Abstract
The earliest blood vessels in mammalian embryos are formed when endothelial cells differentiate from angioblasts and coalesce into tubular networks. Thereafter, the endothelium is thought to expand solely by proliferation of pre-existing endothelial cells. Here we show that a complementary source of endothelial cells is recruited into pre-existing vasculature after differentiation from the earliest precursors of erythrocytes, megakaryocytes and macrophages, the erythro-myeloid progenitors (EMPs) that are born in the yolk sac. A first wave of EMPs contributes endothelial cells to the yolk sac endothelium, and a second wave of EMPs colonizes the embryo and contributes endothelial cells to intraembryonic endothelium in multiple organs, where they persist into adulthood. By demonstrating that EMPs constitute a hitherto unrecognized source of endothelial cells, we reveal that embryonic blood vascular endothelium expands in a dual mechanism that involves both the proliferation of pre-existing endothelial cells and the incorporation of endothelial cells derived from haematopoietic precursors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequence data used in this study have been deposited in the NCBI Gene Expression Omnibus database (accession number GSE117978) and are listed in the Source Data for Fig. 6.
References
Potente, M., Gerhardt, H. & Carmeliet, P. Basic and therapeutic aspects of angiogenesis. Cell 146, 873–887 (2011).
Hirschi, K. K., Ingram, D. A. & Yoder, M. C. Assessing identity, phenotype, and fate of endothelial progenitor cells. Arterioscler. Thromb. Vasc. Biol. 28, 1584–1595 (2008).
Pollard, J. W. Trophic macrophages in development and disease. Nat. Rev. Immunol. 9, 259–270 (2009).
Fantin, A. et al. Tissue macrophages act as cellular chaperones for vascular anastomosis downstream of VEGF-mediated endothelial tip cell induction. Blood 116, 829–840 (2010).
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Förster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999).
de Boer, J. et al. Transgenic mice with hematopoietic and lymphoid specific expression of Cre. Eur. J. Immunol. 33, 314–325 (2003).
Hoeffel, G. et al. C-Myb+ erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678 (2015).
Frame, J. M., McGrath, K. E. & Palis, J. Erythro-myeloid progenitors: “definitive” hematopoiesis in the conceptus prior to the emergence of hematopoietic stem cells. Blood Cells Mol. Dis. 51, 220–225 (2013).
Mass, E. et al. Specification of tissue-resident macrophages during organogenesis. Science 353, aaf4238 (2016).
Gomez Perdiguero, E. et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature 518, 547–551 (2015).
McGrath, K. E. et al. Distinct sources of hematopoietic progenitors emerge before HSCs and provide functional blood cells in the mammalian embryo. Cell Rep. 11, 1892–1904 (2015).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Ginhoux, F. & Guilliams, M. Tissue-resident macrophage ontogeny and homeostasis. Immunity 44, 439–449 (2016).
Hoeffel, G. & Ginhoux, F. Fetal monocytes and the origins of tissue-resident macrophages. Cell. Immunol. 330, 5–15 (2018).
Lux, C. T. et al. All primitive and definitive hematopoietic progenitor cells emerging before E10 in the mouse embryo are products of the yolk sac. Blood 111, 3435–3438 (2008).
Fantin, A. et al. NRP1 acts cell autonomously in endothelium to promote tip cell function during sprouting angiogenesis. Blood 121, 2352–2362 (2013).
Stefater, J. A., III et al. Regulation of angiogenesis by a non-canonical Wnt–Flt1 pathway in myeloid cells. Nature 474, 511–515 (2011).
Deng, L. et al. A novel mouse model of inflammatory bowel disease links mammalian target of rapamycin-dependent hyperproliferation of colonic epithelium to inflammation-associated tumorigenesis. Am. J. Pathol. 176, 952–967 (2010).
Qian, B. Z. et al. CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature 475, 222–225 (2011).
Sasmono, R. T. et al. A macrophage colony-stimulating factor receptor-green fluorescent protein transgene is expressed throughout the mononuclear phagocyte system of the mouse. Blood 101, 1155–1163 (2003).
