Abstract
Ebola virus causes haemorrhagic fever with a high fatality rate in humans and non-human primates. It belongs to the family Filoviridae in the order Mononegavirales, which are viruses that contain linear, non-segmented, negative-sense, single-stranded genomic RNA1,2. The enveloped, filamentous virion contains the nucleocapsid, consisting of the helical nucleoprotein–RNA complex, VP24, VP30, VP35 and viral polymerase1,3. The nucleoprotein–RNA complex acts as a scaffold for nucleocapsid formation and as a template for RNA replication and transcription by condensing RNA into the virion4,5. RNA binding and nucleoprotein oligomerization are synergistic and do not readily occur independently6. Although recent cryo-electron tomography studies have revealed the overall architecture of the nucleocapsid core4,5, there has been no high-resolution reconstruction of the nucleocapsid. Here we report the structure of a recombinant Ebola virus nucleoprotein–RNA complex expressed in mammalian cells without chemical fixation, at near-atomic resolution using single-particle cryo-electron microscopy. Our structure reveals how the Ebola virus nucleocapsid core encapsidates its viral genome, its sequence-independent coordination with RNA by nucleoprotein, and the dynamic transition between the RNA-free and RNA-bound states. It provides direct structural evidence for the role of the N terminus of nucleoprotein in subunit oligomerization, and for the hydrophobic and electrostatic interactions that lead to the formation of the helical assembly. The structure is validated as representative of the native biological assembly of the nucleocapsid core by consistent dimensions and symmetry with the full virion5. The atomic model provides a detailed mechanistic basis for understanding nucleocapsid assembly and highlights key structural features that may serve as targets for anti-viral drug development.
Similar content being viewed by others
Main
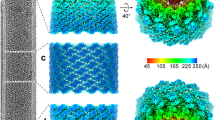
We expressed and purified C-terminally truncated Zaire ebolavirus nucleoprotein (NP), containing residues 1−450 (NP 1−450) in a human cell line (Fig. 1a), in which it sequestered cellular RNA and assembled into a rigid helix indistinguishable from the viral nucleocapsid core3,4,5,7. The sample was imaged without fixation using cryo-electron microscopy (cryo-EM), and its structure was determined by single-particle analysis (SPA) (Fig. 1b, Extended Data Fig. 1a–e, Extended Data Table 1). The complex forms a left-handed helical tube with outer and inner diameters of 295 and 175 Å, respectively, and features a characteristic zipper-like arrangement of parallel C-terminal α-helices (Fig. 1b–d). Its diameter and helical symmetry correspond closely to those of the recently reported Ebola virion reconstructed by cryo-electron tomography (cryo-ET)4,5. Our cryo-EM map enabled generation of an atomic model, including clearly resolved RNA nucleotides (Fig. 1e, Extended Data Figs. 1d, 1f, 2).
a, Structural layout of the Ebola NP sequence. The full-length NP contains 739 amino acids divided into an N-terminal arm domain, an NP core consisting of N- and C-terminal lobes, a disordered linker and a C-terminal tail. b, The iso-electron potential surface of NP–RNA complex reconstruction (contoured at 3σ above average) was calculated from 117,552 single-particle images. The RNA strand and a single NP subunit are highlighted in red and orange, respectively. c, d, The isolated NP subunit (2.5σ) with our atomic model (RCSB Protein Data Bank code (PDB): 5Z9W) in ribbon representation. e, Isolated electron potential map (5.0σ) of RNA superimposed with the atomic model of RNA. RNA bases were modelled as uracil. There are 6 nucleotides per NP. Scale bars, 20 Å.
The transition from the RNA-free to the RNA-bound state requires conformational changes in both RNA and NP. Encapsidation of RNA by NP has been addressed by two leading models. One model has NP encapsidating the RNA in a closed conformation with a hinge-like motion around the junction between N- and C-terminal lobes6,8,9,10,11,12,13,14. The other has RNA and a phosphoprotein competing directly for the RNA-binding cleft15,16 (Extended Data Fig. 3). To discriminate between these possibilities, we compared our atomic model of RNA-bound NP with structures of RNA-free monomeric NP chaperoned by phosphoprotein (VP35)6,8. We found that the relative positions around the junction—specifically, α-helices α9−α12 in the N-terminal lobe and α13−α15 in the C-terminal lobe—do not change significantly, which indicates that the transition is not a simple hinge motion. Binding regions of RNA and the N-terminal peptide of VP35 do not overlap, weakening the case for direct competition. These observations imply that the transition to the closed conformation is a consequence of multiple elements. In Ebola virus NP, the β-hairpin (β3−β4, which is unique to the family Filoviridae) is rotated approximately 90° between RNA-free and RNA-bound states (Fig. 2a, Extended Data Fig. 3, Supplementary Video 1). Simultaneously, the adjacent 310 helix η2 and α-helices α16−α18 are shifted to form a ‘closed’ state (Fig. 2a, Extended Data Fig. 3a). The C-terminal helix α18 clamps the RNA strand and serves as a fastener to cover the RNA in its cleft, as previously reported5. A short 310 helix η5 (residues 329−333), which does not exist in the RNA-free state, shortens and directs the adjacent loop 334–338 to secure the RNA tightly (Fig. 2a, b, Extended Data Fig. 3a).
