Abstract
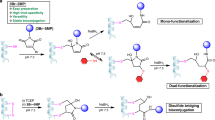
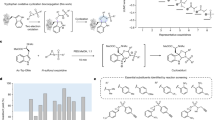
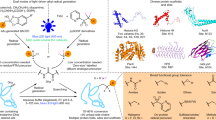
Nature has a remarkable ability to carry out site-selective post-translational modification of proteins, therefore enabling a marked increase in their functional diversity1. Inspired by this, chemical tools have been developed for the synthetic manipulation of protein structure and function, and have become essential to the continued advancement of chemical biology, molecular biology and medicine. However, the number of chemical transformations that are suitable for effective protein functionalization is limited, because the stringent demands inherent to biological systems preclude the applicability of many potential processes2. These chemical transformations often need to be selective at a single site on a protein, proceed with very fast reaction rates, operate under biologically ambient conditions and should provide homogeneous products with near-perfect conversion2,3,4,5,6,7. Although many bioconjugation methods exist at cysteine, lysine and tyrosine, a method targeting a less-explored amino acid would considerably expand the protein functionalization toolbox. Here we report the development of a multifaceted approach to protein functionalization based on chemoselective labelling at methionine residues. By exploiting the electrophilic reactivity of a bespoke hypervalent iodine reagent, the S-Me group in the side chain of methionine can be targeted. The bioconjugation reaction is fast, selective, operates at low-micromolar concentrations and is complementary to existing bioconjugation strategies. Moreover, it produces a protein conjugate that is itself a high-energy intermediate with reactive properties and can serve as a platform for the development of secondary, visible-light-mediated bioorthogonal protein functionalization processes. The merger of these approaches provides a versatile platform for the development of distinct transformations that deliver information-rich protein conjugates directly from the native biomacromolecules.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available within the paper and its Supplementary Information. Raw data are available from the corresponding author on reasonable request.
References
Walsh, C. T., Garneau-Tsodikova, S. & Gatto, G. J. Jr. Protein posttranslational modifications: the chemistry of proteome diversifications. Angew. Chem. Int. Ed. 44, 7342–7372 (2005).
Sletten, E. M. & Bertozzi, C. R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Spicer, C. D. & Davis, B. G. Selective chemical protein modification. Nat. Commun. 5, 4740 (2014).
Koniev, O. & Wagner, A. Developments and recent advancements in the field of endogenous amino acid selective bond forming reactions for bioconjugation. Chem. Soc. Rev. 44, 5495–5551 (2015).
Dawson, P. E. & Kent, S. B. H. Synthesis of native proteins by chemical ligation. Annu. Rev. Biochem. 69, 923–960 (2000).
Lang, K. & Chin, J. W. Cellular incorporation of unnatural amino acids and bioorthogonal labeling of proteins. Chem. Rev. 114, 4764–4806 (2014).
Wang, L., Xie, J. & Schultz, P. G. Expanding the genetic code. Annu. Rev. Biophys. Biomol. Struct. 35, 225–249 (2006).
Vinogradova, E. V., Zhang, C., Spokoyny, A. M., Pentelute, B. L. & Buchwald, S. L. Organometallic palladium reagents for cysteine bioconjugation. Nature 526, 687–691 (2015).
Wright, T. H. et al. Posttranslational mutagenesis: a chemical strategy for exploring protein side-chain diversity. Science 354, aag1465 (2016).
Yang, A. et al. A chemical biology route to site-specific authentic protein modifications. Science 354, 623–626 (2016).
Abegg, D. et al. Proteome-wide profiling of targets of cysteine reactive small molecules by using ethynyl benziodoxolone reagents. Angew. Chem. Int. Ed. 54, 10852–10857 (2015).
Levine, R. L., Moskovitz, J. & Stadtman, E. R. Oxidation of methionine in proteins: roles in antioxidant defense and cellular regulation. IUBMB Life 50, 301–307 (2000).
Cowie, D. B., Cohen, G. N., Bolton, E. T. & De Robichon-Szulmajster, H. Amino acid analog incorporation into bacterial proteins. Biochim. Biophys. Acta 34, 39–46 (1959).
Lin, S. et al. Redox-based reagents for chemoselective methionine bioconjugation. Science 355, 597–602 (2017).
Gross, E. & Witkop, B. Nonenzymatic cleavage of peptide bonds: the methionine residues in bovine pancreatic ribonuclease. J. Biol. Chem. 237, 1856–1860 (1962).
Gundlach, H. G., Stein, W. H. & Moore, S. The nature of the amino acid residues involved in the inactivation of ribonuclease by iodoacetate. J. Biol. Chem. 234, 1754–1760 (1959).