Burnett, S. H. et al. Conditional macrophage ablation in transgenic mice expressing a Fas-based suicide gene. J. Leukoc. Biol. 75, 612–623 (2004).
Tam, S. J. et al. Death receptors DR6 and TROY regulate brain vascular development. Dev. Cell 22, 403–417 (2012).
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013).
Goldie, L. C., Lucitti, J. L., Dickinson, M. E. & Hirschi, K. K. Cell signaling directing the formation and function of hemogenic endothelium during murine embryogenesis. Blood 112, 3194–3204 (2008).
Wilson, C. H. et al. The kinetics of ER fusion protein activation in vivo. Oncogene 33, 4877–4880 (2014).
Palis, J., Robertson, S., Kennedy, M., Wall, C. & Keller, G. Development of erythroid and myeloid progenitors in the yolk sac and embryo proper of the mouse. Development 126, 5073–5084 (1999).
Alharbi, R. A., Pettengell, R., Pandha, H. S. & Morgan, R. The role of HOX genes in normal hematopoiesis and acute leukemia. Leukemia 27, 1000–1008 (2013).
Toshner, M. et al. Transcript analysis reveals a specific HOX signature associated with positional identity of human endothelial cells. PLoS ONE 9, e91334 (2014).
Rössig, L. et al. Histone deacetylase activity is essential for the expression of HoxA9 and for endothelial commitment of progenitor cells. J. Exp. Med. 201, 1825–1835 (2005).
Browning, A. C. et al. Comparative gene expression profiling of human umbilical vein endothelial cells and ocular vascular endothelial cells. Br. J. Ophthalmol. 96, 128–132 (2012).
Nonaka, H., Tanaka, M., Suzuki, K. & Miyajima, A. Development of murine hepatic sinusoidal endothelial cells characterized by the expression of hyaluronan receptors. Dev. Dyn. 236, 2258–2267 (2007).
Majesky, M. W. Developmental basis of vascular smooth muscle diversity. Arterioscler. Thromb. Vasc. Biol. 27, 1248–1258 (2007).
Liu, C. et al. Macrophages mediate the repair of brain vascular rupture through direct physical adhesion and mechanical traction. Immunity 44, 1162–1176 (2016).
Goldman, O. et al. Endoderm generates endothelial cells during liver development. Stem Cell Reports 3, 556–565 (2014).
Matsumoto, K., Yoshitomi, H., Rossant, J. & Zaret, K. S. Liver organogenesis promoted by endothelial cells prior to vascular function. Science 294, 559–563 (2001).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Kawamoto, S. et al. A novel reporter mouse strain that expresses enhanced green fluorescent protein upon Cre-mediated recombination. FEBS Lett. 470, 263–268 (2000).
McKercher, S. R. et al. Targeted disruption of the PU.1 gene results in multiple hematopoietic abnormalities. EMBO J. 15, 5647–5658 (1996).
Scott, E. W., Simon, M. C., Anastasi, J. & Singh, H. Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science 265, 1573–1577 (1994).
Yamazaki, T. et al. Tissue myeloid progenitors differentiate into pericytes through TGF-β signaling in developing skin vasculature. Cell Rep. 18, 2991–3004 (2017).
Kmita, M. et al. Early developmental arrest of mammalian limbs lacking HoxA/HoxD gene function. Nature 435, 1113–1116 (2005).
Klein, S. et al. Interstitial cells of Cajal integrate excitatory and inhibitory neurotransmission with intestinal slow-wave activity. Nat. Commun. 4, 1630 (2013).
Zarkada, G., Heinolainen, K., Makinen, T., Kubota, Y. & Alitalo, K. VEGFR3 does not sustain retinal angiogenesis without VEGFR2. Proc. Natl Acad. Sci. USA 112, 761–766 (2015).
Yoshida, H. et al. The murine mutation osteopetrosis is in the coding region of the macrophage colony stimulating factor gene. Nature 345, 442–444 (1990).
Fantin, A., Vieira, J. M., Plein, A., Maden, C. H. & Ruhrberg, C. The embryonic mouse hindbrain as a qualitative and quantitative model for studying the molecular and cellular mechanisms of angiogenesis. Nat. Protoc. 8, 418–429 (2013).