a, Comparison of an RNA-free, monomeric NP model (PDB: 4YPI, grey) and RNA-bound NP (PDB: 5Z9W, coloured, from this study). The conformational transition to the encapsidated state is orchestrated by a 90° rotation of the β-hairpin, formation of a new secondary structural element (η5) and shift of the C-terminal α-helices. Arrows indicate transitions from RNA-free (open) state to a bound (closed) state. The green arrow indicates formation of a 310 helix η5 and shifting of the polypeptide chain. See also Supplementary Video 1. b–d, Close-up view of the NP–RNA binding region. Blue dashed lines depict predicted hydrogen bonds. b, 310 helix η5, loop 334–338 and helix α18 coordinate RNA bases and flexibly adopt the RNA path. c, Interactions between positively charged NP residues and RNA-backbone phosphates. d, Possible hydrogen bond of E309 with the adjacent RNA pentose hydroxyl group. e, Schematic of RNA nucleotides within the nucleoprotein. Two RNA base triplets with opposing orientations are coordinated by each NP subunit. The RNA bases are numbered sequentially in the 3′ to 5′ direction, representing negative-sense genomic RNA, facing inward (green) towards the helical tube or outward (red). A set of specific electrostatic interactions on both the NP core and α18 determine nucleotide orientation. Secondary structure labels are based on our atomic model (see Extended Data Fig. 2).
The viral RNA is firmly secured in the NP–RNA complex within a positively charged ellipsoid cleft with a cross-section approximately 5 × 11 Å (Extended Data Fig. 4). The conformation of the RNA strand follows an undulating path along the helical NP strand. We found 6 RNA nucleotides clamped in the cleft (Fig. 1c, d), confirming predictions4,5,6,8,9,17. These nucleotides adopt a ‘3-bases-inward, 3-outward’ configuration (Figs. 1e, 2a, b), similar to measles virus and parainfluenza virus 513,18. Coordination is achieved by hydrogen bonding and electrostatic interactions between NP and negatively charged phosphate groups of RNA.
Polar residues on helix η5 and loop 334–338 are within hydrogen-bonding distance of the RNA backbone, adjacent to helix α17, and clamping helix α18 (Fig. 2b). Therefore, this region not only aligns the RNA but also serves as an interface for NP–RNA interactions. Positively charged residues K160, K171, R174, R298 and R401—some of which are important for RNA binding in filovirus NP8,19—were found in close proximity to RNA nucleotides, with their side chains pointing towards RNA-backbone phosphates (Fig. 2c). K248, which is non-essential for RNA binding8,19, faces the RNA bases. These common residues suggest a shared structural motif for RNA interactions within the family Filoviridae. Unlike in measles virus, we found no indication of π–π stacking as a possible cause of the switch between ‘3-in’ and ‘3-out’ nucleotide directions13. The location of R298 on the inner side of the cleft enables it to interact with the RNA backbone. Combined with the restricted space in the cavity and R401 on helix α18 on the outside, these structural constraints may provide an alternative mechanism for the switch. In addition, E309 is within hydrogen-bonding distance of the closest RNA nucleotide on the next NP subunit (Fig. 2d). These specific interactions with the RNA backbone are consistent with the sequence-independent NP–RNA interaction of negative-sense RNA viruses12,13,15,20,21,22,23,24,25 (Fig. 2e).
