Abstract
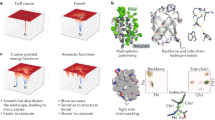
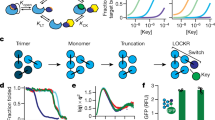
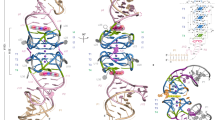
The regular arrangements of β-strands around a central axis in β-barrels and of α-helices in coiled coils contrast with the irregular tertiary structures of most globular proteins, and have fascinated structural biologists since they were first discovered. Simple parametric models have been used to design a wide range of α-helical coiled-coil structures, but to date there has been no success with β-barrels. Here we show that accurate de novo design of β-barrels requires considerable symmetry-breaking to achieve continuous hydrogen-bond connectivity and eliminate backbone strain. We then build ensembles of β-barrel backbone models with cavity shapes that match the fluorogenic compound DFHBI, and use a hierarchical grid-based search method to simultaneously optimize the rigid-body placement of DFHBI in these cavities and the identities of the surrounding amino acids to achieve high shape and chemical complementarity. The designs have high structural accuracy and bind and fluorescently activate DFHBI in vitro and in Escherichia coli, yeast and mammalian cells. This de novo design of small-molecule binding activity, using backbones custom-built to bind the ligand, should enable the design of increasingly sophisticated ligand-binding proteins, sensors and catalysts that are not limited by the backbone geometries available in known protein structures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The atomic coordinates and experimental data of BB1, b10, b11L5F_LGL, mFAP0–DFHBI, and mFAP1–DFHBI crystal structures have been deposited in the RCSB Protein Database with the accession numbers of 6D0T, 6CZJ, 6CZG, 6CZH and 6CZI respectively. All the design models, Illumina sequencing data, sequencing analysis and source data (Figs. 2, 4, Extended Data Figs. 6e, 7, 8a, h) are available at https://doi.org/10.5281/zenodo.1216229.
References
Huang, P.-S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320–327 (2016).
Marcos, E. et al. Principles for designing proteins with cavities formed by curved β sheets. Science 355, 201–206 (2017).
Tinberg, C. E. et al. Computational design of ligand-binding proteins with high affinity and selectivity. Nature 501, 212–216 (2013).
Bick, M. J. et al. Computational design of environmental sensors for the potent opioid fentanyl. eLife 6, e28909 (2017).
Dou, J. et al. Sampling and energy evaluation challenges in ligand binding protein design. Protein Sci. 26, 2426–2437 (2017).
Richardson, J. S. & Richardson, D. C. Natural β-sheet proteins use negative design to avoid edge-to-edge aggregation. Proc. Natl Acad. Sci. USA 99, 2754–2759 (2002).
Polizzi, N. F. et al. De novo design of a hyperstable non-natural protein–ligand complex with sub-Å accuracy. Nat. Chem. 9, 1157–1164 (2017).
LaLonde, J. M., Bernlohr, D. A. & Banaszak, L. J. The up-and-down beta-barrel proteins. FASEB J. 8, 1240–1247 (1994).
Flower, D. R. Structural relationship of streptavidin to the calycin protein superfamily. FEBS Lett. 333, 99–102 (1993).
Richter, A., Eggenstein, E. & Skerra, A. Anticalins: exploiting a non-Ig scaffold with hypervariable loops for the engineering of binding proteins. FEBS Lett. 588, 213–218 (2014).
Toda, M., Zhang, F. & Athukorallage, B. Elastic surface model for beta-barrels: geometric, computational, and statistical analysis. Proteins 86, 35–42 (2018).
Novotný, J., Bruccoleri, R. E. & Newell, J. Twisted hyperboloid (Strophoid) as a model of β-barrels in proteins. J. Mol. Biol. 177, 567–573 (1984).
Naveed, H., Xu, Y., Jackups, R., Jr & Liang, J. Predicting three-dimensional structures of transmembrane domains of β-barrel membrane proteins. J. Am. Chem. Soc. 134, 1775–1781 (2012).