Gory-Fauré, S. et al. Role of vascular endothelial-cadherin in vascular morphogenesis. Development 126, 2093–2102 (1999).
McLaughlin, F., Ludbrook, V. J., Kola, I., Campbell, C. J. & Randi, A. M. Characterisation of the tumour necrosis factor (TNF)-(alpha) response elements in the human ICAM-2 promoter. J. Cell Sci. 112, 4695–4703 (1999).
Morgan, S. M., Samulowitz, U., Darley, L., Simmons, D. L. & Vestweber, D. Biochemical characterization and molecular cloning of a novel endothelial-specific sialomucin. Blood 93, 165–175 (1999).
Albelda, S. M., Muller, W. A., Buck, C. A. & Newman, P. J. Molecular and cellular properties of PECAM-1 (endoCAM/CD31): a novel vascular cell-cell adhesion molecule. J. Cell Biol. 114, 1059–1068 (1991).
Shalaby, F. et al. Failure of blood-island formation and vasculogenesis in Flk-1-deficient mice. Nature 376, 62–66 (1995).
Austyn, J. M. & Gordon, S. F4/80, a monoclonal antibody directed specifically against the mouse macrophage. Eur. J. Immunol. 11, 805–815 (1981).
Ohsawa, K., Imai, Y., Sasaki, Y. & Kohsaka, S. Microglia/macrophage-specific protein Iba1 binds to fimbrin and enhances its actin-bundling activity. J. Neurochem. 88, 844–856 (2004).
Ozerdem, U., Grako, K. A., Dahlin-Huppe, K., Monosov, E. & Stallcup, W. B. NG2 proteoglycan is expressed exclusively by mural cells during vascular morphogenesis. Dev. Dyn. 222, 218–227 (2001).
Goodell, M. A., Brose, K., Paradis, G., Conner, A. S. & Mulligan, R. C. Isolation and functional properties of murine hematopoietic stem cells that are replicating in vivo. J. Exp. Med. 183, 1797–1806 (1996).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Varet, H., Brillet-Guéguen, L., Coppée, J. Y. & Dillies, M. A. SARTools: A DESeq2- and EdgeR-based R pipeline for comprehensive differential analysis of RNA-seq data. PLoS ONE 11, e0157022 (2016).
Acknowledgements
We thank the Biological Resources, FACS, Imaging and Genomics facilities at UCL and E. Scarpa for technical help; D. Saur, A. Mass, D. Duboule, M. Kmita and Y. Kubota for mouse strains; and M. Golding for helpful discussions. This research was supported by grants from the Wellcome Trust (095623/Z/11/Z, 101067/Z/13/Z), Medical Research Council (MR/N011511/1) and British Heart Foundation (FS/17/23/32718).
Reviewer information
Nature thanks L. Iruela-Arispe and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.P., A.F. and C.R. conceived and planned this study, analysed data and co-wrote the manuscript. L.D. performed genetic crosses and genotyping. A.P. and A.F. either performed experiments together or replicated each other’s experiments, except for the cell cycle and Hoxa studies, which were carried out by A.P and A.F., respectively. J.W.P. provided mouse strains. C.R. supervised the project. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Endothelial Csf1r-iCre-targeting is observed with different recombination reporters, and targeted ECs are distinguishable from macrophages and pericytes.