The N-terminal arm domain of NP is essential for NP oligomerization6,8. Our map clearly shows that the N-terminal arms cross-link subunits on the inside of the helical tube (Fig. 3a–c, Extended Data Fig. 5a, d), confirming previous observations5. Furthermore, N-terminal residues have been predicted to associate with the same binding site as the N terminus of VP355,6. VP35 binds to a hydrophobic pocket in the NP C-terminal lobe and acts as a chaperone to keep NP in its monomeric RNA-free state6. To clarify the mechanism of NP oligomerization, we examined the interface between the N-terminal arm and the pocket of adjacent NPs (Fig. 3b, c). The N-terminal arm binds to the same location as VP35 within this pocket via I24, L25 and A27 on its short N-terminal α-helix (Fig. 3c, Extended Data Fig. 5a–d). Whereas VP35 straddles the β-hairpin in the RNA-free monomeric form6,8 (Extended Data Fig. 5a, c), adjacent NP subunits extend their N-terminal arm to interact directly with the rotated β-hairpin and the pocket in the NP–RNA complex (Fig. 3c). Furthermore, the N-terminal arm is tucked behind K257 on the rotated β-hairpin and the pocket (Fig. 3c, d, Extended Data Fig. 5d). Compared with monomeric NP, we discovered that loop 220–227 moves to create a groove between itself and helix α4, which makes space for the N-terminal arm of the adjacent NP (Fig. 3e, Extended Data Fig. 5e). This conformational change exposes two amino acids (D226 and D229) that serve as electrostatic anchors for partners (R37 and R39) on the N-terminal arm. This observation is consistent with a previous report of increased solvent exchange in this area upon oligomerization6.
We refer to the NP–RNA band generically as a ‘strand’, ‘intrastrand interactions’ in the lateral direction and ‘interstrand interactions’ in the axial direction. a, An atomic model of the NP–RNA complex. Larger and smaller dashed boxes outline close-ups of two adjacent NP molecules (b) and interstrand NP–NP interface (g), respectively. b, An NP pair viewed from inside the helix. Dashed boxes delineate respective close-up figures. c, Intrastrand NP–NP interactions between the N-terminal arm and the hydrophobic pocket. The β-hairpin is shown in dark grey. d, Schematics of the hydrophobic pocket illustrating the binding mode of the N-terminal arm in the RNA-bound state (top) and VP35 in the RNA-free state (bottom). e, Contacts between the N-terminal arm and the N-terminal lobe. f, The contact region between α17 of NPn and α18 of NPn + 1 subunits. The blue dashed line indicates a hydrogen bond. g, Interstrand NP–NP interactions (with subunits NPn ± 24 and NPn ± 25) formed by 3 pairs of amino acid side chains, K110–E349, E92–Q371 and R105–Q381.
Lateral assembly of NP monomers relies on NP–NP and NP–RNA interactions. Beside the N-terminal arm, multiple contact regions are essential for RNA encapsidation26. Whereas the N-terminal arm of vesicular stomatitis virus is co-located with the NP–NP interface, contact regions in NP 1–450 are more spatially separated. Stoichiometric excess of VP35 does not dissociate the NP oligomer after the NP–RNA complex has been formed4,6,27. Our data and previous reports show that the NP–RNA complex retains lateral binding of NP subunits to form a flat ribbon-like strand, even where the NP–RNA complex is uncoiled4,6,7,8 (Extended Data Fig. 1a, b). We identified a major NP–NP interface between the parallel C-terminal extended α-helices α17 and α18 (Fig. 3a, b, f), which contains surface patches formed by highly conserved hydrophobic residues (Fig. 3f, Extended Data Fig. 5b). These patches are buried in the NP oligomer. In non-physiological packing within the crystallographic unit cells of filovirus NP, α17 and α18 of adjacent NPs also make contacts in almost identical configurations8,19 (Extended Data Fig. 5f), which emphasizes the high specificity of these helix–helix interactions. The function of the hydrophobic interface may be to align the C-terminal α-helices to form the characteristic zipper-like arrangement throughout the NP strand.
In our structure of NP 1–450, the interstrand interface is uniquely determined by NP–NP interactions. In the axial direction, any arbitrary NPn subunit shares contact regions with two pairs of neighbours located above and below (Fig. 3a). Three pairs of amino acids, K110–E349, E92–Q371 and R105–Q381, are located near enough for electrostatic interaction to occur between the zipper-like C-terminal α-helices and the β-sheet of the N-terminal lobe (Fig. 3g). Of note, most other amino acids at the interstrand interface within 2−6 Å of each other are also charged or polar (Extended Data Fig. 6), suggesting that the nature of interstrand NP–NP interaction is mainly electrostatic. This broad distribution of charged residues may allow distance variation without rupture upon helix bending, and would explain the phenotypic flexibility of filoviruses. The hydrophilic character of the interstrand interface further explains how the NP–RNA complex can easily uncoil, yet remain stable in solution.
Oligomeric NP structures determined at intermediate resolution by cryo-ET5 and SPA28 have previously been reported. The helical parameter of our NP–RNA complex is consistent with the full virion in the cryo-ET study5. However, our structure differs from the bacterially expressed, in vitro-assembled NP (residues 25–457)28, which formed helical tubes of heterodimeric NP subunits with nearly twice the diameter of native Ebola virus. That structure has fundamentally different helical symmetry and contains no RNA density (Extended Data Fig. 7). Nevertheless, this N-terminal truncation mutant can still assemble into a paired back-to-back helical configuration with a partially disordered C-terminal helix located away from the RNA cleft. This further emphasizes the importance of the first 24 N-terminal residues for cooperative control in alignment and oligomerization of NP subunits, the clamping helix α18 and the RNA itself during assembly of the NP–RNA complex under physiological conditions.
