Abstract
The transcriptional co-activator p300 is a histone acetyltransferase (HAT) that is typically recruited to transcriptional enhancers and regulates gene expression by acetylating chromatin. Here we show that the activation of p300 directly depends on the activation and oligomerization status of transcription factor ligands. Using two model transcription factors, IRF3 and STAT1, we demonstrate that transcription factor dimerization enables the trans-autoacetylation of p300 in a highly conserved and intrinsically disordered autoinhibitory lysine-rich loop, resulting in p300 activation. We describe a crystal structure of p300 in which the autoinhibitory loop invades the active site of a neighbouring HAT domain, revealing a snapshot of a trans-autoacetylation reaction intermediate. Substrate access to the active site involves the rearrangement of an autoinhibitory RING domain. Our data explain how cellular signalling and the activation and dimerization of transcription factors control the activation of p300, and therefore explain why gene transcription is associated with chromatin acetylation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Coordinates for the p300 core structure and BΔRP bound to a diacetylated histone H4 peptide are available from the Protein Data Bank (PDB) under accession numbers 6GYR and 6GYT, respectively. Source data are available for Fig. 1b, f and Extended Data Fig. 1d. Figure 1d shows the initial velocities from reactions shown in Extended Data Fig. 1d.
References
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS–STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Panne, D., McWhirter, S. M., Maniatis, T. & Harrison, S. C. Interferon regulatory factor 3 is regulated by a dual phosphorylation-dependent switch. J. Biol. Chem. 282, 22816–22822 (2007).
Zhao, B. et al. Structural basis for concerted recruitment and activation of IRF-3 by innate immune adaptor proteins. Proc. Natl Acad. Sci. USA 113, E3403–E3412 (2016).
Parekh, B. S. & Maniatis, T. Virus infection leads to localized hyperacetylation of histones H3 and H4 at the IFN-β promoter. Mol. Cell 3, 125–129 (1999).
Panne, D., Maniatis, T. & Harrison, S. C. An atomic model of the interferon-beta enhanceosome. Cell 129, 1111–1123 (2007).
Stark, G. R. & Darnell, J. E., Jr. The JAK-STAT pathway at twenty. Immunity 36, 503–514 (2012).
Zhang, J. J. et al. Two contact regions between Stat1 and CBP/p300 in interferon gamma signaling. Proc. Natl Acad. Sci. USA 93, 15092–15096 (1996).
Bedford, D. C. & Brindle, P. K. Is histone acetylation the most important physiological function for CBP and p300? Aging 4, 247–255 (2012).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Visel, A. et al. ChIP–seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
Jin, Q. et al. Distinct roles of GCN5/PCAF-mediated H3K9ac and CBP/p300-mediated H3K18/27ac in nuclear receptor transactivation. EMBO J. 30, 249–262 (2011).
Bedford, D. C., Kasper, L. H., Fukuyama, T. & Brindle, P. K. Target gene context influences the transcriptional requirement for the KAT3 family of CBP and p300 histone acetyltransferases. Epigenetics 5, 9–15 (2010).
Zhao, L. et al. Integrated genome-wide chromatin occupancy and expression analyses identify key myeloid pro-differentiation transcription factors repressed by Myb. Nucleic Acids Res. 39, 4664–4679 (2011).
Waltzer, L. & Bienz, M. Drosophila CBP represses the transcription factor TCF to antagonize Wingless signalling. Nature 395, 521–525 (1998).
Holmqvist, P. H. & Mannervik, M. Genomic occupancy of the transcriptional co-activators p300 and CBP. Transcription 4, 18–23 (2013).
Kasper, L. H., Qu, C., Obenauer, J. C., McGoldrick, D. J. & Brindle, P. K. Genome-wide and single-cell analyses reveal a context dependent relationship between CBP recruitment and gene expression. Nucleic Acids Res. 42, 11363–11382 (2014).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Thompson, P. R. et al. Regulation of the p300 HAT domain via a novel activation loop. Nat. Struct. Mol. Biol. 11, 308–315 (2004).