a–c, Csf1r-iCre;RosaYfp (a), Csf1r-iCre;CAG-Cat-Egfp (b) and Csf1r-iCre;RosatdTom (c) hindbrains (n = 3 each) at the indicated stages were whole-mount labelled with IB4 and for YFP (a) or GFP (b) or are shown with tdTomato fluorescence (c). In a, the white squares indicate areas that were imaged at higher magnification for Fig. 1a. The indicated single channels are also shown individually. d, Csf1r-iCre;RosatdTom E12.5 hindbrains (n = 3), whole-mount labelled for ERG and CDH5 and shown including tdTomato fluorescence to demonstrate that Csf1r-iCre targets ECs that form junctions with neighbouring non-targeted ECs. e, f, E12.5 Csf1r-iCre;RosaYfp hindbrains, labelled for YFP and the microglia marker F4/80 (e) or the pericyte marker NG2 (f) together with IB4, show that Csf1r-iCre-targeted vessel-bound cells are neither microglia nor pericytes; n = 3 each. In e, the boxed area is shown in higher magnification and as single channels adjacent to the panel. In f, a single optical y/z cross section at the position indicated with the yellow line is displayed at higher magnification with single channels. Arrowheads, microglia; arrows, ECs; double arrowheads, pericytes; curved arrow, junctional CDH5 staining; solid and clear symbols indicate the presence or absence of marker expression, respectively. Scale bars: 100 µm (a), 20 µm (b, c, e, f), 50 µm (d).
Extended Data Fig. 2 Endothelial Csf1r-iCre-targeting is not caused by endothelial Csf1r expression and occurs independently of myeloid differentiation.
a, b, Csf1r-Egfp (a) and Csf1r-iCre;RosaYfp (b) E11.5 hindbrains (n = 3 each), whole-mount labelled for CSF1R and EGFP or YFP together with IB4, show lack of Csf1r promoter activity and CSF1R protein in ECs. c, Relative Cdh5 and Csf1r expression levels in our analysis of published E14.5 brain or pooled lung/liver EC microarrays22; n = 5 each; ***P < 0.0001 (two-tailed unpaired t-test). d–g, FACS separation of tdTomato+ cells from Csf1r-iCre;RosatdTom embryos (n = 3) for gene expression analysis, including representative gating strategy to exclude dead cells and doublets in this and subsequent experiments (d) and sorting into PECAM1+CD45− ECs versus CD45+PECAM1− MCs (e). f, Representative RT–qPCR gene amplification graphs for Csf1r and Actb from tdTomato+ MCs and ECs; ΔRn, normalized reporter value for SYBR Green minus baseline instrument signals. g, Graphic representation of the fold change in RT–qPCR amplification of the indicated genes relative to Actb for both cell populations; each data point represents one embryo; *P = 0.0242, ***P < 0.0001 (two-tailed unpaired t-test). h, Csf1r-iCre;RosaYfp P0 striatum on a Pu.1+/+ versus Pu.1−/− background (n = 3 brains each), cryosectioned and labelled for YFP and F4/80 together with IB4 to show that Csf1r-iCre-targeted ECs are PU.1-independent and persist postnatally. Arrowheads, microglia; arrows, YFP+ ECs; clear arrows, YFP+ ECs that are CSF1R− and F4/80−. Scale bars, 20 µm.
Extended Data Fig. 3 Lineage tracing of yolk sac and liver EMPs.
a, b, E8.5 wild-type (a) and Pu.1−/− (b) yolk sacs on a Csf1r-iCre;RosaYfp background (n = 3 yolk sacs each), whole-mount labelled for YFP and KIT, show Csf1r-iCre-targeted KIT+ round cells corresponding to EMPs and MPs as well as Csf1r-iCre-targeted KIT− flat cells corresponding to ECs. Scale bars, 20 µm. c–f, Pregnant Csf1r-Mer-iCre-Mer;RosatdTom (c, d) and KitCreERT2;RosatdTom (e, f) dams were injected with a single tamoxifen dose on the indicated days; E12.5 yolk sacs were whole-mount labelled for the indicated markers to identify Csf1r-iCre-targeted ECs and macrophages (n = 3 yolk sacs for each genotype). Wavy arrows, EMPs; straight arrows, Csf1r-iCre-lineage-traced ECs; arrowheads, macrophages; solid and clear symbols indicate the presence or absence, respectively, of the indicated markers. Scale bars, 20 µm. g–i, Pregnant dams were injected with a single tamoxifen dose on E10.5 (g) before using the indicated markers for FACS analysis of E11.5 Csf1r-Egfp;Csf1r-Mer-iCre-Mer;RosatdTom (h) or Csf1r-Mer-iCre-Mer;RosatdTom control (i) livers (n = 4 each); the CD45highKIT− differentiated MC (blue), CD45lowKIT+ EMP/MP (pink) and CD45−KIT+ (grey) populations were gated further for Csf1r-Egfp and tdTomato. CD45−KIT+ cells were neither MCs nor EMPs, because they lacked CD45, tdTomato and EGFP.