Lasters, I., Wodak, S. J., Alard, P. & van Cutsem, E. Structural principles of parallel beta-barrels in proteins. Proc. Natl Acad. Sci. USA 85, 3338–3342 (1988).
Murzin, A. G., Lesk, A. M. & Chothia, C. Principles determining the structure of β-sheet barrels in proteins. I. A theoretical analysis. J. Mol. Biol. 236, 1369–1381 (1994).
Murzin, A. G., Lesk, A. M. & Chothia, C. Principles determining the structure of β-sheet barrels in proteins. II. The observed structures. J. Mol. Biol. 236, 1382–1400 (1994).
Salemme, F. R. Conformational and geometrical properties of β-sheets in proteins. III. Isotropically stressed configurations. J. Mol. Biol. 146, 143–156 (1981).
Minor, D. L., Jr & Kim, P. S. Measurement of the β-sheet-forming propensities of amino acids. Nature 367, 660–663 (1994).
Fujiwara, K., Ebisawa, S., Watanabe, Y., Toda, H. & Ikeguchi, M. Local sequence of protein β-strands influences twist and bend angles. Proteins 82, 1484–1493 (2014).
Lin, Y.-R. et al. Control over overall shape and size in de novo designed proteins. Proc. Natl Acad. Sci. USA 112, E5478–E5485 (2015).
Kuhlman, B. et al. Design of a novel globular protein fold with atomic-level accuracy. Science 302, 1364–1368 (2003).
Richardson, J. S., Getzoff, E. D. & Richardson, D. C. The beta bulge: a common small unit of nonrepetitive protein structure. Proc. Natl Acad. Sci. USA 75, 2574–2578 (1978).
Chan, A. W., Hutchinson, E. G., Harris, D. & Thornton, J. M. Identification, classification, and analysis of beta-bulges in proteins. Protein Sci. 2, 1574–1590 (1993).
Hemmingsen, J. M., Gernert, K. M., Richardson, J. S. & Richardson, D. C. The tyrosine corner: a feature of most Greek key β-barrel proteins. Protein Sci. 3, 1927–1937 (1994).
Greene, L. H., Hamada, D., Eyles, S. J. & Brew, K. Conserved signature proposed for folding in the lipocalin superfamily. FEBS Lett. 553, 39–44 (2003).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Warner, K. D. et al. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Allison, B. et al. Computational design of protein–small molecule interfaces. J. Struct. Biol. 185, 193–202 (2014).
Zanghellini, A. et al. New algorithms and an in silico benchmark for computational enzyme design. Protein Sci. 15, 2785–2794 (2006).
Rocklin, G. J. et al. Global analysis of protein folding using massively parallel design, synthesis, and testing. Science 357, 168–175 (2017).
Plamont, M.-A. et al. Small fluorescence-activating and absorption-shifting tag for tunable protein imaging in vivo. Proc. Natl Acad. Sci. USA 113, 497–502 (2016).
Meech, S. R. Excited state reactions in fluorescent proteins. Chem. Soc. Rev. 38, 2922–2934 (2009).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Gront, D., Kmiecik, S. & Kolinski, A. Backbone building from quadrilaterals: a fast and accurate algorithm for protein backbone reconstruction from alpha carbon coordinates. J. Comput. Chem. 28, 1593–1597 (2007).
Huang, P.-S. et al. RosettaRemodel: a generalized framework for flexible backbone protein design. PLoS ONE 6, e24109 (2011).
Park, H., DiMaio, F. & Baker, D. The origin of consistent protein structure refinement from structural averaging. Structure 23, 1123–1128 (2015).
Davis, I. W. & Baker, D. RosettaLigand docking with full ligand and receptor flexibility. J. Mol. Biol. 385, 381–392 (2009).
Park, H. et al. Simultaneous optimization of biomolecular energy functions on features from small molecules and macromolecules. J. Chem. Theory Comput. 12, 6201–6212 (2016).