Qin, B. Y. et al. Crystal structure of IRF-3 in complex with CBP. Structure 13, 1269–1277 (2005).
Larabi, A. et al. Crystal structure and mechanism of activation of TANK-binding kinase 1. Cell Rep. 3, 734–746 (2013).
Levy, D. E. & Darnell, J. E., Jr. Stats: transcriptional control and biological impact. Nat. Rev. Mol. Cell Biol. 3, 651–662 (2002).
Shuai, K., Stark, G. R., Kerr, I. M. & Darnell, J. E., Jr. A single phosphotyrosine residue of Stat91 required for gene activation by interferon-gamma. Science 261, 1744–1746 (1993).
Chen, X. et al. Crystal structure of a tyrosine phosphorylated STAT-1 dimer bound to DNA. Cell 93, 827–839 (1998).
Wojciak, J. M., Martinez-Yamout, M. A., Dyson, H. J. & Wright, P. E. Structural basis for recruitment of CBP/p300 coactivators by STAT1 and STAT2 transactivation domains. EMBO J. 28, 948–958 (2009).
Darnell, J. E., Jr. STATs and gene regulation. Science 277, 1630–1635 (1997).
Delvecchio, M., Gaucher, J., Aguilar-Gurrieri, C., Ortega, E. & Panne, D. Structure of the p300 catalytic core and implications for chromatin targeting and HAT regulation. Nat. Struct. Mol. Biol. 20, 1040–1046 (2013).
Park, S. et al. Role of the CBP catalytic core in intramolecular SUMOylation and control of histone H3 acetylation. Proc. Natl Acad. Sci. USA 114, E5335–E5342 (2017).
Lasko, L. M. et al. Discovery of a selective catalytic p300/CBP inhibitor that targets lineage-specific tumours. Nature 550, 128–132 (2017).
Karanam, B., Jiang, L., Wang, L., Kelleher, N. L. & Cole, P. A. Kinetic and mass spectrometric analysis of p300 histone acetyltransferase domain autoacetylation. J. Biol. Chem. 281, 40292–40301 (2006).
Karanam, B. et al. Multiple roles for acetylation in the interaction of p300 HAT with ATF-2. Biochemistry 46, 8207–8216 (2007).
Vitalis, A. & Pappu, R. V. ABSINTH: a new continuum solvation model for simulations of polypeptides in aqueous solutions. J. Comput. Chem. 30, 673–699 (2009).
Liu, X. et al. The structural basis of protein acetylation by the p300/CBP transcriptional coactivator. Nature 451, 846–850 (2008).
Soutoglou, E. et al. Transcription factor-dependent regulation of CBP and P/CAF histone acetyltransferase activity. EMBO J. 20, 1984–1992 (2001).
Bose, D. A. et al. RNA binding to CBP stimulates histone acetylation and transcription. Cell 168, 135–149.e122 (2017).
Matt, T., Martinez-Yamout, M. A., Dyson, H. J. & Wright, P. E. The CBP/p300 TAZ1 domain in its native state is not a binding partner of MDM2. Biochem. J. 381, 685–691 (2004).
Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 17, 487–500 (2016).
Ptashne, M. Epigenetics: core misconcept. Proc. Natl Acad. Sci. USA 110, 7101–7103 (2013).
Rando, O. J. Combinatorial complexity in chromatin structure and function: revisiting the histone code. Curr. Opin. Genet. Dev. 22, 148–155 (2012).
Henikoff, S. & Shilatifard, A. Histone modification: cause or cog? Trends Genet. 27, 389–396 (2011).
Nguyen, U. T. et al. Accelerated chromatin biochemistry using DNA-barcoded nucleosome libraries. Nat. Methods 11, 834–840 (2014).
Rack, J. G. M. et al. The PHD finger of p300 influences its ability to acetylate histone and non-histone targets. J. Mol. Biol. 426, 3960–3972 (2014).
Kimbrel, E. A. et al. Systematic in vivo structure–function analysis of p300 in hematopoiesis. Blood 114, 4804–4812 (2009).
