miyaura reagents and conditions on autoglucosylation activity and one-pot Suzuki–Miyaura autoglucosylation.
a, Effect of different Suzuki–Miyuara reaction (scheme shown) components on GYG-WT activity. Bottom, plots illustrating proportions of glycoforms present before (black) and after (red) glucosylation assay, as calculated from the relative peak intensities of each glycoform in the corresponding LC-MS spectra, and represent single experiments (n = 1) carried out in parallel. Boronic acid and the Pd-scavenger DTT (dithiothreitol) did not appear to cause enzymatic inactivation in isolation. In the presence of palladium with limited Pd-removal/refolding steps, either far more limited activity (Pd + DTT) or no activity (all components) was seen. This highlighted that Pd was the key issue regarding GYG-WT activity. b, The effect of cross-coupling reaction components on GYG-WT structure, as shown by circular dichroism analysis. Neither DTT nor boronic acid (m-CH2OH) caused any substantial alteration to secondary structure. Palladium catalyst, however, caused clear alteration of secondary structure. This effect could however be avoided through minimized Pd concentrations and thorough Pd-scavenging and removal (see c). Circular dichroism plots represent mean averages of three measurements of the same sample (n = 1). c, Demonstration of ‘enzyme-compatible’ Pd-mediated ligation. Through minimizing the Pd concentrations employed, and post-reaction Pd removal, cross-coupling conditions compatible with retention of GYG-WT activity were developed (top row). The plot in the bottom row illustrates proportions of glycoforms present before (black) and after (red) glucosylation assay, as calculated from relative peak intensities of each glycoform in corresponding LC-MS spectra (boxed), and represents a single experiment (n = 1; note however that the ‘1-m-CH2OH’ experiment in Extended Data Fig. 4 is near-identical, differing only in glucosylation time). Circular dichroism (rightmost plot, bottom row) additionally confirmed structural similarity of GYG-WT enzyme before and after subjection to optimized cross-coupling conditions. See also Extended Data Fig. 2 for more details of structural analyses by X-ray crystallography. Circular dichroism plots represent mean averages of three measurements of the same sample (n = 1). d, One-pot SMC of GYG-Y195X and autoglucosylation of GYG-Glc (top row). Autoglucosylation is greater when palladium scavenging is carried out before glucosylation assay quenching (reaction 1) than vice versa (reaction 2). LC-MS data represent single experiments run in parallel (n = 1). e, Average number of glucoses added per enzyme upon treatment of GYG-WT in the presence of varying final concentrations of Pd (0, 0.1, 0.2, 0.4 mM). Data are mean average of three independent replicates (n = 3); error bars are ±1 s.d.
Extended Data Fig. 4 Cross-coupling to ‘simple mimics’ and assessment of catalytic activity of products.
Top row, ‘simple’ substrate templates were introduced using aryl boronic acids 1-o-CH2OH, 1-m-CH2OH, 1-p-CH2OH (exploring different angles of nucleophile display) and 1-m-OH, 1-p-OH (exploring reduced nucleophile length with similar angles) (shown in second row). These proceeded with useful to high conversions (Supplementary Methods, Section 6) to allow the direct creation of systematically altered GYG conjugates bearing substrate mimics: GYG-o-CH2OH, GYG-m-CH2OH, GYG-p-CH2OH, GYG-m-OH, GYG-p-OH. LC-MS analysis showed that none of the cross-coupled products showed autoglucosylation activity (upper scheme, detail in Supplementary Methods, Section 6). Irrespective of the systematically varied nature of the glycan-mimic substrate templates (orientation, length or pKa), none led to efficient mimicry of substrate moiety and hence activation of autoglucosylation. Notably, also, the truncated, ‘linker-only’ variant GYG-allyl-OH was inactive, highlighting further the critical need of an effective mimic moiety for such ‘shunting’. Thus, despite successful Pd-mediated ligation, these ‘simple’ templates provided ineffective mimicry. Identically treated GYG-WT, run in parallel as a positive control in each case, showed detectable activity (central scheme). Bar charts (bottom two rows) are graphical representations of the LC-MS data for treated GYG-WT, showing abundances of each GYG glycoform before (black) and after (red) autoglucosylation assay. Bar charts are representations of single LC-MS experiments (n = 1, Supplementary Tables 13, 14).