Mandell, D. J., Coutsias, E. A. & Kortemme, T. Sub-angstrom accuracy in protein loop reconstruction by robotics-inspired conformational sampling. Nat. Methods 6, 551–552 (2009).
Procko, E. et al. Computational design of a protein-based enzyme inhibitor. J. Mol. Biol. 425, 3563–3575 (2013).
Thyme, S. B. et al. Reprogramming homing endonuclease specificity through computational design and directed evolution. Nucleic Acids Res. 42, 2564–2576 (2014).
Chao, G. et al. Isolating and engineering human antibodies using yeast surface display. Nat. Protocols 1, 755–768 (2006).
Whitehead, T. A. et al. Optimization of affinity, specificity and function of designed influenza inhibitors using deep sequencing. Nat. Biotechnol. 30, 543–548 (2012).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Fowler, D. M., Araya, C. L., Gerard, W. & Fields, S. Enrich: software for analysis of protein function by enrichment and depletion of variants. Bioinformatics 27, 3430–3431 (2011).
Rubin, A. F. et al. A statistical framework for analyzing deep mutational scanning data. Genome Biol. 18, 150 (2017).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. in International Tables for Crystallography, Volume F, 2nd edition (eds Arnold, E. et al.) Ch. 18.11, 539–547 (Wiley, Hoboken, 2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Otwinowski, Z. & Minor, W. in Methods in Enzymology Vol. 276 (Ed. Carter, W. C.) Ch. 3, 307–326 (Academic, Cambridge, 1997).
Merkel, J. S. & Regan, L. Aromatic rescue of glycine in β sheets. Fold. Des. 3, 449–456 (1998).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Conway, P., Tyka, M. D., DiMaio, F., Konerding, D. E. & Baker, D. Relaxation of backbone bond geometry improves protein energy landscape modeling. Protein Sci. 23, 47–55 (2014).
Hauser, C. A. E. et al. Natural tri- to hexapeptides self-assemble in water to amyloid β-type fiber aggregates by unexpected α-helical intermediate structures. Proc. Natl Acad. Sci. USA 108, 1361–1366 (2011).
Acknowledgements
We thank S. R. Jaffrey and T. A. Rapoport for providing experimental materials; A. Kang, S. A. Rettie, K. Lou, D. Sahtoe, D. La, G. J. Rocklin and C. Taylor for their help with experiments and data analysis; D. Alonso, L. Goldschmidt, P. Vecchiato, T. J. Brunette, D. Kim, V. K. Mulligan and T. Linsky for computer support, and the UW Hyak supercomputer and Rosetta@Home volunteers (https://boinc.bakerlab.org) for computing resources. We thank B. Huang, B. Basanta, R. Cacho, G. Daniel, Y. Kipnis, J. Klima and other members of the Baker laboratory for discussions. A.A.V. was supported by Fulbright Commission for Belgium and Luxembourg. E.M. was supported by a Marie Curie International Outgoing Fellowship (FP7-PEOPLE-2011-IOF 298976). B.L.S. is supported by NIH grant R01 GM115545. The Berkeley Center for Structural Biology is supported by the NIH, NIGMS and HHMI. The Advanced Light Source is a DOE User Facility under Contract No. DE-AC02-05CH11231. D.B. is supported by HHMI, WRF and Open Philanthropy.
Reviewer information
Nature thanks R. Campbell and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.A.V., J.D. and D.B. designed the study. W.S. developed RIF docking methods. B.M. developed the parametric design methods, designed and characterized the proteins. A.A.V. developed the β-barrel design methods with help from P.-S.H. L.C. purified proteins, performed SEC–MALS and analysed the results. L.A.D., M.J.B., B.S. and A.A.V. determined crystal structures. J.D. developed the ligand-binding design methodology and designed and optimized the mFAPs. H.P. performed post-design model refinement and docking calculations. G.W.F. and L.A.G. performed in vivo fluorescent imaging experiments. M.Y.L carried out photophysical characterization. E.M. and S.O. provided computational scripts. L.A.D. was supervised by B.L.S.; L.A.G. and M.Y.L were supervised by J.C.V. J.D., A.A.V. and D.B. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
J.D., A.A.V. and D.B. are inventors on a U.S. provisional patent application submitted by the University of Washington that covers the described methods, sequences and applications.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Parametric design: workflow and shortcomings.