Extended Data Fig. 4 Immunostaining controls for cultured Csf1r-iCre-targeted cells.
The indicated cell populations were FACS-isolated from E12.5 Csf1r-iCre;RosatdTom liver or blood with the indicated markers and cultured for three days in methocult (met.) on fibronectin (FN); n = 1 experiment. a, b, Adherent cells from tdTomato+ liver MC (a) and EMP/MP (b) cultures were stained for ERG and VEGFR2 (top) or with secondary antibodies only (bottom). c, Adherent cells from tdTomato+ blood EMP and MP cultures were immunostained for CSF1R together with the myeloid markers CD45 (top) or F4/80 (bottom). In the first panel in each row, the phase contrast and DAPI images were merged. In panels 2–4 in each row, immunolabelled cells were visualized together with tdTomato fluorescence, with single channels for the indicated markers shown separately in greyscale. Arrows, tdTomato+ ECs; arrowheads, tdTomato+ MCs; solid and clear symbols indicate the presence or absence, respectively, of the indicated markers. Scale bars, 20 µm.
Extended Data Fig. 5 Hoxa gene targeting with Csf1r-iCre.
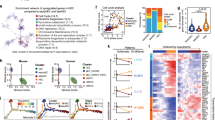
a, Schematic representation of the Hoxa gene cluster and adjacent Evx1 gene using the UCSC Genome Browser with the mouse December 2011 (GRCm38/mm10) Assembly, including position of the LoxP sites used for gene targeting. b, c, Validation of Hoxa targeting. b, FACS strategy to isolate KIT+ cells from E12.5 control (pooled Csf1r-iCre− or Csf1r-iCre+;Hoxa+/+; n = 14), Hoxa+/fl;Csf1r-iCre (n = 6) and Hoxafl/fl;Csf1r-iCre (n = 8) livers. c, qPCR analysis of Hoxa11 gene copy number relative to Evx1; mean ± s.d.; each symbol represents the value for one liver; *P = 0.0156, ***P < 0.001 (one-way ANOVA, Tukey’s multiple comparisons test). d–f, Representative FACS analysis (d) and quantification (e, f) of liver cell populations at E12.5 shows a similar number of total CD45+ and CD45+CD11b+ differentiated myeloid cells in Hoxafl/fl;Csf1r-iCre mutants (n = 7 for CD45+; n = 6 for CD45+CD11b+) versus pooled Csf1r-iCre− and Csf1r-iCre+;Hoxa+/+controls (n = 25 for CD45+, n = 17 for CD45+CD11b+); mean ± s.d. fold change in mutants compared to controls; each data point represents one liver; NS, not significant, P = 0.6519 (e) and P = 0.496 (f) (two-tailed unpaired t-test). g–i, E12.5 hindbrains of the indicated genotypes were immunolabelled to determine vascular complexity and quantify microglia. g, Schematic representation of a whole-mount embryonic hindbrain (left) and location of the hindbrain areas i–iv used for quantification (right); values for the four areas in each hindbrain were averaged to obtain the value for that hindbrain; EC quantifications are shown in Fig. 5c. h, Hindbrains were whole-mount labelled with IB4 and for RFP to visualize tdTomato and for F4/80 to visualize microglia; white boxes indicate areas shown in higher magnification in Fig. 5. i, Quantification of microglia in Hoxafl/fl;Csf1r-iCre mutants (n = 9) versus controls (n = 10, pooled Csf1r-iCre+;Hoxa+/+ and Csf1r-iCre− of any Hoxa genotype); mean ± s.d. fold change in mutant compared to control hindbrain; each data point represents one hindbrain; **P = 0.0055 (two-tailed unpaired t-test). j–l, E11.5 Csf1+/+ and Csf1+/op littermate hindbrains, whole-mount labelled for F4/80 together with IB4 (j) before quantification of microglia number (k) and vascular branchpoints as a measure of vascular complexity (l). Mean ± s.d.; each data point represents one hindbrain, n = 3 each; NS, not significant, P = 0.808, **P = 0.0012 (two-tailed unpaired t-test). Scale bars: 200 µm (h), 100 µm (j).