Picaud, S. et al. Generation of a selective small molecule inhibitor of the CBP/p300 bromodomain for leukemia therapy. Cancer Res. 75, 5106–5119 (2015).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Coleman, R. T. & Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of a Drosophila HOX gene. Science 356, eaai8236 (2017).
Laprell, F., Finkl, K. & Müller, J. Propagation of polycomb-repressed chromatin requires sequence-specific recruitment to DNA. Science 356, 85–88 (2017).
Wang, X. & Moazed, D. DNA sequence-dependent epigenetic inheritance of gene silencing and histone H3K9 methylation. Science 356, 88–91 (2017).
Luger, K., Rechsteiner, T. J. & Richmond, T. J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999).
McGibbon, R. T. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Holehouse, A. S., Garai, K., Lyle, N., Vitalis, A. & Pappu, R. V. Quantitative assessments of the distinct contributions of polypeptide backbone amides versus side chain groups to chain expansion via chemical denaturation. J. Am. Chem. Soc. 137, 2984–2995 (2015).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Kaczmarska, Z. et al. Structure of p300 in complex with acyl-CoA variants. Nat. Chem. Biol. 13, 21–29 (2017).
Capes-Davis, A. et al. Match criteria for human cell line authentication: where do we draw the line? Int. J. Cancer 132, 2510–2519 (2013).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
This work was supported by grant 16-0280 from Worldwide Cancer Research. E.O. was supported by an EMBL Interdisciplinary Postdoctoral (EIPOD) fellowship. S.R. was supported by the Fondation ARC pour la recherche sur le Cancer and by the Fondation FINOVI. A.S.H. is a postdoctoral fellow in the laboratory of R.V. Pappu at Washington University in St. Louis. The computational work was supported by the Human Frontiers Science Program (grant RGP0034/2017 to R.V. Pappu) and the St Jude Collaborative Research Consortium on Membraneless Organelles (to R.V. Pappu). We thank the staff at the European Synchrotron Radiation Facility (ESRF) beamlines ID29; L. Signor for mass spectroscopy analysis; R. Vance for the plasmid encoding GST-STING; and P. Cole for the A-485 inhibitor. S.K. and D.P. were supported by ANR Episperm3 program. S.K. received additional support from Fondation ARC Canc’air project (RAC16042CLA), Plan Cancer (CH7-INS15B66 and ASC16012CSA) and the Université Grenoble Alpes ANR-15-IDEX-02 LIFE and IDEX SYMER.
Reviewer information
Nature thanks L. Chen, V. Hilser and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
E.O. designed and performed most experiments, analysed and validated the data and revised the draft with assistance from S.R., Z.I., N.H. and J.G. A.S.H. performed computational modelling and revised the draft. S.K. provided supervision, funding acquisition and commented on the draft. D.P. was involved in conceptualization, supervision, project administration, funding acquisition and wrote the original and revised drafts.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The effect of IRF3 or STAT1 activation and oligomerization on p300 autoacetylation.
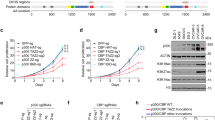
a, The domain structure of IRF3. The truncation construct used is shown at the bottom. b, Size-exclusion chromatography of IRF3 variants. Red, unphosphorylated IRF3; blue, phosphorylated pIRF3; green, C-terminally truncated IRF3ΔC. Representative data of three independent experiments are shown. c, A constant amount of p300s (2 μM) was incubated alone or in the presence of C-terminally truncated IRF3ΔC (2 μM) for the indicated time points. Samples were analysed by SDS–PAGE followed by Coomassie staining and autoradiography. d, Progress curves of HAT scintillation proximity assay. Histone H4 substrate acetylation in the presence (green) or absence (black) of pIRF3 and varying concentrations of [3H]acetyl-CoA. The degree of histone H4 substrate acetylation at different time points and the initial velocity (cpm min−1) at the indicated acetyl-CoA concentrations were determined and plotted in Fig. 1e. Three independent experiments were performed and the mean value and error bars representing the standard deviation are shown. e, The domain structure of STAT1. The truncation constructs used are shown at the bottom, and the Tyr701 phosphorylation site is indicated. f, Uncropped images of SDS–PAGE gels shown in Fig. 1d. The 14C autoacetylation signal of p300s is shown at the bottom. g, Size-exclusion chromatography of STAT1 variants. Black, STAT1ΔNC; green, STAT1ΔN; red, Y701-phosphorylated pSTAT1ΔNC; blue, Y701-phosphorylated pSTAT1ΔN. h, SDS–PAGE analysis of STAT1 variants and analysis by western blotting. Top, Coomassie staining of SDS–PAGE gel; middle, PonceauS staining; bottom, western blot using anti-Phospho-Stat1 (Tyr701). Representative data of three independent experiments are shown. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Crystal packing of the p300 core molecule.