a, Schematic of the parametric approach to generate β-barrel designs. b–d, Comparison between β-barrels of type (n = 8, S = 8) (b), type (n = 8, S = 10) (c) and type (n = 8, S = 12) (d); showing an example of 2D map with residue connectivity (top), the arrangement of the Cβ atoms in the Cβ-strips (middle) and the packing pattern of the core side chains (bottom). Although the three β-barrels have the same number of strands n, the difference in shear number S translates into different overall strand staggering, barrel radii (r) and number of Cβ-strips. The number of core Cβ-strips (top, middle) results in different arrangements of side chains in the core of the barrel. e, f, The parametric designs exhibited distorted hydrogen bonds, reflected by the shear distance (defined in e) between Cα atoms of paired antiparallel β-strand residues. The shear distance in the designs deviates from the distribution observed in native β-sheet proteins (f).
Extended Data Fig. 2 Glycine kinks release strain in β-barrel backbones.
a, Fraction of retained hydrogen-bond interactions after relaxation with Rosetta (‘relax’) of uniform poly-valine backbones (white) and poly-valine backbones with a glycine in the middle of each Cβ-strip (grey). We compare disconnected strand arrangements generated with the parametric hyperboloid model (n = 225 independently generated models), the cylindric model (n = 36 independently generated models), the coiled-coil model (n = 150 independently generated models) and assembled on the basis of a 2D map (n = 144 independently generated models). For all box plots: centre line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum values; points, outliers. b, c, In poly-valine backbones (n = 189 independently generated models) relaxed with constraints to maintain hydrogen bonds between strands, several residues have unfavourable left-handed twist (c). The local strand twist is calculated on a sliding window of four residues along β-strands, as the angle between the vectors Cα1–Cα3 and Cα2–Cα4. The handedness of the twist is defined as the triple scalar product between these two vectors and the central axis of the barrel. Positive and negative values denote right-handed and left-handed twist, respectively (b). d, After relaxation (‘FastRelax’), the valine positions in the middle of each Cβ-strip remained in the β-sheet-specific ABEGO space (right); or were shifted towards the positive Φ space (E ABEGO) if mutated to glycines (bottom). e, A similar torsion angle distribution was observed for glycines in the β-strands of native β-barrels (n = 35 high-resolution crystal structures). f, In comparison with regular β-strands (top), the presence of glycine kinks (bottom) increases the local bending of the strands and creates corners in an otherwise circular barrel cross-section. g, The bending angle α is calculated on a sliding window of three residues.
Extended Data Fig. 3 Placement of β-bulges, β-turns and the tryptophan corner.
a, Change of curvature (from convex to concave) and protrusion (dashed circle) of the longest hairpin associated with the placement of a glycine kink at position 44. b, Relationship between the ‘corners’ in the β-sheet (dashed line) generated by the glycine kinks and the type and position of the β-bulges and β-turns (Supplementary Methods). Cα are shown as spheres and coloured by ABEGO type. The bottom of the barrel was defined as the side of the N and C termini. c, The type I β-turn (‘AA’ ABEGO type) is frequently found at the second position relative to a β-bulge in native proteins and was selected to connect bottom hairpins. d, This choice is further supported by the enrichment of type I (AA) turns over the canonical type I′ turn (GG) in native β-barrels (n = 35 high-resolution crystal structures). e, f, Poly-valine backbones built with β-bulges and the corresponding β-turns (n = 194 independently generated models) retain more hydrogen bonds after relaxation than backbones built without β-bulges and with canonical type I′ β-turns (n = 186 independently generated models) (e) and exhibit better-scored hydrogen bonds per β-strand residue flanking the β-turns (f). g, Superposition of tryptophan corner motifs (n = 41 high-resolution crystal structures) extracted from native β-barrels. h–j, Amino acid preference and torsional constraints derived from the set and used to model the tryptophan corner. Bounded constraints limits are shown as dashed lines.