Here we report the structure of the Ebola virus NP 1–450 RNA complex at near-atomic resolution. Insights from our study provide opportunities for rational design of novel antivirals—for example, creation of peptidomimetics competing with the binding site for the N-terminal arm helix or the C-terminal hydrophobic patches, or by cross-linking the interstrand interface. On the basis of its 6 nucleotides per NP subunit and genome size of the Zaire ebolavirus (18,960 bases), the length of the nucleocapsid necessary to accommodate a single copy of genomic RNA is 950 μm. This dimension is consistent with the average length of the filamentous Ebola virion29,30. Furthermore, a histogram of virion length shows a peak distribution with integer multiples of this distance30. Because the outer surface of mature Ebola nucleocapsid is in contact with matrix protein VP40, the NP–RNA complex may serve as a yardstick to determine virion length, causing pinch-off at its ends. Our structure supports conclusions by previous studies5,6—chiefly, the important role of the N-terminal arm in subunit linking and in competing with the VP35 N-terminal peptide. Encapsidation of the viral genome takes place concomitantly with genome replication, and lateral NP–NP interactions are required for assembly of the nucleocapsid. Therefore, NP oligomerization in open conformation by self-alignment via the hydrophobic interface must precede RNA clamping or occur simultaneously. Because NP oligomerization and NP–RNA binding are cooperative, the end of the RNA strand would terminate assembly.
Methods
Purification of the Ebola NP–RNA complex
Human embryonic kidney 293T cells (tested mycoplasma-free, authenticated by DNA fingerprinting and obtained from Kawaoka Lab at Institute of Medical Science, University of Tokyo) were transfected with a plasmid encoding truncated nucleoprotein (NP residues 1–450) of Zaire ebolavirus (strain Mayinga-76, GenBank code AF499101), the type species of the genus Ebolavirus31. Three days after transfection, cells were rinsed with Tris-HCl buffer (10 mM Tris-HCl (pH 7.8), 150 mM NaCl, 1 mM EDTA), and lysed with 0.1% Nonidet P-40 in buffer supplemented with a protease inhibitor (Roche) and an RNase inhibitor (Promega). The lysate was centrifuged at 20,000g at 4 °C for 10 min to remove insoluble compounds. Supernatant was loaded onto a discontinuous 25–40% (wt/wt) CsCl gradient and centrifuged at 250,000g at 4 °C for 1 h. Absorbance of fractions at 280 nm was measured, and peak fractions were subjected to centrifugation at 200,000g at 4 °C for 30 min. The pellet was suspended in the buffer and stored at 4 °C.
Cryo-EM specimen preparation
Two microlitres purified NP–RNA complex solution at protein concentration of 0.3 mg ml−1 was applied to C-Flat CF-1.2/1.3-4C grids (Protochips), previously treated with H2/O2 plasma (Gatan Solarus). Grids were blotted on a Vitrobot Mark IV (Thermo Fisher Scientific) before plunge freezing in liquid ethane.
Cryo-electron microscopy
Images of the liquid nitrogen-cooled sample were acquired on a Titan Krios cryo-TEM (Thermo Fisher Scientific) equipped with an energy filter (Gatan Quantum 968) at 300 kV in EFTEM nanoprobe mode. Data were acquired automatically as movies with the Leginon System32 on a K2 Summit detector (Gatan). For details, see Extended Data Table 1.
Image processing
Super-resolution movie frames were 2× Fourier-cropped, gain-normalized, aligned, dose-weighted and summed using MOTIONCOR233. Helices were manually traced with E2HELIXBOXER32. Subsequent helical image processing steps were performed with SPRING34 and RELION 2.135,36. Defocus values were estimated with CTFFIND437. Classes with visual details were selected after two rounds of 2D classification in RELION 2.1. Helical symmetry parameters (helical pitch and subunits per turn) were exhaustively explored by multiple real-space correlation searches with the program SEGCLASSRECONSTRUCT of the software package SPRING34. The search provided many correlation peaks corresponding to various combinations of parameters. Then, 3D classification in RELION 2.1 was performed with symmetry search around the respective peaks (search range for pitch ± 10%, rotation ± 0.6 subunits per turn). Eighty-eight per cent of the segments fell into three of four classes with nearly identical helical symmetry and yielded reasonable reconstructions at high resolution. These three classes were combined and submitted to further 3D refinement with an additional symmetry search (resulting in no significant change in helical symmetry). The helical symmetry parameter converged to pitch = 73.56 Å, 24.44 subunits per turn (rise = 3.01 Å, rotation = 14.73°). A SPIDER38-generated cylinder-shaped electron density map was used as an initial alignment reference for 3D classification and refinement in RELION 2.1. The resolution estimate of the final reconstruction was based on Fourier shell correlation (FSC) between independently refined half sets using the high-resolution noise substitution method39 and a soft-edged mask. Local resolution was calculated with RELION 2.1. The locally-sharpened map was normalized with MAPMAN40. Detailed image processing conditions are listed in Extended Data Table 1.