a, There are four p300 molecules (monomers I–IV) in the asymmetric crystallographic unit. The four molecules show an antiparallel arrangement of the BRP-HAT domains. As a result, HAT domains from monomers I and II are closely apposed. Monomers III and IV engage monomer IVsym and monomer IIIsym, respectively, of a neighbouring crystallographic unit, showing that all promoters are in a AIL loop-swap conformation. Black arrows indicate the direction of the AIL. The disordered segment of the AIL is shown as a black dotted line. b, c, Electron density of the AIL. 2Fo − Fc (b) and Fo − Fc (c) difference density omit maps contoured at 0.8 and 2.0 r.m.s.d., respectively. Coloured as in Fig. 3.
Extended Data Fig. 3 Structural analysis of the RING domains.
a, Superposition of the four p300 molecules (monomers I–IV) in the asymmetric crystallographic unit. Whereas the bromodomains (Bd), PHD and HAT domains superpose with a r.m.s.d. of approximately 0.9 Å, the RING domains adopt multiple conformations. b, 2Fo − Fc (blue mesh) and anomalous difference Fourier maps (orange mesh) for the four RING domains contoured around 1σ and 2.5σ, respectively.
Extended Data Fig. 4 Regulation of HAT activity by flanking domains.
a, The domain structure of p300. Sequence conservation of the AIL is shown using WebLogo54. The constructs used are shown. b, Analysis of in vitro expression of the indicated p300 variants. Purified proteins were analysed for autoacetylation by immunoblotting with anti-p300(K1499ac) antibody (left), anti-Flag antibody (middle) and Coomassie staining (right). Representative data of three independent experiments are shown. c, Representative mass spectrometric analysis of BRP_HAT_ZZ_ΔΑΙL after in vitro expression (red) and after SIRT2 mediated deacetylation (black).
Extended Data Fig. 5 Regulation of HAT activity by flanking domains.
a, The AIL contributes to histone substrate acetylation of activated p300. The details of the constructs used are indicated in Extended Data Fig. 4. Defined amounts of p300 variants were incubated with acetyl-CoA and the indicated histones before SDS–PAGE analysis, followed by Coomassie staining and western blotting with the indicated antibodies. b, The indicated amounts of purified p300s variants were incubated with histone octamers as in a, followed by SDS–PAGE and immunoblot analysis with the indicated antibodies. Anti-Kac, pan-acetyl-lysine antibody. Representative data of three independent experiments are shown. c, Crystal structure of the H4(K12ac/K16ac) peptide bound to the BΔRP module containing an in-frame RING deletion. Amino acid residues 1169–1241 were replaced by a single glycine residue. The deletion removes the RING domain (black arrow) and does not adversely affect the structural integrity of the BΔRP module. d, e, Indicated variants of p300 were co-expressed with p53 in H1299 cells and analysed by immunofluorescence with the indicated antibodies (d) or by western blotting (e). Representative data of three independent experiments are shown. Scale bars, 10 μm.