Extended Data Fig. 5 Proposed species observed during cross-couplings to GYG-Y195X and possible mechanisms responsible for their formation.
a, Left to right, unreacted GYG-Y195X (in certain cases), cross-coupled product, de-iodination product, and a species observed uniquely in carbohydrate couplings and proposed to result from ‘reductive substitution’ of the carbohydrate moiety (with ‘hydride’, probably from a hydrido-palladium species). b, Formation of cross-coupling side-products is illustrated using coupling to boronic acid 1-Glc as an example. The relationship of these possible mechanisms to that of productive Suzuki–Miyaura cross-coupling is highlighted. The generally accepted mechanism for de-iodination (red) involves coordination of a β-hydride-containing ligand to Pd following the initial oxidative addition step of cross-coupling; subsequent β-hydride elimination affords a hydrido-palladium species, reductive elimination from which affords de-iodinated side-product. ‘Reductive substitution’, seen here for carbohydrate systems only (blue), could be rationalized by the well documented ability of Pd to cleave Callyl–O bonds, including those of allylic glycosides such as GYG-Glc, to form a π-allyl species. Quenching of the latter with the same hydrido-palladium complex as invoked in de-iodination—such use of a ‘hydride scavenger’ in Pd-catalysed de-allylation is a well documented process36,37,38—would result in a ‘reductive substitution’ product, that is, replacement of the carbohydrate moiety (here, glucose) with hydride. Blue sphere, Glc, ‘Ar–I’, pIPhe195 of GYG.
Extended Data Fig. 6 Distributions of glycoforms in assays of GYG variants and the proposed triphasic mechanism.
a, Species with up to 12 glucose sugar residues attached are observed during autoglucosylation of GYG-WT-0Glc after 900 s reaction time (the key shows reaction time). Glc-0 is slow to decline, while accumulation of later glycoforms (for example, Glc-7, Glc-8) is observed. This is consistent with the slow-fast-slow profile observed for the cross-coupled system, highlighting the relevance of the latter. b, Species with up to 13 sugars attached are observed during autoglucosylation of GYG-Glc (left) and GYG-Glc-Glc (right). c, No species with more than 8 sugars is seen during autogalactosylation of the same enzymes even after 120 min reaction time. Data represent mean averages from n independent replicate kinetic assays (n = 4 for GYG-Glc glucosylation, n = 5 for GYG-Glc-Glc glucosylation, n = 3 for all others). Error bars are ±1 s.d. d, Proposed triphasic mechanism inferred from kinetic experiments. GYG-Glc-Glc exhibits only two distinct phases (fast → slow); the slower initiation step for GYG-Glc represents an additional first (slow) phase before reaching the disaccharide of GYG-Glc-Glc.
Extended Data Fig. 7 Kinetic analyses and comparison of GYG-WT-0Glc with ‘shunted’ GYG-glycomimetics.
a, Left, reaction scheme (top) with R variants under (boxed). Right, overlayed kinetic profiles (left plot) and initial rates (right plot) for GYG-WT-0Glc, GYG-Glc, GYG-Glc-Glc and GYG-Glc6. Data are shown as mean ± s.d. from n independent replicate kinetic assays (n = 4 for GYG-Glc, n = 5 for GYG-Glc-Glc, n = 3 for GYG-WT-0Glc, n = 2 for GYG-Glc6). b, Apparent rate constants for autoglucosylation of GYG-WT-0Glc, GYG-Glc, GYG-Glc-Glc and GYG-Glc6 allowed us to re-construct, using autoglucosylation kinetic parameters of each of the ‘shunted’ glycomimetics (right), an autoglucosylation profile in good agreement with the GYG-WT-0Glc kinetic data (left: top overlay, profile; bottom overlay, nonlinear fit). Data are shown as mean ± s.d. for n independent replicate kinetic assays (n = 4 for GYG-Glc, n = 5 for GYG-Glc-Glc, n = 3 for GYG-WT-0Glc, n = 2 for GYG-Glc6). c, To ensure that there were no potential artefactual catalytic effects from a de-iodinated side-product giving rise to this previously unobserved rapid phase via intermolecular glucosylation, we also explored its effect when added in pure form to reactions (scheme shown left); rather than any enhancement it gave rise only to slight suppression, thereby discounting this possibility. Observed rates (k-values) are essentially independent of levels of GYG-Y195F, which lacks native acceptor capacity, highlighting that our conclusions are not influenced by such cross-coupling side products. Data are shown as mean ± s.d. for n independent replicate kinetic assays (n = 3).