Atomic model building and refinement
Atomic modelling of the NP 1−450 helix used template-based and de novo structure modelling. First, a previously reported crystal structure of Ebola NP residues 38−385 (PDB: 4YPI)8 served as the initial model and was docked as a rigid body into our 3.6 Å-resolution cryo-EM map. Next, N- and C-terminal residues of NP, and RNA nucleotides, were added, and the model was adjusted manually with COOT41. The RNA was built de novo with COOT. During manual model building, stereochemistry of peptide bonds was monitored using a Ramachandran plot42. N- and C-terminal residues 1−18 and 407−450 were not modelled owing to a lack of interpretable map features, although the highly flexible disordered C-terminal region was recognizable in 2D class-average images (Extended Data Fig. 1c). The model was refined including data to 3.6 Å resolution with phenix.real_space_refine43 in the presence of 8 surrounding subunits including an extended RNA strand. Secondary structure elements for protein and base-pair stacking for RNA were estimated by phenix.secondary_structure_restraints43, and used as restraints for the refinement. The model was validated with MolProbity44. The unambiguous assignment of nucleic acid density in our map represents an independent element of map validation. Detailed conditions of model refinement and validation are given in Extended Data Table 1. The electron potential map and atomic model were visualized with UCSF Chimera and ChimeraX45,46.
Statistics and reproducibility
Data collection and atomic model statistics are listed in Extended Data Table 1. The method at the core of this work is cryo-EM and single particle analysis. The reconstruction was assembled from a total number of 117,552 single particle images. These images represent projections of a common 3D object (the NP–RNA complex) and were averaged in three dimensions to result in the present reconstruction. The process used two independent sets, each containing half the number of images. These half-set reconstructions were compared to each other using a FSC, which measures the degree of correspondence as a function of resolution. Using the established criterion for resolution measurement (FSC = 0.143 for independent datasets), the two half-set reconstructions agreed up to a spatial resolution of 3.6 Å.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Change history
06 January 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41586-021-04304-6
References
Lamb, R. A. in Fields Virology Vol. 1 (eds Knipe, D. M. & Howley, P. M.) Ch. 30, 880–884 (Wolters Kluwer/Lippincott Williams & Wilkins, 2013).
Ruigrok, R. W., Crepin, T. & Kolakofsky, D. Nucleoproteins and nucleocapsids of negative-strand RNA viruses. Curr. Opin. Microbiol. 14, 504–510 (2011).
Huang, Y., Xu, L., Sun, Y. & Nabel, G. J. The assembly of Ebola virus nucleocapsid requires virion-associated proteins 35 and 24 and posttranslational modification of nucleoprotein. Mol. Cell 10, 307–316 (2002).
Bharat, T. A. et al. Structural dissection of Ebola virus and its assembly determinants using cryo-electron tomography. Proc. Natl Acad. Sci. USA 109, 4275–4280 (2012).
Wan, W. et al. Structure and assembly of the Ebola virus nucleocapsid. Nature 551, 394–397 (2017).
Kirchdoerfer, R. N., Abelson, D. M., Li, S., Wood, M. R. & Saphire, E. O. Assembly of the Ebola virus nucleoprotein from a chaperoned VP35 complex. Cell Reports 12, 140–149 (2015).
Noda, T., Hagiwara, K., Sagara, H. & Kawaoka, Y. Characterization of the Ebola virus nucleoprotein–RNA complex. J. Gen. Virol. 91, 1478–1483 (2010).
Leung, D. W. et al. An intrinsically disordered peptide from ebola virus VP35 controls viral RNA synthesis by modulating nucleoprotein-RNA interactions. Cell Reports 11, 376–389 (2015).
Dong, S. et al. Insight into the Ebola virus nucleocapsid assembly mechanism: crystal structure of Ebola virus nucleoprotein core domain at 1.8 Å resolution. Protein Cell 6, 351–362 (2015).
Yabukarski, F. et al. Structure of Nipah virus unassembled nucleoprotein in complex with its viral chaperone. Nat. Struct. Mol. Biol. 21, 754–759 (2014).