Extended Data Fig. 4 Biochemical and structural characterizations of designs BB1–4.
a, Results of experimental characterization of the nonfunctional designs (BB1–4). Reproducibility is described in the Methods. †E value is calculated by BLAST, the non-redundant protein database. b, Far-ultraviolet CD spectra of designs BB2 and BB3 at 25 °C. c, SEC–MALS analysis showed a major monomer peak for BB1 and a major tetramer peak for BB2. d, Variants of BB1 with residues of the tryptophan corner and glycine kinks mutated to alanine were purified and sized. SEC traces are superimposed on the SEC trace of wild-type BB1 (WT). The mutations of all residues of the tryptophan corner eliminate the monomeric peak. Most of the glycine kink mutations negatively affect the monomeric species. The exceptions are Gly53 and Gly55, which are next to each other on the fourth strand. One glycine kink per strand might be sufficient to introduce enough negative twist to remove strain in the β-barrel. e–f, Deviations between BB1 design model and crystal structure. e, One of the three bottom turns of the crystal structure (grey) deviates from the design model (magenta) and forms additional crystal contacts (indicated by a dashed circle). f, Three phenylalanine side chains have different rotameric states. In the crystal structure, Phe41 interacts with Gly53 (which shows the most backbone deviation between the crystal structure and the design) to form an aromatic rescue motif53. It is likely that the discrepancies in the Phe rotamers reflect a scoring and sampling challenge to accurately capture such aromatic rescue; molecular dynamics simulation starting from the crystal structure (cyan) was also unable to recover the correct Phe41–Gly53 interaction. g, Biophysical properties (absorbance or fluorescence spectra, quantum yield and binding affinity) of mFAP1 and mFAP2 in complex with DFHBI. Mean values from three biological replicates were used for the nonlinear regression to determine the KD. The error estimates are the standard deviation from the fitting calculation. *λabs is peak absorbance wavelength, λex is peak excitation wavelength and λem is peak emission wavelength. †Absolute quantum yield is measured with an integrating sphere; relative quantum yield is measured using acridine yellow and fluorescein as the standards. ‡Previously reported value26. §Taken from previously published work54. ||Taken from previously published work31.
Extended Data Fig. 5 RIF docking grid-based search algorithm, β-barrel scaffold construction and post-design ligand-docking simulations.
a, Illustration of grid-based hierarchical search strategy in RIF docking. After generating an ensemble of interactions for the target ligand (Fig. 3), each one of the selected scaffold is docked into the fixed RIF using the grid-based hierarchical searching algorithm. This search procedure starts from coarse sampling grids to fine sampling grids in 3D space. An example 2D grid scheme is shown in the upper row, from the lowest resolution (coarse sampling, left) to the highest resolution (fine sampling, right). At each searching stage, the backbone is assigned to different grids on the basis of its relative position and the resulting docking configurations are scored. The top-scored backbone positions (highlighted by cyan circles in the 2D scheme) are shown as 3D structures in the lower row for each searching resolution and are continued for the next grid search and scoring. The 3D structure example shown here was streptavidin structure (PDB ID: 1STP) with grid searching resolutions of 8.0 Å, 4.0 Å, 2.0 Å, and 1.0 Å. b–d, β-barrel scaffold construction for small-molecule binding. Three geometric constraints (b) were used to describe each backbone hydrogen bond and drive the backbone assembly during Rosetta low-resolution centroid modelling. Backbones generated with all three constraints had a very narrow Φ/Ψ distribution as a result of strong constraints (c, Ramachandran plot in upper left, set 1, density coloured in blue); by omitting N–H–O angle constraint, backbone torsion diversity slightly improved (c, upper right, set 2). These two raw backbone sets yielded few non-redundant RIF docking solutions (d, blue bars). After two rounds of sequence design calculation using Rosetta full-atom force field (Supplementary Methods), regularized backbones (peptide bonds with proper dihedral geometry) and broadened Φ/Ψ distribution (c, Ramachandran plot in the lower row, density coloured in orange) yielded more unique RIF docking solutions (d, orange bars). e, Computed metrics for 42 designs ordered and tested. Results from ab initio folding simulation were scaled to 0.0 to 1.0, in which 1.0 represents a funnel-shaped folding landscape55. f, Alternative ligand-binding conformations revealed by post-design ligand-docking simulations. The lowest-energy docking conformation using the design model (by simply taking out the ligand from the pocket) was similar to the designed DFHBI-binding mode (top left, grey; designed binding mode was circled in grey in the energy landscape in the lower row). Docking simulations using an apo-protein model refined by molecular dynamics simulations revealed an alternative equal-energy docking conformation (top right, green) that is indicated by a green circle in the docking energy landscapes (bottom). Both binding modes rely on three hydrogen-bonding residues from RIF docking (top).