Vithayathil, P. J. & Richards, F. M. Modification of the methionine residue in the peptide component of ribonuclease-S. J. Biol. Chem. 235, 2343–2351 (1960).
Kramer, J. R. & Deming, T. J. Preparation of multifunctional and multireactive polypeptides via methionine alkylation. Biomacromolecules 13, 1719–1723 (2012).
Kramer, J. R. & Deming, T. J. Reversible chemoselective tagging and functionalization of methionine containing peptides. Chem. Commun. 49, 5144–5146 (2013).
Stang, P. J. & Zhdankin, V. V. Organic polyvalent iodine compounds. Chem. Rev. 96, 1123–1178 (1996).
Weiss, R., Seubert, J. & Hampel, F. α-Aryliodonio diazo compounds: SN reactions at the α-C atom as a novel reaction type for diazo compounds. Angew. Chem. Int. Edn Engl. 33, 1952–1953 (1994).
Schnaars, C., Hennum, M. & Bonge-Hansen, T. Nucleophilic halogenations of diazo compounds, a complementary principle for the synthesis of halodiazo compounds: experimental and theoretical studies. J. Org. Chem. 78, 7488–7497 (2013).
Kim, Y. et al. Efficient site-specific labeling of proteins via cysteines. Bioconjug. Chem. 19, 786–791 (2008).
Mülhberg, M. et al. Orthogonal dual-modification of proteins for the engineering of multivalent protein scaffolds. Beilstein J. Org. Chem. 11, 784–791 (2015).
Staudinger, H. & Lüscher, G. Über darstellung und reaktionen von phosphazinen. Helv. Chim. Acta 5, 75–86 (1922).
Prier, C. K., Rankic, D. A. & MacMillan, D. W. C. Visible light photoredox catalysis with transition metal complexes: applications in organic synthesis. Chem. Rev. 113, 5322–5363 (2013).
Chen, Y., Kamlet, A. S., Steinman, J. B. & Liu, D. R. A biomolecule-compatible visible-light-induced azide reduction from a DNA-encoded reaction-discovery system. Nat. Chem. 3, 146–153 (2011).
Huang, W. & Cheng, X. Hantzsch esters as multifunctional reagents in visible-light photoredox catalysis. Synlett 28, 148–158 (2017).
Fukuzumi, S., Hironaka, K. & Tanaka, T. Photoreduction of alkyl halides by an NADH model compound. An electron-transfer chain mechanism. J. Am. Chem. Soc. 105, 4722–4727 (1983).
Hedstrand, D. M., Kruizinga, W. H. & Kellog, R. M. Light induced and dye accelerated reductions of phenacyl onium salts by 1,4-dihydropyridines. Tetrahedron Lett. 19, 1255–1258 (1978).
Krause, G., Lundström, J., Barea, J. L., Pueyo de la Cuesta, C. & Holmgren, A. Mimicking the active site of protein disulfide-isomerase by substitution of proline 34 in Escherichia coli thioredoxin. J. Biol. Chem. 266, 9494–9500 (1991).
Acknowledgements
We thank M. Nappi and C. Guerot for advice and useful discussions. We thank the Marie Curie Actions program (M.T.T. and M.G.S.), AstraZeneca and EPRSC (J.E.N.), and the European Research Council (ERC-SRG-259711), EPSRC (EP/100548X/1) and the Royal Society (Wolfson Merit Award) for fellowships (M.J.G.). We are grateful to J. Chin, N. Huguen, M. Skehel, H. Lewis and M. Edgeworth for assistance with protein purification and mass spectrometry experiments.
Reviewer information
Nature thanks A. Spokoyny and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.J.G., M.T.T., J.E.N. and M.G.S. conceived the project and designed the experiments. M.J.G., M.T.T., J.E.N. and M.G.S. performed and analysed the experiments. M.J.G., M.T.T. and J.E.N. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
This file contains Supplementary Text, which includes Supplementary Figs. S1–S83 and Supplementary Tables S1–S8.
Rights and permissions
About this article
Cite this article
Taylor, M.T., Nelson, J.E., Suero, M.G. et al. A protein functionalization platform based on selective reactions at methionine residues. Nature 562, 563–568 (2018). https://doi.org/10.1038/s41586-018-0608-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0608-y
Keywords
This article is cited by
-
Non-symmetric stapling of native peptides
Nature Reviews Chemistry (2024)
-
Ring expansion of indene by photoredox-enabled functionalized carbon-atom insertion
Nature Catalysis (2024)
-
Selenium chemistry for spatio-selective peptide and protein functionalization
Nature Reviews Chemistry (2024)
-
Chemoselective umpolung of thiols to episulfoniums for cysteine bioconjugation
Nature Chemistry (2024)
-
Recent advances in chemical protein synthesis: method developments and biological applications
Science China Chemistry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.