Yabukarski, F. et al. Ensemble structure of the highly flexible complex formed between vesicular stomatitis virus unassembled nucleoprotein and its phosphoprotein chaperone. J. Mol. Biol. 428, 2671–2694 (2016).
Renner, M. et al. Nucleocapsid assembly in pneumoviruses is regulated by conformational switching of the N protein. eLife 5, e12627 (2016).
Gutsche, I. et al. Near-atomic cryo-EM structure of the helical measles virus nucleocapsid. Science 348, 704–707 (2015).
Guryanov, S. G., Liljeroos, L., Kasaragod, P., Kajander, T. & Butcher, S. J. Crystal structure of the measles virus nucleoprotein core in complex with an N-terminal region of phosphoprotein. J. Virol. 90, 2849–2857 (2015).
Green, T. J., Zhang, X., Wertz, G. W. & Luo, M. Structure of the vesicular stomatitis virus nucleoprotein–RNA complex. Science 313, 357–360 (2006).
Leyrat, C. et al. Structure of the vesicular stomatitis virus N0–P complex. PLoS Pathog. 7, e1002248 (2011).
Weik, M., Enterlein, S., Schlenz, K. & Muhlberger, E. The Ebola virus genomic replication promoter is bipartite and follows the rule of six. J. Virol. 79, 10660–10671 (2005).
Alayyoubi, M., Leser, G. P., Kors, C. A. & Lamb, R. A. Structure of the paramyxovirus parainfluenza virus 5 nucleoprotein–RNA complex. Proc. Natl Acad. Sci. USA 112, E1792–E1799 (2015).
Liu, B. et al. Structural insight into nucleoprotein conformation change chaperoned by VP35 peptide in Marburg virus. J. Virol. 91, (2017).
Albertini, A. A. et al. Crystal structure of the rabies virus nucleoprotein–RNA complex. Science 313, 360–363 (2006).
Tawar, R. G. et al. Crystal structure of a nucleocapsid-like nucleoprotein–RNA complex of respiratory syncytial virus. Science 326, 1279–1283 (2009).
Hastie, K. M. et al. Crystal structure of the Lassa virus nucleoprotein–RNA complex reveals a gating mechanism for RNA binding. Proc. Natl Acad. Sci. USA 108, 19365–19370 (2011).
Raymond, D. D., Piper, M. E., Gerrard, S. R., Skiniotis, G. & Smith, J. L. Phleboviruses encapsidate their genomes by sequestering RNA bases. Proc. Natl Acad. Sci. USA 109, 19208–19213 (2012).
Ariza, A. et al. Nucleocapsid protein structures from orthobunyaviruses reveal insight into ribonucleoprotein architecture and RNA polymerization. Nucleic Acids Res. 41, 5912–5926 (2013).
Niu, F. et al. Structure of the Leanyer orthobunyavirus nucleoprotein–RNA complex reveals unique architecture for RNA encapsidation. Proc. Natl Acad. Sci. USA 110, 9054–9059 (2013).
Zhang, X., Green, T. J., Tsao, J., Qiu, S. & Luo, M. Role of intermolecular interactions of vesicular stomatitis virus nucleoprotein in RNA encapsidation. J. Virol. 82, 674–682 (2008).
Noda, T., Kolesnikova, L., Becker, S. & Kawaoka, Y. The importance of the NP:VP35 ratio in Ebola virus nucleocapsid formation. J. Infect. Dis. 204, S878–S883 (2011).
Su, Z. et al. Electron cryo-microscopy structure of Ebola virus nucleoprotein reveals a mechanism for nucleocapsid-like assembly. Cell 172, 966–978 (2018).
Geisbert, T. W. & Jahrling, P. B. Differentiation of filoviruses by electron microscopy. Virus Res. 39, 129–150 (1995).
Beniac, D. R. et al. The organisation of Ebola virus reveals a capacity for extensive, modular polyploidy. PLoS ONE 7, e29608 (2012).
Watanabe, S., Noda, T. & Kawaoka, Y. Functional mapping of the nucleoprotein of Ebola virus. J. Virol. 80, 3743–3751 (2006).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Desfosses, A., Ciuffa, R., Gutsche, I. & Sachse, C. SPRING—an image processing package for single-particle based helical reconstruction from electron cryomicrographs. J. Struct. Biol. 185, 15–26 (2014).
He, S. & Scheres, S. H. W. Helical reconstruction in RELION. J. Struct. Biol. 198, 163–176 (2017).
Scheres, S. H. Processing of structurally heterogeneous cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Kleywegt, G. J. et al. The Uppsala electron-density server. Acta Crystallogr. D 60, 2240–2249 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Ramachandran, G. N., Ramakrishnan, C. & Sasisekharan, V. Stereochemistry of polypeptide chain configurations. J. Mol. Biol. 7, 95–99 (1963).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Brasseur, R. Differentiation of lipid-associating helices by use of three-dimensional molecular hydrophobicity potential calculations. J. Biol. Chem. 266, 16120–16127 (1991).