Extended Data Fig. 6 Biochemical and structural characterization of design b10, b32 and b11.
a, Size-exclusion chromatogram of His6-tagged b10 and b32 after Ni-NTA affinity purification. The monodispersed peaks of absorbance at 280 nm of b10 and b32 (cyan and lavender, respectively) have an elution volume compatible with the monomeric β-barrel (14 kDa), on the basis of their relative position to the protein standard peaks (dashed line). Biological replicates were performed with similar observation: n = 4 for b10, n = 5 for b32. b, Comparison of the ligand-binding pocket in the b10 design model (middle, grey) with the crystal structure (left, cyan). The side chain disagreements are highlighted with a dashed black circle on the right panel. c, d, The designed disulfide bond as a stabilizing mechanism. SEC curves of His6-tagged b11 (purple line) and b38 (dark yellow line) were overlaid to show the appearance of a monomer peak for b11 (the same standard as in a was applied here). A disulfide bond connecting the N-terminal helix to a β-strand (Q1C and M59C, circled in d) and four mutations of neighbouring residues were introduced into design b38 (dark yellow) to make design b11 (purple). Biological replicates were performed with similar observation: n = 3 for b38, n = 5 for b11. e, Far-ultraviolet CD spectra of b10, b32 and b11. Left, spectra at different temperatures within one heating–cooling cycle. Right, thermal melting curves (the CD signal of b10 was monitored at 220 nm; b32 and b11 at 226 nm). b11 probably forms an amyloid-like beta structure at 95 °C (left, bottom row) with a negative peak around 226 nm56 and refolds back after cooling to 25 °C. The thermal stability of b11 decreases when the disulfide was reduced with 1 mM tris(2-carboxyethyl) phosphine (TCEP) (right, bottom). Measurements were performed once for each design (n = 1). f, Fluorescence emission spectra of b32, b11 and b11L5F in complex with DFHBI. With 200 μM proteins, b32, b11 and b11L5F can activate 10 μM DFHBI fluorescence by 8-, 12- and 18-fold, respectively. Two biological replicates were performed with similar results. g, The residues designed to interact with DFHBI contribute to b11 and b32 activity. Single or double knockouts of hydrogen-bonding residues (Y71, S23, N17 and T95) and a hydrophobic-packing residue (M15) showed decreased fluorescence intensity at 500 nm in comparison with the wild-type b11 or b32 (WT). Mutants were purified once for activity measurement. h, i, Re-designed five-residue fifth turn in b11L5F. The original bulge-containing ‘AAG’ β-turn in b11 (Extended Data Fig. 3b) was redesigned into a five-residue turn. b11L5F was detected by yeast surface display and flow cytometry (i and Supplementary Data). Yeast cells displaying b11 and b11L5F showed an increased 520-nm fluorescence signal (excited by 488-nm laser, i). Three biological replicates were performed with similar observation.