Extended Data Fig. 8 Glycogenin dynamics, glycoform mimics and active site structure considering acceptors of different length.
a, Root mean square fluctuation (RMSF) of the enzymatic Cα atoms (top). Results are obtained from the MD simulations of the intra ‘UDP-Glc + GYG-WT-Glc3’ Michaelis complex. Coloured segments in plot correspond to the regions coloured in the structure below (see Supplementary Information for details). b, Top, modelled complex of the GYG glycoform mimic (right column), in comparison with the WT complex (left column). Blue balls represent Glc units, and the red and thin rectangle represents the allyl moiety. Bottom, normalized distributions of the donor–acceptor C1–O4 distances. Both inter and intra conformations gave similar results. c, Structural superposition of intra and inter conformations for ‘UDP-Glc + GYG-WT-Glcn’ complexes, with n = 0 to n = 5 Glc units (six left-hand panels). The orange loop corresponds to the acceptor arm of the same subunit of the active site that is displayed (that is, intra), and the white loop is the acceptor arm of the opposite subunit (inter). The protein loop coloured in blue hinders the approach of short acceptors in intra conformations. Specifically, the loop clashes with Tyr195 for the n = 0 (intra), causing it to move away from the donor, as indicated by the black arrow. The n = 1 (intra) is also affected by the loop, as reflected in the shift of the corresponding C1–O4 frequency peak. Frequency distributions are shown in the respective six right-hand panels. Each distribution was obtained from 0.4 μs of simulation data. The maximum frequency peak for intra/inter conformations correspond to 6.4/3.8 Å (n = 0), at 3.6/3.2 Å (n = 1), 3.3/3.3 Å (n = 2) and 3.2/3.2 Å (n = 3). Hydrogen atoms have been omitted for clarity.
Extended Data Fig. 9 Glycogenin plasticity and simulations of the glucosylation reaction mechanism.
a, Modelled intra ‘UDP-Glc + (Glc)2-Tyr195’ complexes (panels on left) and normalized distribution of the C1–O4 distances considering donor and acceptor Gal variants (panels on right). The change of Glc (blue) to Gal (yellow) in the acceptor displaces the reactive hydroxyl by 0.5 Å (from 3.3 Å to 3.8 Å). The Gal modification at the donor site displays alternative conformations (not shown) in which the OH-2 and OH-3 substituents interact with D163. Hydrogen atoms have been omitted for clarity. b, Computed free-energy landscape for the intra ‘UDP-Glc + GYG-WT-Glc3’ reaction catalysed by GYG (contour lines at 1 kcal mol−1; left panel) and atomic rearrangement along the reaction pathway (six panels on the right). Hydrogen atoms have been omitted for clarity, except OH-2, OH-3 and OH-4 of the acceptor sugar, the OH-2 of the donor sugar and those of the side-chain amide NH2 of Q164. Bonds being broken/formed are represented as dashed red lines (snapshots 1 and 3). c, Hydrogen-bond interactions (dashed lines; from PDBs 3T7O, 3U2V and 3U2U) that were restrained during the first steps of the initial classical MD simulations. Q164, interacting with the acceptor OH-3, is not shown for clarity.
Extended Data Fig. 10 Other methods considered for generation of GYG glycoforms.
Illustrated here are hypothetical strategies for generating a GYG-monosaccharide conjugate based on tags incorporated through amber codon suppression to ensure site-specific glycosylation; the corresponding natural glycoform is shown for comparison. The top boxed scheme chosen in this study is compared with other possible and useful methods shown below. Subjective views on perceived, possible advantages of our chosen strategy (Suzuki–Miyaura coupling; boxed) are highlighted, along with reasons for our selection over alternative approaches.
Supplementary information
Supplementary Information
Supplementary Methods and Supplementary Tables 1-14.
Supplementary Video
Supplementary Video of QM/MM analysis of 'Extension Phase 2’.
Rights and permissions
About this article
Cite this article
Bilyard, M.K., Bailey, H., Raich, L. et al. Palladium-mediated enzyme activation suggests multiphase initiation of glycogenesis. Nature 563, 235–240 (2018). https://doi.org/10.1038/s41586-018-0644-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0644-7
Keywords
This article is cited by
-
Molecular basis for bacterial N-glycosylation by a soluble HMW1C-like N-glycosyltransferase
Nature Communications (2023)
-
Two distinct catalytic pathways for GH43 xylanolytic enzymes unveiled by X-ray and QM/MM simulations
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.