Acknowledgements
We thank Y. Takizawa, A. Maigné and M. Collins for helpful discussions and S. D. Aird for editing the manuscript. This work was supported by a Grant-in-Aid for Research Activity Start-up from MEXT (Grant no. 26892028) (to Y.S.), by Joint Research Project of Institute of Medical Science, University of Tokyo (to Y.S. and T.N.), by Joint Research Program of Institute for Frontier Life and Medical Sciences, Kyoto University (to Y.S.), by JST (Grant no. JPMJPR13L9), by AMED (Grant no. 17fk0108128h0001), from JSPS Core-to-Core Program A and from Daiichi Sankyo Foundation of Life Science (to T.N.), by J-PRIDE for Global Epidemic from AMED (JP17fm0208101j0001), by RPERID from AMED (JP17fk0108029h0002) and by a Grant-in-Aid for Scientific Research on Innovative Areas from MEXT (Grant no. 16H06429, 16K21723 and 16H06434) (to Y.K.). Y.S. and M.W. were supported by direct funding from Okinawa Institute of Science and Technology Graduate University.
Reviewer information
Nature thanks I. Gutsche, M. Luo and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.S., T.N. and M.W. designed experiments. Y.S. purified NP and prepared specimen. Y.S. and M.W. collected cryo-EM data and reconstructed the 3D structure. Y.S. and H.M. built and refined the atomic model. Y.K. and T.N. provided Ebola viral genes and valuable advice. M.W. supervised the project. Y.S. wrote the initial draft and created all figures. Y.S. and M.W. wrote the paper with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM analysis and atomic model building of NP 1−450.
a, One representative image of 2,467 digital micrographs showing NP helices in amorphous ice. The dashed box depicts the close-up in b. b, Magnified image from a, and a schematic of the interface between the ordered helical array and the uncoiled protein–RNA strand (right). c, Two-dimensional class-average image from ~9,000 aligned segmented projections (top, middle), back-projection from the 3D reconstruction (top, right) and their corresponding power spectra (below). The left-most power spectrum is the squared Fourier transform of the vertically aligned longest-helix image from the digital micrograph in a. Blurry disordered C-terminal regions are visible in the class-average image, protruding radially outward (arrows). Power spectra of the helix image and the class average are twofold binned (Nyquist limit at edge: 5.56 Å). All contain visible layer lines. The highest-resolution layer line in the unbinned power spectrum extends to 3.7 Å. d, FSC from independently refined datasets indicates overall map resolution of 3.63 Å (FSC = 0.143) (red curve), which is validated by the cross-FSC between cryo-EM map and model-generated electron-density map (black curve, 3.91 Å at FSC = 0.5). e, Local-resolution map. f, Parts of the cryo-EM electron potential map superimposed upon the atomic model of the N-terminal arm, α9, and C-terminal α18. All maps are contoured at 2.5σ above average. The map was sharpened with a B-factor of −100 Å2. Scale bars, 500 Å in a; 50 Å in b, c, e.
Extended Data Fig. 2 Secondary structure assignment of Ebola viral NPs aligned with filovirus NPs.
The secondary structure assignments from our atomic model of NP 19−406 (PDB: 5Z9W) in the presence of RNA, and the crystal structure of monomeric RNA-free NP 38−385 (PDB: 4YPI) are depicted at the top and second row, respectively. Sequence alignment of nucleoproteins from Zaire ebolavirus (EBOV), Lake Victoria marburgvirus (MARV), and Lloviu cuevavirus (LLOV) (UniProt: P18272, P35263 and G8EFI1) was performed with ClustalW2.147. Conserved and similar residues are boxed in red and white, respectively. The figure was prepared with phenix.secondary_structure_restraints43 and ESPript3.148.
Extended Data Fig. 3 Conformational changes upon RNA binding of nucleoproteins (NP or N) chaperoned by phosphoproteins (VP35 or P) in the order Mononegavirales.
Structures were rendered in ribbon and tube representations (α-helices) based on their atomic coordinates. Only core domains (N- and C-terminal lobes, beige and green) and RNA (red) are shown, with the exception of P in vesicular stomatitis virus (VSV, orange). The two models were aligned relative to the N-terminal lobe. Schematic and summary on the right. The monomeric, RNA-free state (grey) is superimposed on the RNA-bound oligomeric state (colour). a, Ebola virus (EBOV) (NP-VP35, PDB: 4YPI8; NP–RNA, PDB: 5Z9W, this study) in the family Filoviridae. b, c, Measles virus (MeV) (N–P, PDB: 5E4V14; N–RNA, PDB: 4UFT13) (b) and human metapneumovirus (HMPV) (N–P, PDB: 5FVD; N–RNA, PDB: 5FVC)12 (c) in the family Paramyxoviridae. d, VSV (N–P, PDB: 3PMK16; N–RNA, PDB: 2GIC12) in the family Rhabdoviridae.