Extended Data Fig. 7 Deep mutational scanning maps for b11L5F.
a, The complete function (left) and protease stability (middle and right) landscapes of b11L5F. Fluorescence activation scores, trypsin and chymotrypsin stability scores were calculated as described in Supplementary Methods and demonstrated in the Supplementary Data (b11L5F_DMS_analysis.ipy). Data are from two biological replicates with more than tenfold sequencing coverage. Red colour represents beneficial effect whereas mutations coloured in blue are detrimental (relative to the wild-type b11L5F). Wild-type residues at each position are indicated by black dots. b, b11L5F backbone model coloured one the basis of the average stability scores. Glycine backbone Cα are shown as spheres. c, d, Mutational scanning maps of glycine kinks (G25, G43, G53, G55, G81 and G105) and tryptophan corner positions (G9, W9 and R109) (c), and of glycines in the β-turns and prolines (d). e, Statistics of the fluorescence activation and stability scores. The standard deviation between the two replicates used for calculating fluorescence activation scores is smaller than two for most of the data points (left); 95% confidence interval calculated for the proteolysis/stability analysis is less than 0.25 for most the experimental protease half maximal effective concentration (EC50) values (middle and right).
Extended Data Fig. 8 Experimental and computational improvement on the basis of b11L5F.
a–c, Incorporation of point mutations from deep mutational scanning. Beneficial mutations that improve fluorescence activity without compromising protein stability (positive scores relative to wild-type b11L5F; a, left, n = 2 biological replicates) were mapped onto b11L5F backbone model (a, right). b, Purified b11L5F variants incorporating those single, double or triple mutations showed consistently improved fluorescence activity. Binding titration curves were obtained for all six possible triple mutants (right, n = 1 biological measurement). c, b11L5F with V103L, V95A, V83I, C59V and C1S mutations was renamed as ‘b11L5F.1’. d, Characterization of five designs from the second round of design calculation. Three of the five designs (nC1–5) that were based on b11L5F showed improved binding activities by titrating purified proteins into 0.5 μM DFHBI (n = 1 biological sample was used for the measurement). The best variant (nC5) was renamed ‘b11L5F.2’. e, Ligand-docking simulations with the molecular dynamics-refined apo b11L5F.2. Energy landscape was plotted by comparing all the docking conformations to the design model (left). The lowest-energy docking conformations (highlighted in green circle) match the design model (right, design mode in silver and docking model in green). f, g, Characterization of three best variants (mFAP0–2) from combinatorial library selections. f, Yeast cells displaying mFAP proteins incubated with 5 μM DFHBI analysed by flow cytometry (excited by 488-nm laser, n = 1 biological sample was used for the measurement with proper controls). g, Purified proteins showed up to 100-fold fluorescence activation (5 μM protein + 0.5 μM DFHBI, excited at 450 nm and monitored at 500 nm and 510 nm in a plate reader, n = 1 biological measurement). h, Far-UV CD characterization of b11L5F.1, b11L5F.2, mFAP0, mFAP1 and mFAP2. Left, spectra at different temperatures within one heating–cooling cycle. Right, thermal melting curves (CD signals were monitored at 226 nm, spectra were recorded once (n = 1) with internal noise estimation).
Extended Data Fig. 9 Crystal structure of b11L5F_LGL, mFAP0 and mFAP1.