Extended Data Fig. 6 Csf1r-iCre-targeted ECs proliferate in vivo.
a, b, E12.5 Csf1r-iCre;RosatdTom yolk sac (a) or hindbrain (b), whole-mount stained for the proliferation marker pHH3 and VEGFR2 or for pHH3 together with IB4, respectively, and shown together with tdTomato fluorescence (n = 3 each). Areas indicated with white squares were imaged at higher magnification and are shown below the corresponding panels, with tdTomato and pHH3 channels also shown separately in greyscale. Arrows, proliferating tdTomato+pHH3+ ECs; solid and clear symbols indicate the presence or absence, respectively, of tdTomato fluorescence; wavy arrow, a tdTomato−pHH3+ neural progenitor. Scale bars: 100 µm (top), 20 µm (bottom). c–e, Cell cycle distribution of tdTomato+ and tdTomato− ECs. c, FACS strategy to isolate tdTomato+ and tdTomato−PECAM1+ ECs from E12.5 Csf1r-iCre;RosatdTom embryos (n = 3 embryos). d, Cell cycle distribution based on Hoechst 33342 fluorescence as a measure of DNA content; low and high staining intensity is observed in cells with a DNA ploidy of 2n (G0/G1 phase) or 4n (G2/M phase), respectively; intermediate staining intensity corresponds to S phase. e, Mean ± s.d. proportion of tdTomato+ and tdTomato− ECs in G1, S and G2/M based on the area of the corresponding peaks in d; NS, not significant, P > 0.9999 (two-way ANOVA, Bonferroni’s multiple comparisons test).
Extended Data Fig. 7 Validation of gene expression data from RNA-Seq and microarray studies.
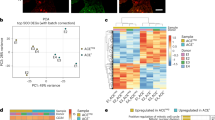
ECs were FACS-isolated from E12.5 Csf1r-iCre;RosatdTom embryos (n = 3) as in Fig. 6a to validate the RNA-seq and microarray data shown in Fig. 6d–f. Slc2a1 was analysed as a representative brain EC-enriched transcript/differentiation marker, and Mrc1 and Oit3 as representative liver EC-enriched transcripts. a, Relative transcript levels of the Gt(ROSA)26Sor (tdTomato) transcript by RNA-seq of the E12.5 tdTomato+ and tdTomato− EC populations (analysis presented in Fig. 6a–f); mean ± s.d. of normalized counts, n = 3 each; **P = 0.0085 (two-sided unpaired t-test). b, RT–qPCR analysis for the indicated genes in tdTomato+ versus tdTomato− ECs isolated from whole E12.5 embryos (n = 5) to validate genes identified by RNA-seq in Fig. 6e, f as differentially expressed. Mean ± s.d. of fold change; ***P < 0.0001 (Slc2a1), ***P = 0.0008 (Mrc1) **P = 0.0056 (Oit3) (two-sided unpaired t-test). c, RT–qPCR analysis for the indicated genes in tdTomato− ECs isolated from the E12.5 brain versus liver (n = 3 for each organ) to validate organ-specific transcript enrichment identified via microarray analysis shown in Fig. 6f. Mean ± s.d. of fold change; *P = 0.019, **P = 0.0082, ***P < 0.0001 (two-sided unpaired t-test); ND, not detectable. d, RT–qPCR analysis for the indicated genes to directly compare the expression levels of brain and liver EC differentiation markers in tdTomato+ versus tdTomato− ECs isolated from brain (n = 3) or liver (n = 5). Mean ± s.d. of fold change; NS, not significant, P = 0.9398 (liver Slc2a1), P = 0.8045 (liver Mrc1), P = 0.6327 (liver Oit3), **P = 0.0073 (brain Slc2a1) (two-sided unpaired t-test); ND, not detectable.