Extended Data Fig. 4 NP–RNA interactions.
a, Electrostatic surface surrounding the RNA cleft. Scale ranging from −10 (red) to +10 kcal mol−1 e−1 (blue), calculated from the atomic model of our NP monomer (PDB: 5Z9W). b, Close-up view with dimensions of the cleft shown in a without showing the RNA.
Extended Data Fig. 5 Intrastrand NP–NP interactions.
a, Comparison of NP structures. The VP35-bound, RNA-free monomeric state (PDB: 4YPI, orange and red, and 4ZTG, sky blue and blue) and the RNA-bound oligomeric state (PDB: 5Z9W, grey and yellow) were aligned relative to the whole polypeptide. VP35 and the N-terminal arm bind to the same hydrophobic pocket. b–d, Molecular lipophilicity potential maps (MLPP, Brasseur method49) of our structure (PDB: 5Z9W), coloured blue (hydrophilic) to orange (hydrophobic). c, A close-up of VP35-bound NP. d, A close-up of NP bound by the N-terminal arm. e, The contact region of the N-terminal arm and the N-terminal lobe of two adjacent NPs. f, A comparison of two adjacent NPs in the NP–RNA complex (PDB: 5Z9W, pink and red) and NP crystal packing (PDB: 4YPI). The two models were aligned relative to α17.
Extended Data Fig. 6 Interstrand NP–NP interface.
A close-up of the interstrand NP–NP interface shared among NPn, NPn−24 and NPn−25 subunits. Amino acid side chains within a distance of ≤ 6 Å between NPn and NPn−25 and between NPn and NPn−24 are shown with their labels.
Extended Data Fig. 7 Structural comparison of the currently available NP assemblies.
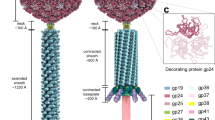
a, Summary of the comparison among reported NP structures. b, Overall and close-up iso-potential surface views of helical NP assemblies (NP 1−450 by our study, NP 1−450 and nucleocapsid in virion5 and NP 25−45728). Experimental cryo-EM maps were normalized and contoured at 2.5σ, 2.5σ, 2.0σ and 2.5σ above average, respectively. Two adjacent NP subunits are coloured according to the assignment in Fig. 1, using their corresponding atomic models within 4 Å distance (PDB: 5Z9W, 6EHL, 6EHM and 6C54). The approximate location of RNA is indicated by red dashed boxes. VP24 in the virus reconstruction is coloured in white (density is clipped at higher radius to reveal the RNA). Scale bar, 500 Å. c, Comparison between NP 1−450 structures from our study (coloured) and ref. 5 (grey). The structures agree in their overall fold. However, the course of the backbone deviates in regions with poor density—in particular, at the N terminus, the β-hairpin and around the RNA interface. The 310 helix η5, which had not previously been described, is missing. Root mean square deviation (calculated with UCSF Chimera) among all 387 atoms pairs is 1.97 Å.
Supplementary information
Video 1
Animated visualization of conformational changes upon RNA binding. Animation of the dynamic transition between the RNA-free open state and the RNA-bound closed state of the truncated Ebola virus nucleoprotein. The atomic coordinates are rendered as a ribbon model (α-helices as rods, β-strands as arrows). The conformational transition to the encapsidated (closed) state is not a simple hinge motion, but can be summarized as coordinated movements of multiple components: 90º rotation of the β-hairpin, formation of a new secondary structural element (η5), and shift of the C-terminal α-helices.
Rights and permissions
About this article
Cite this article
Sugita, Y., Matsunami, H., Kawaoka, Y. et al. Cryo-EM structure of the Ebola virus nucleoprotein–RNA complex at 3.6 Å resolution. Nature 563, 137–140 (2018). https://doi.org/10.1038/s41586-018-0630-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0630-0
Keywords
This article is cited by
-
Structural landscape of the respiratory syncytial virus nucleocapsids
Nature Communications (2023)
-
Poxvirus under the eyes of electron microscope
Applied Microscopy (2022)
-
Structural insight into Marburg virus nucleoprotein–RNA complex formation
Nature Communications (2022)
-
The two-stage interaction of Ebola virus VP40 with nucleoprotein results in a switch from viral RNA synthesis to virion assembly/budding
Protein & Cell (2022)
-
Structural plasticity of mumps virus nucleocapsids with cryo-EM structures
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.