Extended Data Fig. 6 Autoacetylation changes the hydrodynamic properties of p300.
a, Simulations of the AIL in the context of the loop-swapped dimer. Left, cartoon of the trajectory of the AIL (dashed line). Right, representative conformations with the AIL Cα backbone atoms are coloured according to charge. b, SEC–MALLS analysis of deacetylated (blue) and acetylated (yellow) p300 core. Note the decrease in elution volume upon acetylation. c, SEC-MALLS analysis of deacetylated (blue), acetylated (red) BRP_HAT_CH3 and deacetylated (black) and acetylated (green) BRP_HAT_CH3 ΔAIL. There is no increase in elution volume upon acetylation of the ΔAIL construct. d, Comparison of acetylated and deacetylated BRP_HAT and BRP_HAT_CH3. The deacetylated BRP_HAT (green) and deacetylated BRP_HAT_CH3 (blue) elute at the same position, which is indicative of a similar hydrodynamic radius. The acetylated BRP_HAT (yellow) and BRP_HAT_CH3 (red) elute at a larger elution volume. The normalized refractive index is plotted as a function of elution volume from an S200 column coupled to a MALLS detector. Calculated molecular masses are plotted as a function of volume for each eluted peak. The experiment was carried out at least three times with similar results. One representative example of each sample is shown. e, Mass spectrometry analysis (electrospray ionization) of the BRP_HAT before (blue) and after (yellow) autoacetylation. The molecular mass and the number of acetylation events are indicated. f, Mass spectrometry analysis of BRP_HAT_CH3 before (blue) and after (red) autoacetylation. g, Mass spectrometry analysis of BRP_HAT_CH3_ΔΑΙL before (black) and after (green) autoacetylation.
Extended Data Fig. 7 Molecular model and controls showing that p300 acetyltransferase activity is not stimulated by eRNA.
a, p300 is maintained in the inactive state by deacetylases such as SIRT2. IRF3 is autoinhibited by a C-terminal segment in the IAD domain. b, TBK1 phosphorylation activates and dimerizes IRF3. The activated IRF3 dimer engages the IBID domain of p300. c, Recruitment of two molecules of p300 results in trans-autoacetylation in the AIL loop and HAT activation. d, Activated p300 can acetylate chromatin and engage acetylated substrates via the bromodomain. e, A constant amount of p300s (2 μM) was incubated in [14C]acetyl-CoA alone or in the presence of 2 μM Klf6 eRNA for the indicated time points. Samples were analysed by SDS–PAGE followed by Coomassie staining (top) and autoradiography (bottom). f, As in e but in the presence of 0.5 mM EDTA. The experiment was carried out at least twice with consistency. One representative example is shown. g, Quality control of Klf6 RNA. 3 μg Klf6 was deposited on a 1% agarose gel or a 14% 6 M urea PAGE gel and detected by SYBR Safe stain. M, 100-bp DNA ladder.
Supplementary information
Supplementary Information
This file contains the uncropped gels.
Video 1: Simulations of the deacetylated AIL.
All backbone and side chain dihedral angles in the AIL were fully sampled using all-atom Monte Carlo Simulations. Normalized distances between pairs of amino acids of the AIL and the p300 core were plotted in Fig. 5a.
Video 2: Simulations of the acetylated AIL.
Distances between pairs of amino acids of the AIL and the p300 core were plotted in Fig. 5b. After acetylation, lysine-mediated electrostatic interactions are lost.
Rights and permissions
About this article
Cite this article
Ortega, E., Rengachari, S., Ibrahim, Z. et al. Transcription factor dimerization activates the p300 acetyltransferase. Nature 562, 538–544 (2018). https://doi.org/10.1038/s41586-018-0621-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0621-1
Keywords
This article is cited by
-
Tissue-specific RNA Polymerase II promoter-proximal pause release and burst kinetics in a Drosophila embryonic patterning network
Genome Biology (2024)
-
The cell biology of HIV-1 latency and rebound
Retrovirology (2024)
-
Tamoxifen exerts anti-peritoneal fibrosis effects by inhibiting H19-activated VEGFA transcription
Journal of Translational Medicine (2023)
-
Mild internet use is associated with epigenetic alterations of key neurotransmission genes in salivary DNA of young university students
Scientific Reports (2023)
-
Bromodomain and extraterminal (BET) proteins: biological functions, diseases, and targeted therapy
Signal Transduction and Targeted Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.