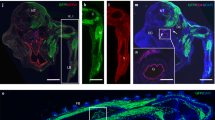
Extended Data Fig. 8 Csf1r-iCre-targeted ECs contribute to embryonic vasculature in multiple organs.
a, 20-µm cryosections of the indicated E12.5 Csf1r-iCre;RosatdTom organs (n = 3 each) were immunolabelled for the indicated EC markers together with antibodies for RFP to identify tdTomato protein (top and bottom) or are shown with tdTomato fluorescence (middle); single channels are shown in greyscale. The white boxes indicate the positions of areas shown in higher magnification in Fig. 6g; some areas selected for higher magnification are not contained entirely within the field of view, and accordingly the boxes are shown incomplete. Scale bars, 200 µm. b, Gating strategy for FACS analysis of tdTomato+ and tdTomato− ECs from E12.5 Csf1r-iCre;RosatdTom brain, lung, heart and liver versus control organs lacking iCre, using antibodies for CD11b, CD41, CD45, KIT and PECAM1; associated EC quantifications are shown in Fig. 6i. An analogous strategy was used for the quantifications shown in Fig. 6j and in Extended Data Fig. 9b.
Extended Data Fig. 9 Csf1r-iCre-targeted ECs contribute to organ vasculature in late-stage embryos and adults.
a, 20-µm cryosections of the indicated organs from E18.5 Csf1r-iCre;RosaYfp mice (n = 2 each) were immunolabelled for YFP, PECAM1 and IBA1; single channels are shown in greyscale. Arrowheads, YFP+IBA1+ macrophages; solid and empty arrows, ECs that are YFP+ and lack IBA1 expression, respectively. Scale bars, 20 µm. b, FACS analysis of dissociated cells from the indicated organs of E18.5 Csf1r-iCre;RosatdTom embryos after staining with antibodies for CD11b, CD41, CD45, KIT and PECAM1, using the gating strategy shown in Extended Data Fig. 8b; mean ± s.d., n = 5 each; ***P < 0.0001 (one-way ANOVA, Tukey’s multiple comparisons test). c, 20-µm cryosections of the indicated organs from 6-month-old adult Csf1r-iCre;RosaYfp mice (n = 3 organs each) were immunolabelled for YFP, PECAM1 and F4/80; single channels are shown in greyscale. Arrowheads and arrows as in a. Scale bars, 20 µm.
Extended Data Fig. 10 Csf1r-iCre-targeted ECs contribute to adult organ vasculature.
a, 20-µm cryosections of 3-month-old adult Csf1r-iCre;RosatdTom livers (n = 3) were immunolabelled for RFP, VEGFR2 and F4/80 or MRC1 and then counterstained with DAPI; single channels are shown in greyscale. The white box indicates an area shown in higher magnification in Fig. 6h. Scale bars, 100 µm. b, Working model for the role of EMPs in generating extra-embryonic yolk sac and intra-embryonic organ ECs alongside their known role in generating myeloid and erythrocyte/megakaryocyte lineage cells. It is not yet known whether EMP-derived and non-EMP-derived ECs have different functions to regulate normal organ physiology or pathological vascular responses in the adult.
Supplementary information
Rights and permissions
About this article
Cite this article
Plein, A., Fantin, A., Denti, L. et al. Erythro-myeloid progenitors contribute endothelial cells to blood vessels. Nature 562, 223–228 (2018). https://doi.org/10.1038/s41586-018-0552-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0552-x
Keywords
This article is cited by
-
Endothelial cells and macrophages as allies in the healthy and diseased brain
Acta Neuropathologica (2024)
-
The niche matters: origin, function and fate of CNS-associated macrophages during health and disease
Acta Neuropathologica (2024)
-
Macrophages and endothelial cells in the neurovascular unit
Acta Neuropathologica (2024)
-
Testicular macrophages are recruited during a narrow fetal time window and promote organ-specific developmental functions
Nature Communications (2023)
-
Flow-induced reprogramming of endothelial cells in atherosclerosis
Nature Reviews Cardiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.