a–g, b11L5F_LGL crystal structure. Protein samples of all six triple mutants in Extended Data Fig. 8b (right) were prepared for crystallization. b11L5F_LGL with V83L/V95G/V103L combination was successfully crystallized. Crystal contacts between protein copies in one asymmetric unit (yellow) were mediated by two tyrosines (stick representation, grey dashed circle); contacts between three asymmetric units (yellow, blue and green) were formed between β-turns (black dashed circle), which might have displaced one of the top β-turns (c). Overall backbone and side chain conformations in the design model matched the crystal structure with a backbone Cα r.m.s.d. of 1.02 Å (b, crystal in yellow and design model in silver), and the designed disulfide bond was present in the crystal structure (d). Ligand density in the crystal structure was ambiguous: 2Fo − Fc omit map showing the electron density after refinement without placing DFHBI (e), the best ligand placement to match the density (f), and designed ligand-binding interactions (silver) overlaid with the crystallized binding pocket (g). h, i, Crystal contacts in the DFHBI-bound structures of mFAP0 (h) and mFAP1 (i). Contacts between protein copies in one asymmetric unit were formed around 40V and 54Y (grey dashed circle) that were introduced for helping crystallization (Extended Data Fig. 10a). Contacts between asymmetric units were formed between β-turns (black dashed circle). j, 2Fo − Fc omit electron density of DFHBI in the mFAP0–DFHBI complex crystal structure. DFHBI density contoured at 1.0σ is clear and matches the planar conformation of the ligand (right). k, Superposition of mFAP0 design model (silver) and the crystal structure (magenta). Hydrogen bonds are indicated with dashed lines. l, Helical capping interactions mediated by P62D mutation in mFAP1 crystal structure.
Extended Data Fig. 10 Mapping of mutations introduced into b11 to yield the final brighter variants, biophysical characterization of mFAP1 and 2, and epifluorescent images.
a, Sequence alignment of b11-based DFHBI-binding fluorescence-activating proteins. Orange boxes indicate mutations or loop insertions introduced by computational design; purple boxes highlight mutations rationally introduced on the basis of the deep mutational scanning maps (Extended Data Figs. 7, 8); green boxes indicate mutations or loop insertions that were incorporated during combinatorial library selections; K40V and K54Y in light blue boxes were introduced to help crystal formation (Extended Data Fig. 9h, i). Despite having hydrophobic residues on the surface, mFAP2 remains soluble at 150 mg ml−1. b, Mutations in the mFAPs mapped on the design models. Common mutations in all three mFAPs are highlighted in bold. c, Absorbance spectra for DFHBI, and the mFAP1–DFHBI and mFAP2–DFHBI complexes (n = 4 biological replicates with similar observations). d, Extinction coefficient determination for DFHBI at 418 nm. e, Normalized absorbance and fluorescence spectra of the mFAP1–DFHBI and mFAP2–DFHBI complex. Data are representative of two biological replicates with similar observations. f, g, Widefield epifluorescence (bottom) and brightfield (top) images of E. coli and yeast cells with 20 μM DFHBI. Untransformed E. coli Lemo21 cells (f, left, n = 2 biological replicates with similar observation) and yeast EBY100 cells displaying ZZ domain (g, left, n = 2 biological replicates with similar observation) were treated with the same amount of DFHBI and imaged in the same way (1000 mA 470-nm LED and 200-ms exposure time).
Supplementary information
Supplementary Information
This file contains a detailed Supplementary Methods section, Supplementary Tables 1-18 and a Supplementary Data section including essential computational command lines and scripts. More supplementary materials can be found on https://dx.doi.org/10.5281/zenodo.1216229
Video 1
Widefield epifluorescence video of live COS-7 cells expressing sec61β-mFAP1. Contrast was adjusted to highlight the peripheral endoplasmic reticulum. Video duration was 5 minutes with one 200 ms exposure every 5 sec. n=2 biological replicates were performed with similar observation
Video 2
Widefield epifluorescence video of live COS-7 cells expressing mito-mFAP2. Video duration was 5 minutes with one 200 ms exposure every 5 sec. n=2 biological replicates were performed with similar observation
Rights and permissions
About this article
Cite this article
Dou, J., Vorobieva, A.A., Sheffler, W. et al. De novo design of a fluorescence-activating β-barrel. Nature 561, 485–491 (2018). https://doi.org/10.1038/s41586-018-0509-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0509-0
Keywords
This article is cited by
-
Opportunities and challenges in design and optimization of protein function
Nature Reviews Molecular Cell Biology (2024)
-
Sparks of function by de novo protein design
Nature Biotechnology (2024)
-
Design of complicated all-α protein structures
Nature Structural & Molecular Biology (2024)
-
Machine learning for functional protein design
Nature Biotechnology (2024)
-
Hallucination of closed repeat proteins containing central pockets
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.